 Back
BackMicrobial Metabolism: Principles, Pathways, and Energy Conservation
Study Guide - Smart Notes

Microbial Metabolism
Defining the Requirements for Life
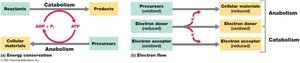
Microbial metabolism encompasses all biochemical reactions necessary for life, including both energy-releasing and energy-synthesizing processes. These reactions are divided into catabolism (breaking down molecules to obtain energy) and anabolism (building cellular material). Metabolic processes rely on the transfer of electrons from electron donors to electron acceptors, and cells conserve energy by converting it into forms that can perform cellular work, primarily through the generation of adenosine triphosphate (ATP).
Catabolism: Energy-releasing metabolic reactions.
Anabolism: Energy-consuming metabolic reactions that synthesize cellular components.
ATP: The universal energy currency in cells, produced by substrate-level and oxidative phosphorylation.
Energy Classes of Microorganisms
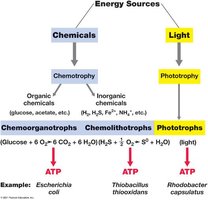
Microorganisms are classified based on their energy and carbon sources. These classifications help in understanding their ecological roles and metabolic diversity.
Chemoorganotrophs: Conserve energy from organic chemicals.
Chemolithotrophs: Oxidize inorganic compounds (e.g., H2, H2S, NH4+).
Phototrophs: Convert light energy into ATP.
Heterotrophs: Obtain carbon from organic compounds.
Autotrophs: Obtain carbon from CO2.
Catalysis and Enzymes
Activation Energy and Catalysis
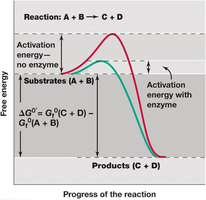
For a chemical reaction to occur, reactant molecules must overcome an energy barrier known as activation energy. Catalysts, including biological catalysts called enzymes, lower this barrier, increasing the rate of reaction without being consumed or altering the reaction's equilibrium.
Activation Energy: Minimum energy required for molecules to become reactive.
Catalyst: Substance that facilitates a reaction, lowers activation energy, and increases reaction rate.
Enzyme Structure and Function
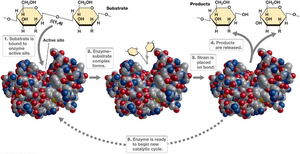
Enzymes are highly specific biological catalysts, typically proteins, that bind substrates at their active site. The shape and chemical environment of the active site are crucial for substrate binding and catalysis. Enzymes can be denatured by high temperatures or extreme pH, which disrupts their structure and function.
Active Site: Region of the enzyme where substrate binding and catalysis occur.
Specificity: Enzymes are highly specific to their substrates.
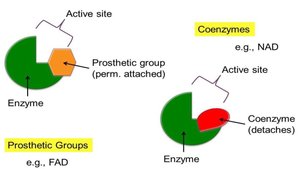
Enzyme Cofactors: Prosthetic Groups and Coenzymes
Many enzymes require non-protein molecules for activity. Prosthetic groups are tightly and permanently bound to the enzyme, while coenzymes are loosely bound and often derived from vitamins.
Prosthetic Groups: Covalently and permanently attached (e.g., heme in cytochromes).
Coenzymes: Loosely attached, often vitamin derivatives (e.g., NAD).
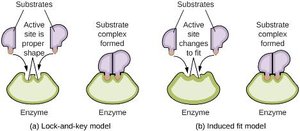
Models of Enzyme-Substrate Interaction
Enzyme catalysis depends on substrate binding and positioning within the active site. Two models describe this interaction: the lock-and-key model and the induced fit model.
Lock-and-Key Model: The active site is a perfect fit for the substrate.
Induced Fit Model: The active site changes shape to fit the substrate upon binding.
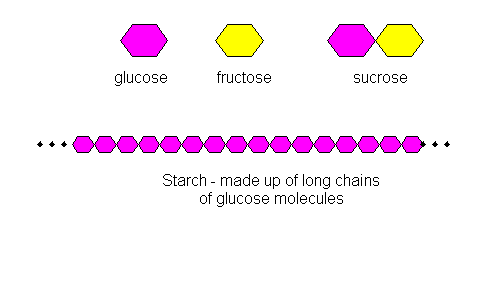
Energy-Rich Compounds and Storage
Long-Term Energy Storage
Microorganisms store energy in the form of insoluble polymers that can be oxidized to generate ATP. These storage compounds differ between prokaryotes and eukaryotes.
Prokaryotes: Glycogen, poly-β-hydroxybutyrate, elemental sulfur.
Eukaryotes: Starch, lipids (simple fats).
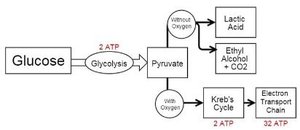
Glycolysis and Fermentation
Overview of Glycolysis and Fermentation
Chemoorganotrophs conserve energy through two main pathways: fermentation and respiration. Fermentation is an anaerobic process where organic compounds serve as both electron donors and acceptors, while respiration involves the oxidation of a donor with O2 (aerobic) or another compound (anaerobic) as the electron acceptor.
Fermentation: Anaerobic catabolism; organic compounds donate and accept electrons.
Respiration: Aerobic or anaerobic catabolism; donor is oxidized with O2 or other acceptors.
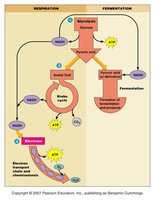
Glycolysis (Embden–Meyerhof–Parnas Pathway)
Glycolysis is a central pathway for glucose catabolism, producing two ATP molecules per glucose via substrate-level phosphorylation. The pathway can lead to fermentation (without O2) or respiration (with O2).
Substrate-Level Phosphorylation: Direct transfer of a phosphate group from a high-energy compound to ADP, forming ATP.
Products: 2 ATP, 2 NADH, and 2 pyruvate per glucose.
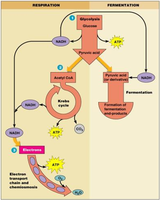
Fermentation Types and Organisms
Fermentation results in various end products depending on the organism and substrate. Common types include alcoholic, homolactic, heterolactic, mixed acid, butyric acid, butanol, caproate/butyrate, and acetogenic fermentations.
Type | Reaction (simplified) | Organisms |
|---|---|---|
Alcoholic | Hexose → 2 ethanol + 2 CO2 | Yeast, Zymomonas |
Homolactic | Hexose → 2 lactate | Streptococcus, some Lactobacillus |
Heterolactic | Hexose → lactate + ethanol + CO2 | Leuconostoc, Oenococcus, some Lactobacillus |
Mixed acid | Hexose → lactate + succinate + ethanol + acetate + formate + CO2 + H2 | Enteric bacteria (Escherichia, Salmonella, Shigella, Klebsiella, Enterobacter) |
Butyric acid | Hexose → butyrate + 2 CO2 + 2 H2 | Clostridium butyricum |
Butanol | Hexose → butanol + acetone + CO2 + H2 | Clostridium acetobutylicum |
Caproate/butyrate | Hexose → caproate + butyrate + CO2 + H2 | Clostridium kluyveri |
Acetogenic | Fructose → 2 acetate + 2 CO2 + 4 H2 | Clostridium aceticum |
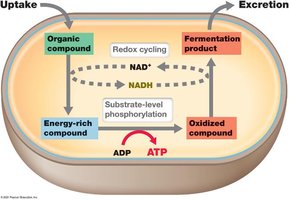
Respiration: Electron Transport Chain (ETC) and Proton Motive Force
Electron Donors and Acceptors
In eukaryotes, NADH is the primary electron donor for the ETC, with molecular oxygen as the final electron acceptor. In prokaryotes, a variety of electron donors and acceptors are used, reflecting metabolic diversity.
NADH: Key electron donor in eukaryotic ETC.
O2: Final electron acceptor in aerobic respiration.
Prokaryotes: Can use nitrate, sulfate, ferric iron, CO2, and organic compounds as electron acceptors.
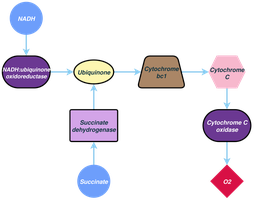
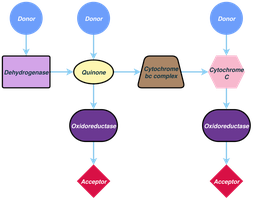
Electron Transport and the Proton Motive Force
The electron transport system is embedded in the cytoplasmic membrane of prokaryotes and the mitochondrial membrane of eukaryotes. As electrons are transferred through the chain, protons are pumped across the membrane, generating a proton motive force (PMF)—an electrochemical gradient used to synthesize ATP.
Proton Motive Force: Results in a pH gradient and electrical potential across the membrane.
ATP Synthesis: Most ATP is generated during electron transport via oxidative phosphorylation.
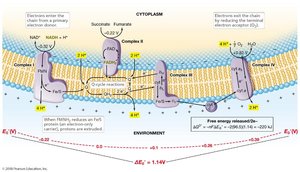
Electron Carriers in the ETC
Electron transport chains involve various carriers, including NADH dehydrogenases, flavoproteins, iron–sulfur proteins, cytochromes, and quinones. Cytochromes contain heme groups and transfer electrons, while quinones are nonprotein carriers that shuttle electrons and protons within the membrane.
Cytochromes: Proteins with heme groups, transfer electrons.
Quinones: Nonprotein molecules, transfer electrons and protons.
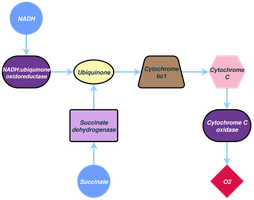
Options for Energy Conservation
Anaerobic Respiration
Some microorganisms use electron acceptors other than oxygen, such as nitrate, ferric iron, sulfate, carbon dioxide, or fumarate. Anaerobic respiration conserves less energy than aerobic respiration but still generates a proton motive force and uses ATPase for ATP synthesis.
Chemolithotrophy
Chemolithotrophs use inorganic chemicals (e.g., H2S, H2, Fe2+, NH4+) as electron donors. These organisms are typically autotrophic, using CO2 as a carbon source, and their metabolism often begins with the oxidation of the inorganic donor.
Phototrophy
Phototrophs use light as an energy source. Photophosphorylation is the light-mediated synthesis of ATP. Photoautotrophs use ATP and CO2 for biosynthesis, while photoheterotrophs use ATP and organic carbon.
Biosynthesis of Cellular Components
Sugars and Polysaccharides
Prokaryotic polysaccharides are synthesized from glucose derivatives such as uridine diphosphoglucose (UDPG). Gluconeogenesis allows synthesis of glucose from non-carbohydrate precursors, and the pentose pathway produces 5-carbon sugars for nucleic acid synthesis.
Amino Acids
Amino acid biosynthesis involves multistep pathways, with carbon skeletons derived from glycolysis or the citric acid cycle. The amine group is incorporated from ammonia and transferred by transaminase enzymes.
Fatty Acids and Lipids
Fatty acid biosynthesis varies with species and temperature. Bacteria and eukarya assemble lipids by adding fatty acids to glycerol, while archaea use phytanyl side chains. All membranes require polar groups for proper architecture.
Low temperature: Shorter, more unsaturated fatty acids.
High temperature: Longer, more saturated fatty acids.
Summary Table: Key Metabolic Pathways
Pathway | Main Function | ATP Yield |
|---|---|---|
Glycolysis | Glucose breakdown to pyruvate | 2 ATP (substrate-level) |
Fermentation | Anaerobic energy production | 2 ATP (substrate-level) |
Aerobic Respiration | Complete oxidation of glucose | ~38 ATP (oxidative phosphorylation) |
Anaerobic Respiration | Oxidation using non-O2 acceptors | Variable, less than aerobic |
Chemolithotrophy | Oxidation of inorganic compounds | Variable |
Phototrophy | Light-driven ATP synthesis | Variable |
Example: Escherichia coli is a chemoorganotroph, Thiobacillus thiooxidans is a chemolithotroph, and Rhodobacter capsulatus is a phototroph.
Additional info: These notes provide a comprehensive overview of microbial metabolism, including energy conservation, metabolic diversity, enzyme function, and biosynthetic pathways, suitable for college-level microbiology students.
