 Back
BackMicrobial Metabolism: Fundamentals, Pathways, and Biosynthesis
Study Guide - Smart Notes

Microbial Metabolism
Fundamentals of Metabolism
Microbial metabolism encompasses all biochemical reactions necessary for life, including both catabolic (energy-yielding) and anabolic (biosynthetic) processes. These reactions are tightly coupled through the flow of energy and electrons, enabling cells to grow, reproduce, and maintain their structures.
Metabolism: The sum of all chemical reactions in a cell, divided into catabolism (breakdown of molecules to release energy) and anabolism (synthesis of cellular components).
Energy is conserved primarily in the form of adenosine triphosphate (ATP), which fuels cellular processes.
Electron donors transfer electrons to electron acceptors, driving metabolic reactions.
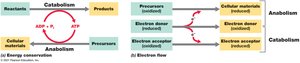
Requirements for Life
All cells require water, carbon, nutrients, free energy, and reducing power to sustain metabolism.
Free energy (ΔG): The energy available to do work. Reactions can be exergonic (release energy, ΔG < 0) or endergonic (require energy, ΔG > 0).
Reducing power: The ability to donate electrons, essential for both catabolic and anabolic reactions.
Catabolic pathways are exergonic and generate ATP, while anabolic pathways are endergonic and consume ATP.
Redox Reactions and Electron Flow
Redox (reduction-oxidation) reactions are central to metabolism, involving the transfer of electrons from donors to acceptors. The difference in reduction potential between these pairs determines the amount of energy released.
Electron donor: Substance that loses electrons (is oxidized).
Electron acceptor: Substance that gains electrons (is reduced).
Example: In aerobic respiration, glucose is oxidized and O2 is reduced.
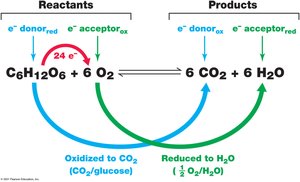
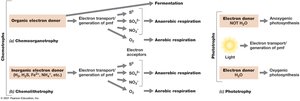
Metabolic Classes of Microorganisms
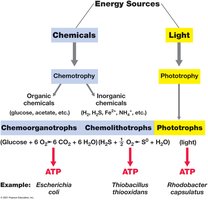
Microorganisms are classified based on their energy and carbon sources:
Phototrophs: Obtain energy from light (photosynthesis).
Chemotrophs: Obtain energy from chemical compounds.
Chemoorganotrophs: Use organic compounds as energy and electron sources.
Chemolithotrophs: Use inorganic compounds as energy and electron sources.
Heterotrophs: Obtain carbon from organic compounds.
Autotrophs: Obtain carbon from CO2 (primary producers).
Electron Transfer Reactions
Reduction Potentials and the Redox Tower
The reduction potential (E0') measures a substance's tendency to accept electrons. The redox tower arranges redox couples from most negative (best electron donors) to most positive (best electron acceptors).
Electrons flow from donors with low (negative) E0' to acceptors with high (positive) E0'.
The greater the difference in E0' between donor and acceptor, the more energy is released.
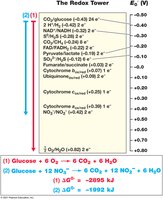

Redox Couple | E0' (V) |
|---|---|
CO2/glucose | -0.43 |
2H+/H2 | -0.42 |
NAD+/NADH | -0.32 |
NO3-/NO2- | +0.42 |
1/2 O2/H2O | +0.82 |
... (see full table for more) | ... |

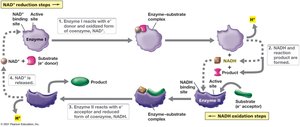
Electron Carriers: NAD+/NADH and NADP+/NADPH
Electron carriers such as NAD+ and NADP+ shuttle electrons between metabolic reactions. NADH is a strong electron donor, while NAD+ is a weak acceptor.
Reduction: NAD+ + 2e- + 2H+ → NADH + H+
These coenzymes allow diverse electron donors and acceptors to interact in metabolism.
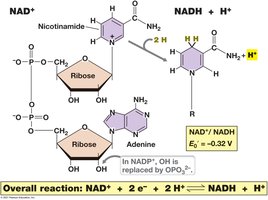
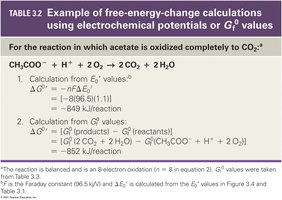
Calculating Changes in Free Energy
Relationship Between Redox Potential and Free Energy
The amount of free energy released in a redox reaction can be calculated using the difference in reduction potentials:
Equation: where n = number of electrons transferred, F = Faraday constant (96.5 kJ/V·mol), and ΔE0' = difference in reduction potential.
Alternatively, ΔG0 can be calculated from the free energy of formation (Gf0) of reactants and products.


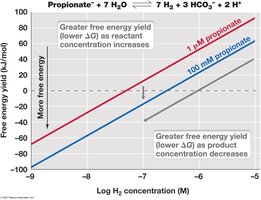
Free Energy in Natural Conditions
Actual free energy change (ΔG) in cells depends on the concentrations of reactants and products, not just standard conditions. Only exergonic reactions (ΔG < 0) can be used by cells to conserve energy.
Cellular Energy Conservation
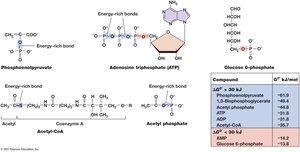
ATP and Energy-Rich Compounds
ATP is the primary energy currency in cells, containing two high-energy phosphate bonds. Other energy-rich compounds include phosphoenolpyruvate and acetyl-CoA.
Mechanisms of ATP Synthesis
Substrate-level phosphorylation: Direct transfer of a phosphate group to ADP from a high-energy substrate.
Oxidative phosphorylation: ATP synthesis driven by a proton motive force generated by electron transport.
Photophosphorylation: Light-driven generation of a proton motive force for ATP synthesis.
Catalysis and Enzymes
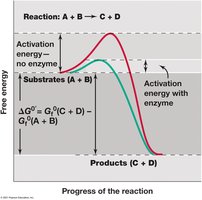
Enzyme Function and Activation Energy
Enzymes are biological catalysts that lower the activation energy of reactions, increasing their rate without being consumed. They are highly specific, typically proteins, and function by binding substrates at their active sites.
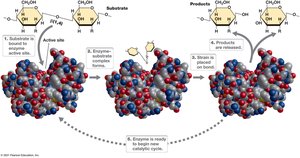
Some enzymes require nonprotein molecules for activity:
Prosthetic groups: Tightly bound, often permanent (e.g., heme in cytochromes).
Coenzymes: Loosely bound, often derived from vitamins (e.g., NAD+).
Enzyme-catalyzed reactions are theoretically reversible, but highly exergonic or endergonic reactions require different enzymes for the reverse process.
Catabolism: Chemoorganotrophs
Glycolysis (Embden–Meyerhof–Parnas Pathway)
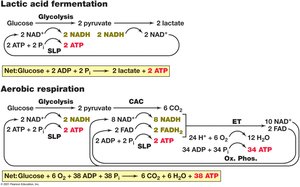
Glycolysis is a nearly universal pathway for glucose catabolism, converting glucose to pyruvate and generating ATP and NADH. It consists of two stages: a preparatory phase and a payoff phase.
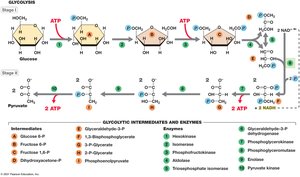
Net yield per glucose: 2 ATP, 2 NADH, 2 pyruvate.
Redox balance is maintained by further reduction of pyruvate (fermentation) or by respiration.
The Citric Acid Cycle (CAC)
The CAC oxidizes pyruvate to CO2, generating NADH, FADH2, and ATP. It also provides precursors for biosynthesis.
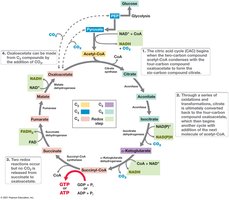
Per pyruvate: 2 CO2, 3 NADH, 1 FADH2, 1 ATP (or GTP).
Cycle intermediates are used for amino acid, nucleotide, and other biosyntheses.
The Glyoxylate Cycle
The glyoxylate cycle enables the catabolism of C2 compounds (e.g., acetate) by bypassing the decarboxylation steps of the CAC, allowing the net synthesis of four-carbon compounds for biosynthesis.
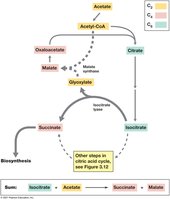
Fermentation
Principles of Fermentation
Fermentation is an anaerobic process in which organic compounds serve as both electron donors and acceptors. ATP is generated by substrate-level phosphorylation, and redox balance is maintained by reducing pyruvate or its derivatives.
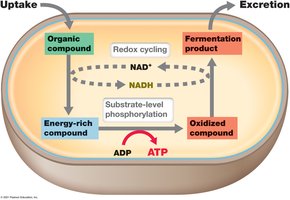
Examples: Ethanol fermentation (yeast), lactic acid fermentation (lactic acid bacteria).
Fermentation products are important in food production and human health.
Respiration: Electron Carriers and Proton Motive Force
Electron Transport Chain Components
Respiration involves the transfer of electrons from NADH and FADH2 to terminal electron acceptors via membrane-bound carriers, generating a proton motive force (pmf).
NADH dehydrogenases: Accept electrons from NADH.
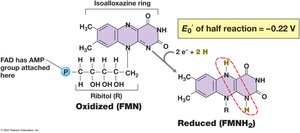
Flavoproteins: Contain FMN or FAD as prosthetic groups.
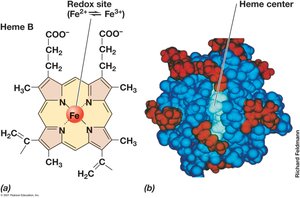
Cytochromes: Proteins with heme groups, transfer electrons via iron atoms.
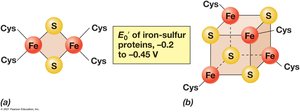
Iron-sulfur proteins: Contain Fe-S clusters, transfer electrons only.
Quinones: Small, hydrophobic, mobile electron carriers.
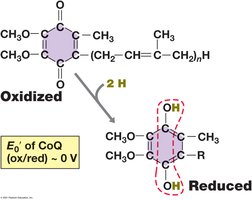
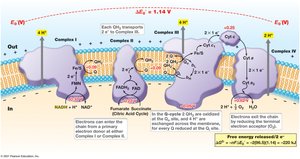
Generation of the Proton Motive Force
Electron transport chains are organized in the cytoplasmic membrane, with electrons flowing through complexes of increasing reduction potential. This flow is coupled to the translocation of protons across the membrane, creating an electrochemical gradient (pmf) used to synthesize ATP.
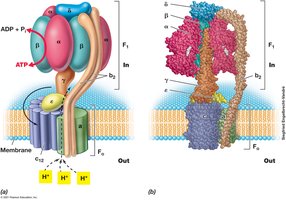
ATP synthase (ATPase) uses the pmf to catalyze ATP formation from ADP and Pi.
In E. coli, about 3.3 H+ are required per ATP synthesized.
Metabolic Diversity: Anaerobic Respiration, Chemolithotrophy, and Phototrophy
Anaerobic Respiration
Some microorganisms can respire in the absence of oxygen by using alternative terminal electron acceptors (e.g., nitrate, sulfate). Anaerobic respiration is less energy-efficient than aerobic respiration.
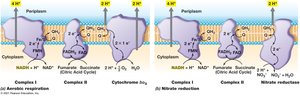
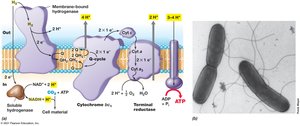
Chemolithotrophy
Chemolithotrophs use inorganic compounds (e.g., H2S, H2, Fe2+, NH4+) as electron donors. They often fix CO2 as a carbon source and use reverse electron transport to generate reducing power.
Phototrophy
Phototrophs use light energy to generate a proton motive force and synthesize ATP via photophosphorylation. Oxygenic phototrophs (e.g., cyanobacteria) produce O2, while anoxygenic phototrophs (e.g., purple bacteria) do not.
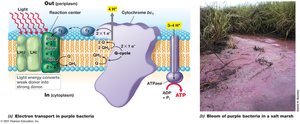
Biosynthesis: Autotrophy, Nitrogen Fixation, and Macromolecule Synthesis
Autotrophy and the Calvin Cycle
Autotrophs fix CO2 into organic molecules using the Calvin cycle, which requires ATP, NADPH, and the enzyme RubisCO. Six CO2 molecules are needed to synthesize one hexose.
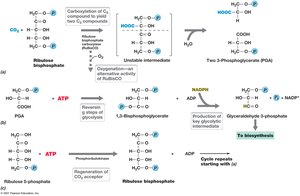
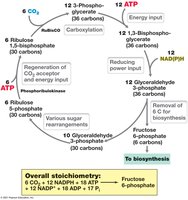
Nitrogen Fixation
Nitrogen fixation is the reduction of atmospheric N2 to ammonia (NH3) by the enzyme nitrogenase. This process is energy-intensive and sensitive to oxygen.
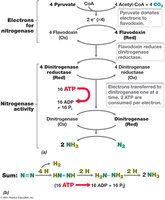
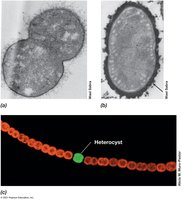
Sugars, Polysaccharides, and Gluconeogenesis
Polysaccharides are synthesized from activated glucose derivatives (e.g., UDP-glucose, ADP-glucose). Gluconeogenesis allows the synthesis of glucose from non-carbohydrate precursors.
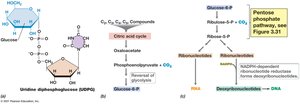
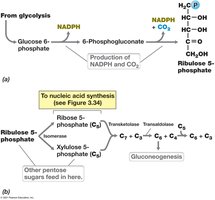
Pentose Phosphate Pathway
This pathway generates pentoses for nucleic acid synthesis and NADPH for biosynthetic reactions.
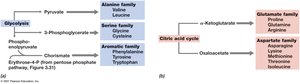
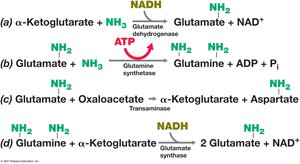
Amino Acid and Nucleotide Biosynthesis
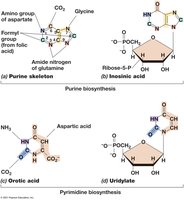
Amino acids are synthesized from intermediates of glycolysis and the citric acid cycle. Nucleotides are built from multiple carbon and nitrogen sources, with purines and pyrimidines following distinct biosynthetic routes.
Fatty Acid and Lipid Biosynthesis
Fatty acids are synthesized two carbons at a time by the action of acyl carrier protein (ACP). Lipids are assembled by attaching fatty acids to glycerol (in Bacteria and Eukarya) or by using isoprenoids (in Archaea).
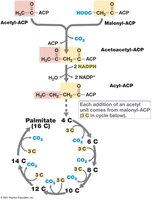
Fatty acid composition varies with temperature and species.
Complex lipids include polar groups for membrane structure.
