 Back
BackMicrobial Metabolism: Catabolic and Anabolic Pathways, Metabolic Diversity, and Biochemical Identification
Study Guide - Smart Notes

Microbial Metabolism
Other Catabolic Pathways for Oxidizing Nutrients
Microorganisms possess diverse catabolic pathways to oxidize nutrients, allowing them to adapt to various environments and substrates. While glycolysis is the most common pathway, alternatives such as the Entner-Doudoroff and Pentose Phosphate Pathways are also utilized.
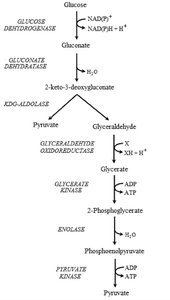
Entner-Doudoroff Pathway: Catabolizes glucose using unique enzymes, primarily found in Gram-negative, obligate aerobic bacteria. This pathway is less efficient, yielding only one ATP per glucose molecule.
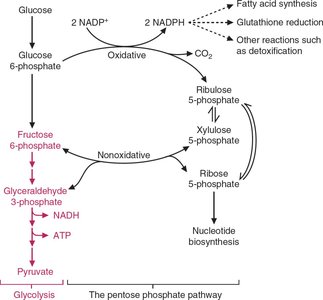
Pentose Phosphate Pathway: Converts 5-carbon sugars to 3- or 6-carbon sugars, which can enter glycolysis. It also provides precursors for nucleotide and amino acid biosynthesis and is used by many bacteria and some eukaryotes.
Key Point: These pathways allow bacteria to metabolize a wider range of carbohydrates and adapt to different ecological niches.
Fermentation
Fermentation is a metabolic process used by organisms that lack a final electron acceptor or a respiratory chain. It allows cells to regenerate NAD+ and sustain ATP production via glycolysis in the absence of oxygen.
Anaerobic Respiration vs Fermentation: Both are anaerobic, but fermentation does not use a respiratory chain, while anaerobic respiration uses an inorganic substance as the final electron acceptor.
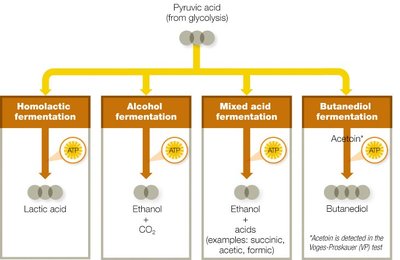
Fermentation Pathways: Use pyruvic acid from glycolysis to regenerate NAD+, enabling continued glycolytic ATP production.
Key Point: Fermentation is essential for energy production in environments lacking oxygen or a functional electron transport chain.
Lactic Acid Fermentation
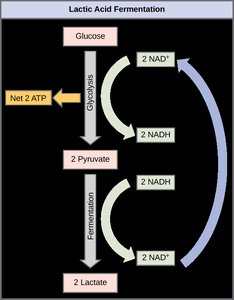
Homolactic fermentation reduces pyruvic acid to lactic acid, regenerating NAD+ and sustaining ATP production. This pathway is used by certain bacteria and human muscle cells under low oxygen conditions.
Example: Lactobacillus bulgaricus and Streptococcus thermophilus (yogurt bacteria), human muscle cells.
Alcohol Fermentation
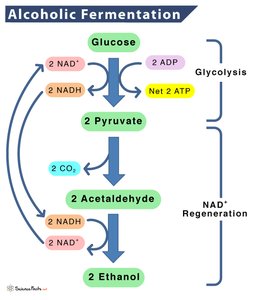
Alcohol fermentation converts pyruvic acid to ethanol and CO2, regenerating NAD+ and sustaining ATP production. This pathway is used by some bacteria, fungi, and plant cells, notably Saccharomyces cerevisiae (yeast).
Example: Bread rising, beer and wine production.
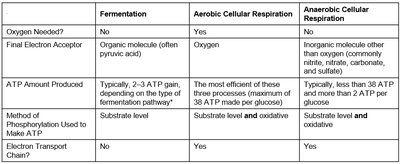
Fermentation vs Aerobic Respiration vs Anaerobic Respiration
Cells utilize different pathways to harvest energy, each with distinct characteristics regarding oxygen use, electron acceptors, and ATP yield.
Fermentation | Aerobic Cellular Respiration | Anaerobic Cellular Respiration | |
|---|---|---|---|
Oxygen Needed? | No | Yes | No |
Final Electron Acceptor | Organic molecule (often pyruvic acid) | Oxygen | Inorganic molecule other than oxygen (e.g., nitrate, nitrite, carbonate, sulfate) |
ATP Amount Produced | Typically, 2-3 ATP gain | Up to 38 ATP per glucose | Less than 38 ATP, more than 2 ATP per glucose |
Method of Phosphorylation | Substrate level | Substrate level and oxidative | Substrate level and oxidative |
Electron Transport Chain? | No | Yes | Yes |
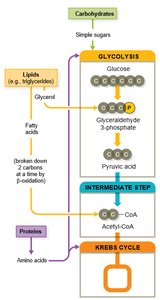
Catabolism of Macromolecules
Microbes can catabolize lipids, proteins, and nucleic acids in addition to carbohydrates. Initial breakdown often occurs extracellularly via exoenzymes, followed by intracellular catabolic pathways.
Exoenzymes: Secreted by bacteria to degrade large macromolecules, aiding in nutrient acquisition and identification of microbes.
Key Point: Detection of exoenzymes (e.g., gelatinase) is useful in microbial identification.

Catabolizing Lipids
Lipids are broken down by lipases into glycerol and fatty acids. Glycerol enters glycolysis, while fatty acids undergo beta-oxidation to form acetyl-CoA, which enters the Krebs cycle.
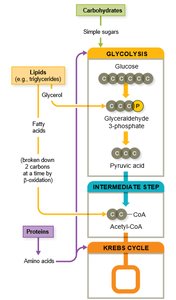
Catabolizing Proteins
Proteins are degraded by proteases and peptidases into amino acids, which can be recycled or further catabolized via deamination and entry into the Krebs cycle.
Catabolizing Nucleic Acids
Nucleic acids are broken down by nucleases into nucleotides. While not a major energy source, organisms often salvage nucleotides for biosynthesis.
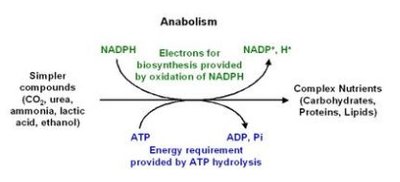
Anabolic Reactions: Biosynthesis
Biosynthetic Pathways
Anabolic reactions build complex biological molecules using ATP and reducing power from NADPH. These pathways are essential for cell growth and maintenance.
NADPH vs NADH: NADPH is primarily used in anabolic reactions, while NADH is used in catabolic reactions.
Key Point: The phosphate group in NADPH allows enzymes to distinguish it from NADH.
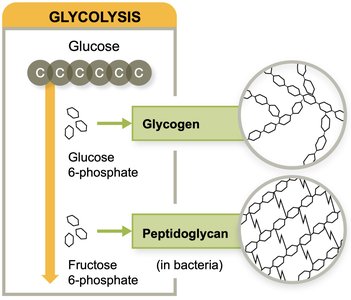
Polysaccharide Biosynthesis
Polysaccharide biosynthesis starts with simple sugars. Gluconeogenesis builds glucose from non-sugar precursors, while intermediates from glycolysis and the Krebs cycle are used to synthesize glycogen and peptidoglycan.
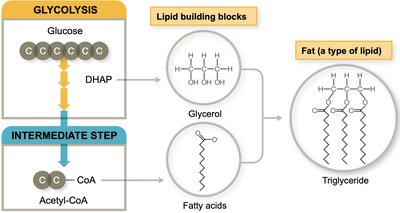
Lipid Biosynthesis
Lipid biosynthesis begins with carbohydrate catabolism intermediates. Glycerol is made from DHAP (a glycolysis intermediate), and fatty acids are synthesized by linking acetyl-CoA molecules.
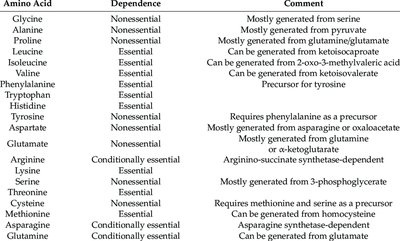
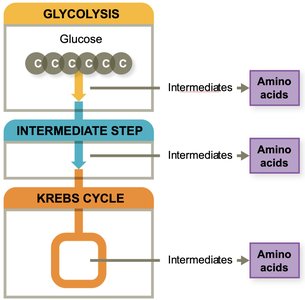
Amino Acid Biosynthesis
Cells synthesize nonessential amino acids via amination, while essential amino acids must be obtained from the environment. The degree of dependence varies among species.
Example: Most E. coli strains can synthesize all amino acids, while Lactobacillus species require many from their environment.

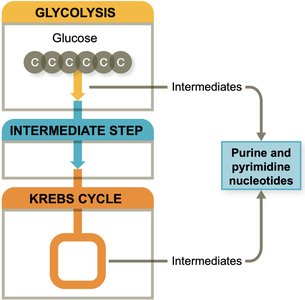
Nucleic Acid Biosynthesis
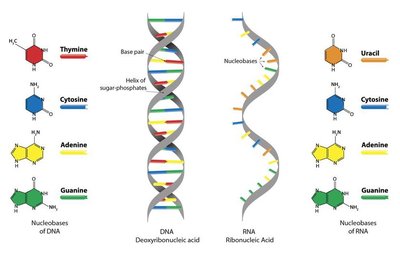
Purines (adenine, guanine) and pyrimidines (uracil, thymine, cytosine) are essential for nucleic acids and energy molecules like ATP. While often recycled, they can be synthesized de novo.
The Interconnected Web of Metabolism
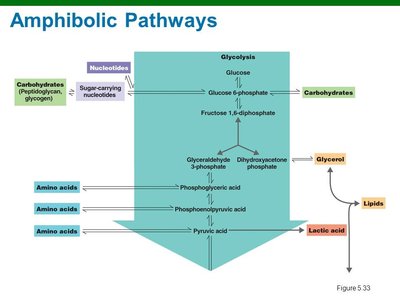
Amphibolic Pathways
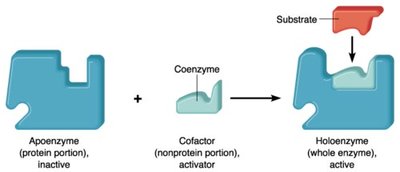
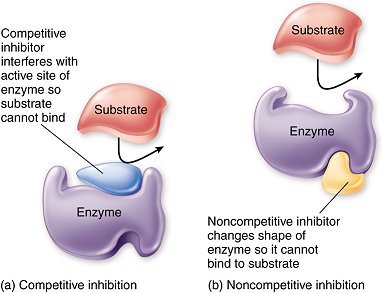
Amphibolic pathways function in both anabolism and catabolism, allowing cells to balance energy production and biosynthesis. Regulation occurs via cofactors and enzyme activity.
Key Point: NAD+ is used in catabolic pathways, NADP+ in anabolic pathways. Enzyme activity is regulated by feedback inhibition, allosteric mechanisms, and phosphorylation.
Metabolic Diversity
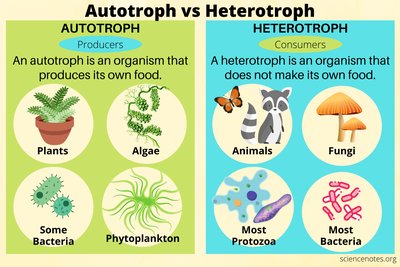
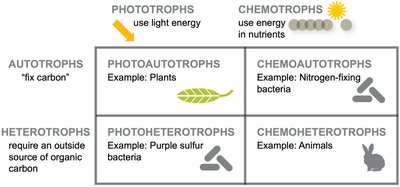
Carbon Fixation: Autotrophs vs Heterotrophs
Microbes are classified based on their ability to fix carbon. Autotrophs synthesize organic carbon from inorganic sources, while heterotrophs require external organic carbon.
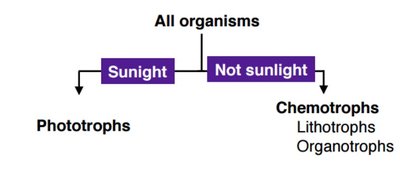
Energy Acquisition: Phototrophs, Chemotrophs, Mixotrophs
Microbes obtain energy through various mechanisms. Phototrophs harvest energy from light, chemotrophs from chemical bonds, and mixotrophs can switch between modes.
Metabolic Diversity Table
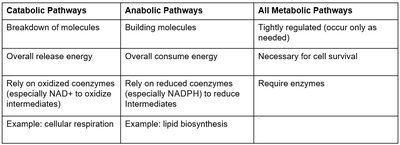
Catabolic Pathways | Anabolic Pathways | All Metabolic Pathways |
|---|---|---|
Breakdown of molecules | Building molecules | Tightly regulated (occur only as needed) |
Overall release energy | Overall consume energy | Necessary for cell survival |
Rely on oxidized coenzymes (especially NAD+) | Rely on reduced coenzymes (especially NADPH) | Require enzymes |
Example: cellular respiration | Example: lipid biosynthesis |
Using Metabolic Properties to Identify Bacteria
Biochemical Tests
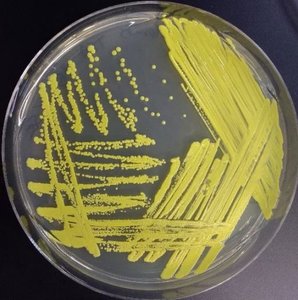
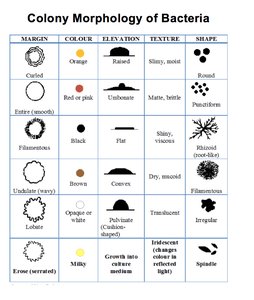
Biochemical tests are essential for identifying unknown microbes by detecting metabolic end products, intermediates, or specific enzymes. These tests range from specialized media to molecular and genetic procedures.
Key Point: Metabolic profiles act as biochemical fingerprints for microbial identification.
Amino Acid Catabolism Tests
These tests detect deaminases and decarboxylases, useful for identifying enteric bacteria such as Salmonella and Proteus species.
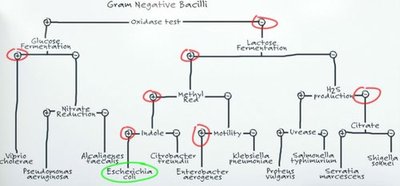
Fermentation Tests
Fermentation tests use media containing protein, a single carbohydrate, a pH indicator, and sometimes a Durham tube to capture gas. Acidic end products lower the pH, changing the color of the medium.
Example: Methyl red/Voges-Proskauer (MRVP) test distinguishes mixed acid and butanediol fermentation.
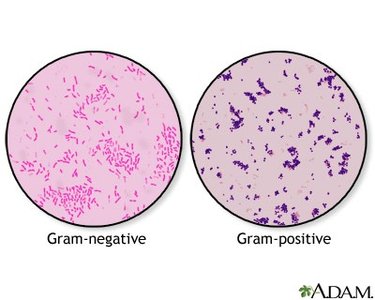
MacConkey agar: Selective and differential medium for gram-negative, enteric bacteria, differentiating based on lactose fermentation.
Oxidase and Catalase Tests
Oxidase test detects cytochrome c oxidase, while catalase test identifies the enzyme catalase, which breaks down hydrogen peroxide. These tests help differentiate bacterial species.
Rapid Identification Techniques
Systems such as the Analytical Profile Index (API®) and Enterotubes allow for rapid identification of microbes based on metabolic properties.
Summary
Microbial metabolism encompasses a wide range of catabolic and anabolic pathways, enabling microbes to adapt to diverse environments and substrates. Understanding these pathways and their regulation is crucial for identifying microbes and appreciating their metabolic diversity.
