 Back
BackMicrobial Metabolism: Core Concepts and Pathways
Study Guide - Smart Notes

Microbial Metabolism
Introduction to Metabolism
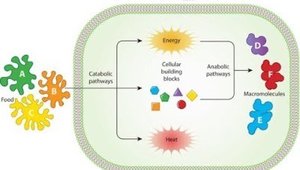
Metabolism encompasses all controlled biochemical reactions that occur within a microbe. The ultimate function of metabolism is to enable the reproduction and survival of the organism by managing energy and molecular resources.
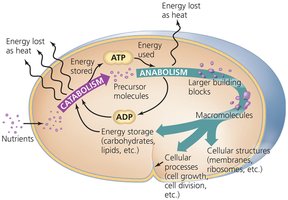
Catabolism: The breakdown of larger molecules into smaller products, releasing energy (exergonic).
Anabolism: The synthesis of large molecules from smaller products, requiring energy input (endergonic).
These processes are interconnected, with catabolic reactions providing the energy and building blocks for anabolic reactions.
Basic Chemical Reactions Underlying Metabolism
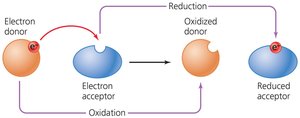
Oxidation and Reduction (Redox) Reactions
Redox reactions involve the transfer of electrons from an electron donor to an electron acceptor. These reactions are fundamental to energy transfer in cells and always occur simultaneously.
Electron carriers are molecules that transport electrons within the cell, often in the form of hydrogen atoms.
Key electron carriers include NAD+, NADP+, and FAD.
The Roles of Enzymes in Metabolism
Enzyme Structure and Function
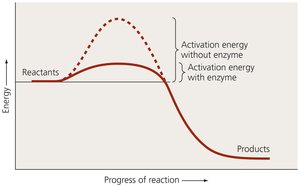
Enzymes are organic catalysts that increase the likelihood of chemical reactions by lowering activation energy. They are not permanently altered during reactions and can be used repeatedly.
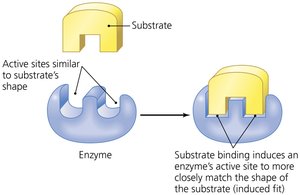
Enzymes are highly specific for their substrates.
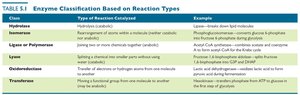
They are classified into six categories based on their mode of action: Hydrolases, Isomerases, Ligases/Polymerases, Lyases, Oxidoreductases, and Transferases.
Class | Type of Reaction Catalyzed | Example |
|---|---|---|
Hydrolases | Hydrolysis (catabolic) | Lipase—breaks down lipid molecules |
Isomerases | Rearrangement of atoms within a molecule | Phosphoglucoisomerase—converts glucose-6-phosphate into fructose-6-phosphate during glycolysis |
Ligases/Polymerases | Joining two molecules (anabolic) | Acetyl-CoA synthetase—combines acetate and CoA to form acetyl-CoA for the Krebs cycle |
Lyases | Splitting a chemical into smaller parts without hydrolysis | Fructose-1,6-bisphosphate aldolase—splits fructose-1,6-bisphosphate into G3P and DHAP in glycolysis |
Oxidoreductases | Transfer of electrons or hydrogen atoms from one molecule to another | Lactic acid dehydrogenase—oxidizes lactic acid to form pyruvic acid during fermentation |
Transferases | Moving a functional group from one molecule to another | Hexokinase—transfers phosphate from ATP to glucose in the first step of glycolysis |
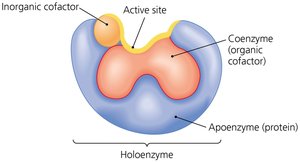
Cofactors and Coenzymes
Many enzymes require non-protein helpers called cofactors to function. These can be inorganic ions or organic molecules (coenzymes).
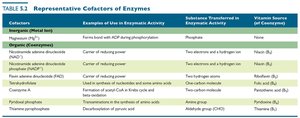
Cofactor | Example of Use | Substance Transferred | Vitamin Source |
|---|---|---|---|
Magnesium (Mg2+) | Forms bond with ATP during phosphorylation | Phosphate | None |
NAD+ (coenzyme) | Center of redox power | Two electrons and a hydrogen ion | Niacin (B3) |
FAD (coenzyme) | Center of redox power | Two hydrogen atoms | Riboflavin (B2) |
Enzyme Activity and Regulation
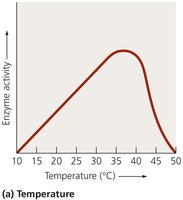
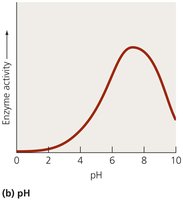
Enzyme activity is influenced by several factors, including temperature, pH, substrate concentration, and the presence of inhibitors or activators.
Enzymes have optimal temperature and pH ranges for activity.
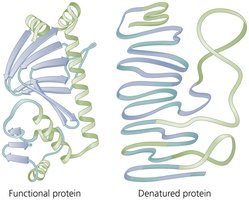
High temperatures or extreme pH can denature enzymes, rendering them inactive.
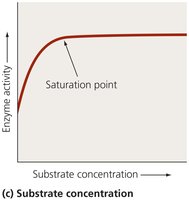
Enzyme activity increases with substrate concentration until a saturation point is reached.
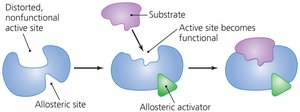
Enzyme Regulation: Allosteric Control and Inhibition
Enzymes can be regulated by molecules that bind to sites other than the active site (allosteric sites), causing conformational changes that activate or inhibit the enzyme. Inhibitors can be competitive (binding to the active site) or noncompetitive (binding elsewhere).
Carbohydrate Catabolism
Glycolysis
Glycolysis is the metabolic pathway that breaks down glucose into pyruvic acid, generating ATP and NADH. It occurs in the cytoplasm and consists of three stages: energy-investment, lysis, and energy-conserving.
Net gain: 2 ATP, 2 NADH, and 2 pyruvic acid molecules per glucose.
Substrate-level phosphorylation is the direct transfer of phosphate to ADP to form ATP.
Cellular Respiration
Cellular respiration is the complete oxidation of substrates to produce ATP. It includes three main stages: synthesis of acetyl-CoA, the Krebs cycle, and the electron transport chain (ETC).
Synthesis of acetyl-CoA: Converts pyruvate to acetyl-CoA, producing NADH and CO2.
Krebs cycle: Occurs in the cytosol (prokaryotes) or mitochondrial matrix (eukaryotes), generating ATP, NADH, FADH2, and CO2.
Electron Transport Chain (ETC): Series of redox reactions that generate a proton gradient used to produce ATP via chemiosmosis.
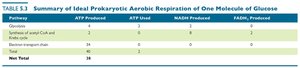
Pathway | ATP Produced | ATP Used | NADH Produced | FADH2 Produced |
|---|---|---|---|---|
Glycolysis | 4 | 2 | 2 | 0 |
Synthesis of acetyl-CoA and Krebs cycle | 2 | 0 | 8 | 2 |
Electron transport chain | 34 | 0 | 0 | 0 |
Net Total | 38 | 2 | 10 | 2 |
Alternative Pathways: Entner-Doudoroff and Pentose Phosphate
Some bacteria use the Entner-Doudoroff (ED) pathway instead of glycolysis, producing 1 ATP, 1 NADH, and 1 NADPH per glucose. The pentose phosphate pathway is another alternative, generating precursor metabolites and NADPH for biosynthesis.
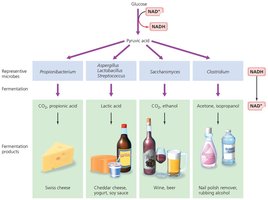
Fermentation
Fermentation is an anaerobic process that allows cells to regenerate NAD+ by transferring electrons to organic molecules, producing various end products such as lactic acid, ethanol, and propionic acid.
Aerobic Respiration | Anaerobic Respiration | Fermentation | |
|---|---|---|---|
Oxygen Required? | Yes | No | No |
Type of Phosphorylation | Substrate-level and oxidative | Substrate-level and oxidative | Substrate-level |
Final Electron Acceptor | Oxygen | NO3-, SO42-, CO32-, or other inorganic molecules | Cellular organic molecules |
ATP Yield (per glucose) | 38 (prokaryotes) | 2-36 | 2 |

Other Catabolic Pathways
Lipids and proteins can also be catabolized to generate energy and precursor metabolites, feeding into glycolysis and the Krebs cycle.
Photosynthesis
Overview and Structures
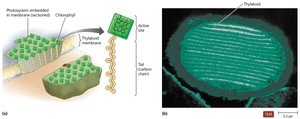
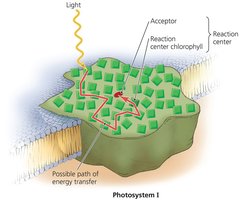
Photosynthesis is the process by which organisms synthesize organic molecules from CO2 and H2O using light energy. Chlorophylls are the main pigments involved, and photosystems are complexes that capture light energy.
Photosystems are embedded in thylakoid membranes (prokaryotes: cytoplasmic membrane invaginations; eukaryotes: chloroplasts).
Photosystems are arranged in stacks called grana, with the stroma as the surrounding space.
Light-Dependent and Light-Independent Reactions
Photosynthesis consists of two main stages:
Light-dependent reactions: Use light energy to generate ATP and NADPH via photophosphorylation (cyclic or noncyclic).
Light-independent reactions: Use ATP and NADPH to fix CO2 into glucose through the Calvin-Benson cycle (fixation, reduction, regeneration of RuBP).
Type | Source of Phosphate | Source of Energy | Location in Eukaryotic Cell | Location in Prokaryotic Cell |
|---|---|---|---|---|
Substrate-Level Phosphorylation | Organic molecule | High-energy phosphate bond of donor | Cytosol and mitochondrial matrix | Cytosol |
Oxidative Phosphorylation | Inorganic phosphate (PO43-) | Proton motive force | Inner membrane of mitochondrion | Cytoplasmic membrane |
Photophosphorylation | Inorganic phosphate (PO43-) | Proton motive force | Thylakoid of chloroplast | Thylakoid of membrane |

Other Anabolic Pathways
Anabolic reactions synthesize complex molecules from simpler ones, requiring energy (usually from ATP) and precursor metabolites. Many anabolic pathways are the reverse of catabolic pathways and are termed amphibolic if they can proceed in both directions.
Metabolite | Pathway That Generates | Macromolecules Synthesized | Functional Use |
|---|---|---|---|
Glucose 6-Phosphate | Glycolysis | Polysaccharides, nucleic acids | Cell wall, energy storage |
Ribose 5-Phosphate | Pentose phosphate pathway | DNA, RNA, amino acids | Genomic information |
Acetyl-CoA | Krebs cycle | Fatty acids, isoprenoids | Membranes, energy storage |

Integration and Regulation of Metabolic Function
Cells regulate metabolism by controlling enzyme synthesis and activity, substrate availability, and feedback inhibition. Eukaryotic cells compartmentalize metabolic pathways within organelles, and cells often use allosteric regulation and feedback inhibition to maintain metabolic balance.
Control of gene expression: Regulates the amount and timing of enzyme production.
Control of metabolic expression: Regulates the activity of enzymes once produced.
