 Back
BackMicrobial Genome Evolution, Mutations, and Horizontal Gene Transfer
Study Guide - Smart Notes

Transposable Elements in Genetic Analysis
Structure and Function of Transposable Elements
Transposable elements, also known as "jumping genes," are DNA sequences that can change their position within the genome. They play a crucial role in genetic variation and genome evolution. - Transposase: An enzyme that catalyzes the movement of transposable elements. - Inverted repeats: Short, identical sequences at both ends of the transposon, necessary for recognition and movement. - Antibiotic resistance gene: Often included in engineered transposons for selection purposes (e.g., kanamycin resistance). - Origin of replication: Allows the plasmid containing the transposon to replicate in host cells. 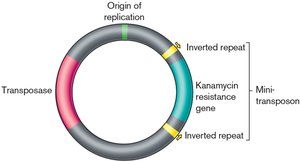
Transposon Mutagenesis and Functional Genomics
Transposon mutagenesis is a technique used to disrupt genes and study their function. Pools of transposon mutants can be introduced into model organisms (such as mice) to identify genes involved in virulence or survival. - Mutant pool generation: Random insertion of transposons creates a library of mutants. - Functional screening: Mutant pools are introduced into a host, and their abundance is measured before and after infection to identify essential genes. - DNA sequencing (TnSeq): Used to quantify the abundance of each mutant and determine which genes are required for survival or virulence. 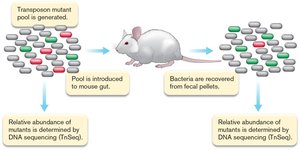
Signature-Tagged Mutagenesis (STM)
STM is a method for identifying virulence genes by tagging transposons with unique DNA sequences. Mutants are pooled, introduced into a host, and recovered to determine which tags (and thus which genes) are missing, indicating loss of function. - Tag insertion: Unique oligonucleotide tags are inserted into transposons. - Amplification and selection: Mutants are amplified and selected for transposon insertion events. - Screening: Mutant pools are screened for loss of function in host environments. 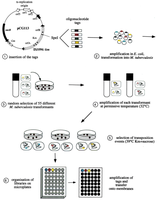
Genome Evolution in Microbes
Mechanisms of Genome Evolution
Microbial genomes are dynamic and evolve through several mechanisms, primarily driven by natural selection. - Horizontal gene transfer (HGT): The movement of genetic material between organisms, often across species or domains. - Gene duplication and divergence: Duplication of genes followed by mutation allows new functions to evolve.
Homology, Duplications, and Divergence
Homologous genes can arise from duplication events, leading to paralogous and orthologous relationships. - Paralogous genes: Genes within the same organism that arose from duplication and have diverged in function. - Orthologous genes: Genes in different species that originated from a common ancestor and retain similar functions. 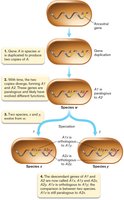
Horizontal Gene Transfer (HGT)
HGT is a major driver of microbial genome evolution, allowing the acquisition of new traits such as antibiotic resistance or pathogenicity. - Interdomain transfer: Genes can be transferred between bacteria, archaea, and eukaryotes. - Genomic islands: Regions of the genome acquired through HGT, often containing clusters of genes for pathogenicity, symbiosis, or fitness. 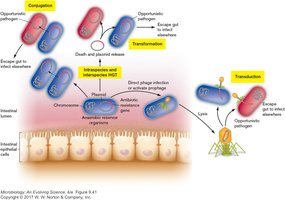
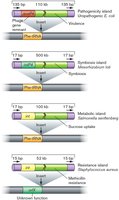
Genome Reduction
Genome reduction involves the loss of genes, often seen in obligate intracellular pathogens. - Pseudogenes: Nonfunctional remnants of genes no longer needed. - Chromosomal black holes: Regions lacking genes compared to closely related species.
Types of Gene Transfer in Bacteria
Transformation, Conjugation, and Transduction
Bacteria can exchange genetic material through three main mechanisms: - Transformation: Uptake of free DNA from the environment. - Conjugation: Transfer of DNA following direct cell-to-cell contact, often via plasmids. - Transduction: Transfer of DNA via bacteriophages (viruses that infect bacteria). 
Mutations and Their Effects
Point Mutations
Point mutations are changes in a single nucleotide and can be classified as: - Transition: Purine to purine (A ↔ G) or pyrimidine to pyrimidine (C ↔ T). - Transversion: Purine to pyrimidine or vice versa (A/G ↔ C/T).
Classes of Mutations
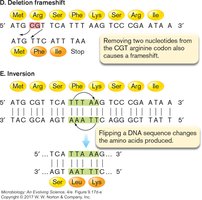
Mutations can have various effects on gene expression and protein function: - Missense mutation: Changes one amino acid to another. - Nonsense mutation: Changes a codon to a stop codon, truncating the protein. - Frameshift mutation: Insertion or deletion of nucleotides alters the reading frame. - Inversion: Flipping a DNA segment changes the amino acids produced.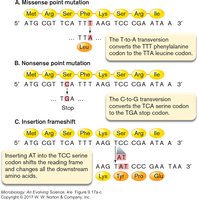 
Mutation Effects on Progeny
A single mutation prior to DNA replication can result in 50% of progeny carrying the mutant gene, assuming no further mutations and equal reproduction rates.
Methyl Mismatch Repair
During methyl mismatch repair, the unmethylated, newly synthesized strand is cleaved, as it is presumed to contain the error.
Defense Mechanisms Against Phage
Restriction-Modification Systems and CRISPR-CAS
Bacteria defend against phage infection using: - Restriction-modification systems: Enzymes that cut foreign DNA and protect self-DNA via methylation. - CRISPR-CAS: Adaptive immune system that targets and destroys invading phage DNA.
Gene Duplication and Divergence
Role in Genome Evolution
Gene duplication allows for the evolution of new functions by freeing one copy from selective constraints. - Superfamilies: Groups of proteins with structural similarities but diverse functions, such as ABC transporters. 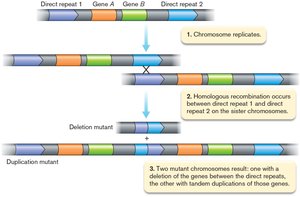

Summary Table: Types of Gene Transfer
Type | Mechanism | Key Features |
|---|---|---|
Transformation | Uptake of free DNA | Requires competent cells |
Conjugation | Cell-to-cell contact | Plasmid transfer |
Transduction | Bacteriophage-mediated | Virus transfers DNA |
Summary Table: Classes of Mutations
Mutation Type | Effect |
|---|---|
Missense | Changes amino acid |
Nonsense | Creates stop codon |
Frameshift | Alters reading frame |
Inversion | Reverses DNA segment |
Summary Table: Paralogous vs. Orthologous Genes
Gene Type | Origin | Function |
|---|---|---|
Paralogous | Gene duplication within species | Divergent functions |
Orthologous | Speciation event | Conserved function across species |
Key Takeaways
- Microbial genomes evolve through horizontal gene transfer and gene duplication/divergence. - Transposable elements are powerful tools for genetic analysis and mutagenesis. - Mutations can have diverse effects, from subtle changes to complete loss of function. - Defense mechanisms such as restriction-modification and CRISPR-CAS protect bacteria from phage infection. - Understanding these processes is essential for studying microbial evolution, pathogenesis, and biotechnology.
