 Back
BackMicrobial Metabolism, Catabolism, Anabolism, and Genetic Regulation
Study Guide - Smart Notes

CHAPTER 10: METABOLISM
Overview of Metabolism
Metabolism encompasses all chemical reactions occurring within a cell, including both energy-producing and energy-consuming processes. These reactions are essential for cellular function and survival.
Redox Reactions: Involve oxidation (loss of electrons) and reduction (gain of electrons).
Electron Transport Chain (ETC): Central to ATP production via oxidative phosphorylation.
Types of Cellular Work:
Chemical work: Synthesis of molecules.
Transport work: Movement of substances across membranes.
Mechanical work: Cellular movement (e.g., flagella, cytoplasmic streaming).
ATP: The universal energy currency, composed of adenine, ribose, and three phosphate groups.
Thermodynamics in Metabolism
Cellular reactions obey the laws of thermodynamics, which govern energy transformations.
First Law: Energy cannot be created or destroyed.
Second Law: Energy transfers increase entropy (disorder).
Free Energy (ΔG): Determines spontaneity of reactions.
: Exergonic (spontaneous)
: Endergonic (non-spontaneous)
Standard Free Energy (ΔG°′): Indicates spontaneity under standard conditions.
Redox Reactions and Electron Flow
Reduction Potential (E₀′): Measures tendency to gain/lose electrons.
More negative E₀′: Better electron donor.
More positive E₀′: Better electron acceptor.
Electrons flow from negative to positive E₀′.
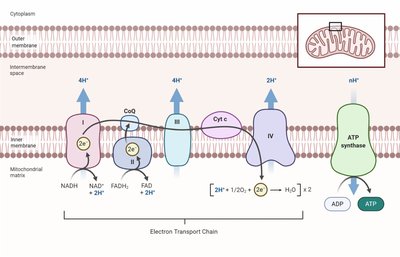
Electron Transport Chain (ETC)
The ETC is a series of electron carriers that transfer electrons, release energy, and synthesize ATP.
Location: Plasma membrane (prokaryotes), mitochondria/chloroplasts (eukaryotes).
Carriers: NAD⁺/NADP⁺, FAD/FMN, Coenzyme Q, cytochromes, iron-sulfur proteins.
Biochemical Pathways
Linear: Direct progression from start to end.
Cyclic: Repeats (e.g., TCA cycle).
Branched: Multiple products.
Enzymes and Regulation
Enzymes are protein catalysts that speed up reactions without being consumed.
Structure: Apoenzyme (protein), cofactor (helper), holoenzyme (active form).
Activity depends on: Substrate concentration, pH, temperature.
Regulation:
Competitive inhibition: Competes for active site.
Noncompetitive inhibition: Binds elsewhere.
Allosteric regulation: Positive/negative effectors.
Covalent modification: Adds/removes groups (e.g., phosphate).
Feedback inhibition: End product inhibits pathway.
Ribozymes and Metabolic Regulation
Ribozymes: RNA molecules acting as enzymes (e.g., self-splicing RNA).
Regulation Mechanisms:
Metabolic channeling: Organizes enzymes spatially.
Gene regulation: Controls enzyme synthesis.
Enzyme activity control: Modifies enzyme function.
CHAPTER 11: CATABOLISM
Catabolic Processes
Catabolism involves breaking down complex molecules to simpler ones, generating ATP, NADH, and building blocks for biosynthesis.
Sources:
Carbon: Autotrophs (CO₂), heterotrophs (organic carbon).
Energy: Phototrophs (light), chemotrophs (chemicals).
Electrons: Lithotrophs (inorganic), organotrophs (organic).
Nutritional Types:
Photolithoautotroph: Light, CO₂, inorganic electron donor.
Photoorganoheterotroph: Light, organic carbon, organic electron donor.
Chemolithoautotroph: Inorganic electron donor, CO₂.
Chemolithoheterotroph: Inorganic electron donor, organic carbon.
Chemoorganoheterotroph: Organic electron donor, organic carbon (most pathogens).
Fueling Reactions
Respiration: Uses ETC; aerobic (O₂ as final acceptor), anaerobic (other acceptors). Produces most ATP.
Fermentation: No ETC; uses organic molecules as acceptor. Produces less ATP.
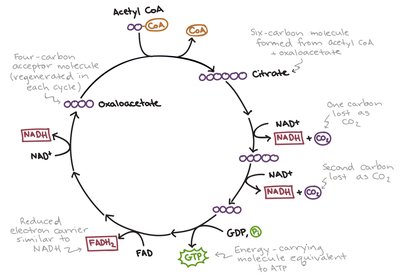
Glycolysis and TCA Cycle
Glycolysis: Glucose is converted to 2 pyruvate.
TCA Cycle: Per acetyl-CoA: 3 NADH, 1 FADH₂, 1 GTP (ATP), 2 CO₂.
ETC and ATP Production
Oxidative Phosphorylation: Most ATP produced.
Chemiosmosis: ETC pumps H⁺ out, creating proton motive force (PMF).
ATP Synthase: H⁺ flows back in to make ATP.
ATP Yield: 32 ATP per glucose in eukaryotes.
Anaerobic Respiration and Fermentation
Anaerobic Respiration: Uses ETC, final acceptor not O₂. Less ATP than aerobic.
Fermentation: No ETC, regenerates NAD⁺, ATP from substrate-level phosphorylation.
Lactic acid: Produces lactate.
Alcoholic: Produces ethanol and CO₂.
Mixed acid: Produces multiple acids.
Butanediol: Produces butanediol.
Catabolism of Other Molecules
Carbohydrates: Converted to glycolysis intermediates.
Lipids: Glycerol enters glycolysis; fatty acids undergo beta oxidation to acetyl-CoA.
Proteins: Deamination removes NH₂ group.
Photosynthesis
Light Reactions: Produce ATP and NADPH.
Dark Reactions: Convert CO₂ to organic molecules.
Oxygenic: Produces O₂, electron source is H₂O.
Anoxygenic: Does not produce O₂, electron source is H₂S.
Cyclic: Produces ATP only.
Noncyclic: Produces ATP and NADPH.
Rhodopsin Phototrophy: Light pumps H⁺ to PMF, no ETC.
CHAPTER 12: ANABOLISM
Anabolic Processes
Anabolism uses energy from catabolism to build complex molecules via biosynthetic pathways.
Principles:
Builds from small to large molecules (monomers to macromolecules).
Many enzymes function in both directions.
Cofactors: Catabolism uses NADH; anabolism uses NADPH.
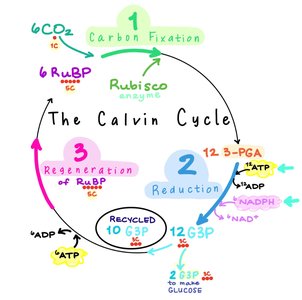
Calvin-Benson Cycle
The Calvin-Benson cycle is the primary pathway for carbon fixation in photoautotrophs.
Carboxylation Phase: CO₂ + RuBP forms 3-PGA (enzyme: RuBisCO).
Reduction Phase: 3-PGA is reduced to G3P using ATP and NADPH.
Regeneration Phase: Reforms RuBP.
Per CO₂: 3 ATP + 2 NADPH used.
Precursor Metabolites and Biosynthesis
Precursor Metabolites: Small molecules from glycolysis and TCA cycle used to build amino acids, nucleotides, sugars.
Carbohydrate Synthesis: Gluconeogenesis makes glucose from non-carbohydrates; uses many glycolysis enzymes and three unique bypass steps.
UDP-glucose used for sugar synthesis.
ATP and UTP required.
Peptidoglycan Synthesis: Uses UDP derivatives and bactoprenol; cross-linking via transpeptidation (antibiotics target this step).
Amino Acid, Nucleotide, and Lipid Synthesis
Amino Acids: Built from carbon skeleton + NH₃ (+ sometimes S).
Nitrogen assimilation: Sources are ammonia and nitrate; fixation uses nitrogenase (requires ATP).
Sulfur assimilation: Source is sulfate; converted to cysteine.
Nucleotide Synthesis:
Purines: Two rings (A, G), built on ribose.
Pyrimidines: One ring (C, T, U), ring formed first then attached.
Nucleoside: Base + sugar; nucleotide: nucleoside + phosphate.
Lipids:
Fatty acids made from acetyl-CoA + malonyl-CoA (uses NADPH and ACP).
Saturated (no double bonds), unsaturated (double bonds).
Major lipids: Triacylglycerols (storage), phospholipids (membranes), sterols (eukaryotes), isoprenoids (archaea).
LPS: Gram-negative bacteria (Lipid A + core + O-antigen).
CHAPTER 13: BACTERIAL GENOME REPLICATION AND EXPRESSION
Discovery of Genetic Material
Frederick Griffith: Demonstrated transformation in bacteria.
Oswald Avery: Proved DNA is genetic material.
Hershey and Chase: Confirmed DNA carries genetic information.
DNA, RNA, and Protein Structure
DNA: Double-stranded, deoxyribose, bases A, T, G, C.
RNA: Single-stranded, ribose, bases A, U, G, C.
Base Pairing: A-T (2 bonds), G-C (3 bonds).
Protein Structure:
Primary: Sequence of amino acids.
Secondary: Alpha helix/beta sheet.
Tertiary: 3D shape.
Quaternary: Multiple chains.
DNA Replication
Semiconservative: One old and one new strand.
Bidirectional: From one origin.
Key Enzymes:
Helicase: Unwinds DNA.
SSB: Stabilizes strands.
Primase: Makes RNA primer.
DNA polymerase: Synthesizes DNA (5’ to 3’).
Ligase: Joins fragments.
Leading Strand: Continuous synthesis.
Lagging Strand: Okazaki fragments.
Gene Structure and Expression
Promoter: RNA polymerase binding site.
Leader: Not translated.
Coding Region: Encodes protein.
Start Codon: AUG.
Stop Codons: UAA, UAG, UGA.
Transcription and Translation
Transcription: DNA to RNA (enzyme: RNA polymerase).
Initiation: Sigma factor binds promoter.
Elongation: RNA synthesized 5’ to 3’.
Termination: Rho-dependent or independent.
Operon: Multiple genes under one promoter.
Genetic Code: Codon (3 bases) specifies 1 amino acid; degenerate (multiple codons per amino acid).
Translation: mRNA to protein at ribosome.
tRNA: Carries amino acids (anticodon matches codon).
Ribosome sites: A (entry), P (growing chain), E (exit).
Initiation, elongation, termination.
First amino acid: fMet (bacteria).
Secretion Systems: Move proteins out of cell.
CHAPTER 14: REGULATION OF CELLULAR PROCESSES
Gene Regulation
Cells regulate gene expression to adapt to environmental changes and conserve resources.
Constitutive Genes: Always ON.
Regulated Genes: Turned ON/OFF as needed.
Inducible Genes: Turned ON by inducer (e.g., lactose metabolism).
Repressible Genes: Turned OFF by corepressor (e.g., amino acid synthesis).
Transcriptional Control
Negative Control: Repressor blocks RNA polymerase.
Positive Control: Activator helps RNA polymerase bind.
Operon: Group of genes controlled together.
Lac, Trp, and Ara Operons
Lac Operon: Controls lactose metabolism.
No lactose: Repressor binds, transcription OFF.
Lactose present: Allolactose binds repressor, transcription ON.
Glucose present: Low cAMP, CAP inactive, transcription LOW.
Low glucose: High cAMP, CAP active, transcription HIGH.
Trp Operon: Controls tryptophan synthesis.
No tryptophan: Operon ON.
Tryptophan present: Acts as corepressor, operon OFF.
Ara Operon: Positive and negative control by AraC protein depending on arabinose presence.
Translation Regulation and Global Control
Riboswitches: mRNA changes shape to block ribosome binding.
RNA Thermometers: Temperature changes RNA structure, affecting translation.
Small RNAs (sRNA): Bind mRNA to inhibit/enhance translation.
Global Regulation: Controls many genes at once (regulon).
Two-component system: Sensor kinase and response regulator.
Phosphorelay: Multi-step signaling pathway.
Sigma factors: Direct RNA polymerase to specific genes.
Second messengers: cAMP, ppGpp, c-di-GMP.
Other Regulatory Mechanisms
Chemotaxis: Movement toward/away from stimuli (MCP, CheA, CheY proteins).
Quorum Sensing: Cell communication for virulence, biofilm production.
Sporulation: Triggered by starvation, controlled by sigma factors and phosphorelay (Spo0A regulator).
Defense Systems: Restriction-modification (cuts foreign DNA), CRISPR-Cas (adaptive immunity).
Summary Table: Metabolic Pathways and Regulation
Pathway | Main Function | Key Products | Regulation |
|---|---|---|---|
Glycolysis | Glucose breakdown | ATP, NADH, pyruvate | Feedback inhibition |
TCA Cycle | Energy production, biosynthesis | NADH, FADH₂, GTP, CO₂ | Allosteric regulation |
ETC | ATP synthesis | ATP, H₂O | Substrate availability |
Calvin Cycle | Carbon fixation | G3P, RuBP | Enzyme activity |
Lac Operon | Lactose metabolism | Enzymes for lactose use | Inducer (allolactose), CAP/cAMP |
Trp Operon | Tryptophan synthesis | Tryptophan | Corepressor (tryptophan), attenuation |
Additional info: Academic context was added to clarify pathway functions, regulatory mechanisms, and enzyme roles for completeness.
