 Back
BackMicrobial Phylogeny, Diversity, and Functional Groups: Study Notes
Study Guide - Smart Notes

Origin and Evolution of Cellular Life
Prebiotic Chemistry and Early Cellular Life
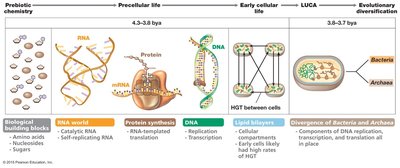
The origin of cellular life is rooted in prebiotic chemistry, where simple biological building blocks such as amino acids, nucleotides, and sugars formed the basis for more complex molecules. The transition from precellular life to early cellular life involved the development of self-replicating RNA, protein synthesis, and eventually DNA-based replication and transcription. The Last Universal Common Ancestor (LUCA) marks the divergence of Bacteria and Archaea, with cellular compartments and horizontal gene transfer (HGT) playing key roles in early evolution.
RNA World: Early life likely relied on catalytic RNA for replication and self-organization.
Protein Synthesis: RNA-templated translation enabled the production of proteins.
DNA: Provided stable genetic information, allowing replication and transcription.
LUCA: The population of early cells from which modern Bacteria and Archaea diverged.
Evolutionary Diversification: Comparison of DNA replication, transcription, and translation mechanisms led to the separation of Bacteria and Archaea.
Photosynthesis, Oxygen, and Ozone in Microbial Diversity
Impact of Photosynthesis and Oxygen Production
Photosynthesis, particularly by Cyanobacteria, was a major driver of microbial diversity. Oxygenic photosynthesis led to the accumulation of O2 in the atmosphere, which in turn enabled the formation of the ozone layer. This shielded terrestrial life from harmful UV radiation and allowed organisms to colonize new habitats.
Phototrophs: Use light energy to oxidize molecules and synthesize organic compounds from CO2.
Cyanobacteria: Earliest oxygen-producing organisms; oxygenic phototrophs.
Ozone Layer: Formed from O2, absorbs UV radiation, protecting DNA and enabling terrestrial colonization.
Microbial Diversification and Endosymbiosis
Evolution of Eukaryotic Cells
The development of an oxic atmosphere spurred the evolution of new metabolic pathways and organelle-containing eukaryotic microorganisms. The endosymbiosis hypothesis posits that mitochondria and chloroplasts originated from symbiotic associations of prokaryotes within other cells.
Endosymbiosis: Mitochondria and chloroplasts arose from prokaryotes living inside another cell.
Metabolic Pathways: Oxic conditions enabled more energy-efficient metabolic processes.
Microbial Evolutionary Processes
Mechanisms of Microbial Evolution
Microbial evolution is driven by changes in allele frequencies within populations, resulting from mutation, recombination, natural selection, and genetic drift. These processes contribute to genetic diversity and adaptation.
Mutation: Random changes in DNA sequence; can be neutral, deleterious, or beneficial.
Recombination: DNA segments are broken and rejoined, creating new genetic combinations.
Selection: Fitness-based process favoring beneficial mutations.
Genetic Drift: Random changes in gene frequencies, especially in small populations.
Phylogenetic Analysis and 16S rRNA
Phylogeny and Molecular Clocks
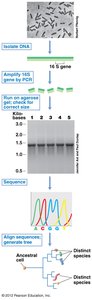
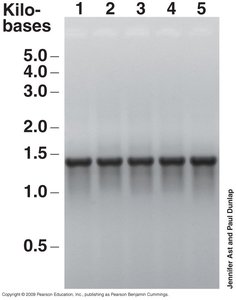
Phylogeny refers to the evolutionary history of organisms, inferred from nucleotide sequence data. Molecular clocks, such as small-subunit ribosomal RNA (SSU rRNA) genes, are used to measure evolutionary change. The 16S rRNA gene is the gold standard for phylogenetic comparisons in prokaryotes due to its universality, functional constancy, and slow rate of change.
16S rRNA: Found in all prokaryotes, mitochondria, and chloroplasts; highly conserved.
18S rRNA: Used for eukaryotes.
Limitations: Poor resolution for closely related species; multi-gene approaches improve discrimination.
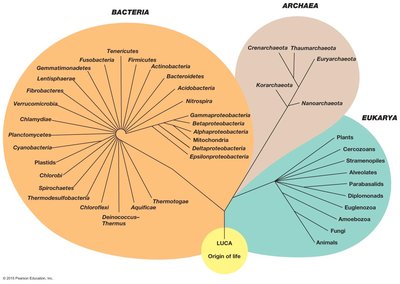
Universal Phylogenetic Tree
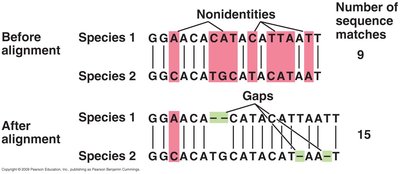
The universal phylogenetic tree, based on SSU rRNA genes, illustrates the genealogy of all life, showing the relationships among Bacteria, Archaea, and Eukarya. Branches represent evolutionary lineages, and branch lengths indicate the number of genetic changes.
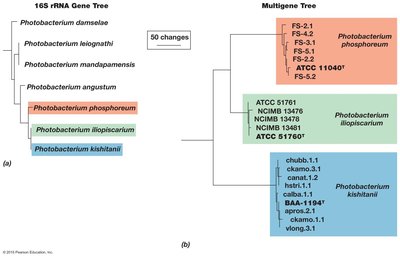
Multigene Phylogenetic Analysis
Multi-gene approaches, such as combining 16S rRNA with other genes (e.g., gyrB, luxABFE), provide clearer resolution for distinguishing species and genera.
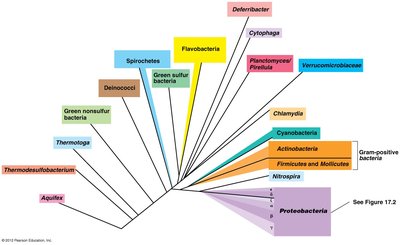
Microbial Diversity: Phylogenetic and Functional Perspectives
Phylogenetic Diversity
Phylogenetic diversity refers to the evolutionary relationships among organisms, defined by ribosomal RNA phylogeny. It encompasses diversity at the phylum, genus, and species levels, as well as genetic and genomic diversity.
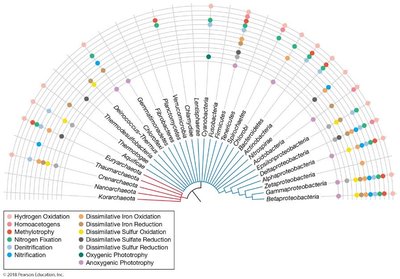
Functional Diversity
Functional diversity relates to microbial physiology and ecology, including metabolic, ecological, and morphological traits. Functional groups are useful for understanding microbial roles in ecosystems, gene loss, convergent evolution, and horizontal gene transfer.
Phototrophic Bacteria and Cyanobacteria
Overview of Phototrophic Bacteria
Phototrophic bacteria conserve energy from light, using chlorophyll-like pigments and membrane-bound reaction centers. Anoxygenic phototrophs do not generate O2, while oxygenic phototrophs (Cyanobacteria) do. Pigments are often found in intracellular membrane systems, and many phototrophs fix carbon.
Cyanobacteria: Structure, Function, and Ecology
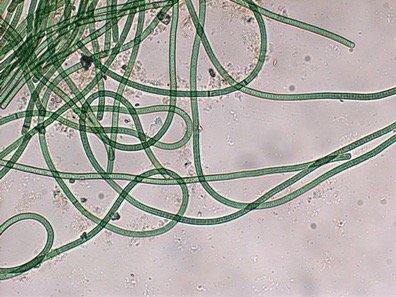
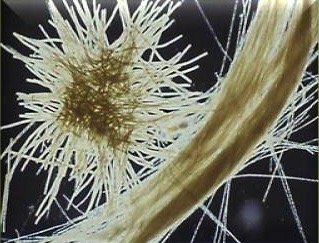
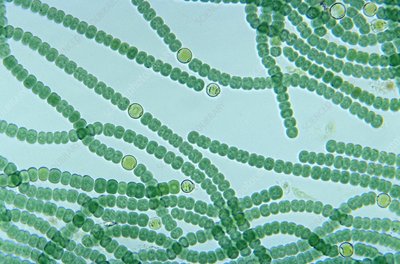
Cyanobacteria are a diverse group of oxygenic phototrophic bacteria, ranging from unicellular to filamentous forms. They possess specialized membrane systems called thylakoids for photosynthesis, produce pigments (chlorophyll a and phycobilins), and can fix CO2 and N2. Cyanobacteria are important for marine and terrestrial productivity, nitrogen fixation, and can form symbiotic relationships (e.g., lichens).
Key Genera: Prochlorococcus, Crocosphaera, Synechococcus, Trichodesmium, Oscillatoria, Anabaena
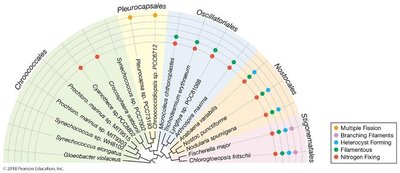
Morphological Groups: Chroococcales (unicellular), Pleurocapsales (colonial), Oscillatoriales (filamentous), Nostocales (filamentous, can differentiate), Stigonematales (branching filaments)
Physiology: Oxygenic photosynthesis, nitrogen fixation, vitamin synthesis, photoheterotrophy, anoxygenic photosynthesis (some species)
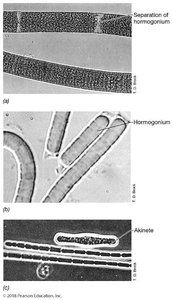
Motility: Gliding motility, phototaxis, gas vesicles for buoyancy, sheaths, hormogonia, akinetes, cyanophycin storage
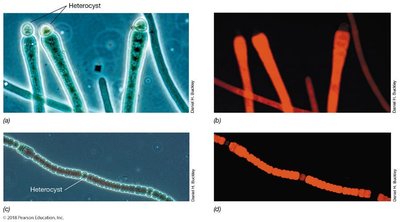
Heterocysts: Specialized cells for nitrogen fixation, provide an anoxic environment, exchange materials with adjacent cells
Ecology: Dominant marine phototrophs, tolerant of extreme environments, important for nitrogen input and metabolic products
Major Lineages and Functional Groups of Bacteria
Proteobacteria and Functional Groups
Proteobacteria represent the most diverse and largest phylum of Bacteria, encompassing a wide range of metabolic and morphological diversity. Functional groups include phototrophic, nitrifying, sulfur-oxidizing, hydrogen-oxidizing, and microbial predators.
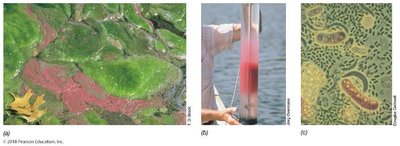
Purple Phototrophic Bacteria: Anoxygenic photosynthesis, found in anoxic zones with H2S.
Nitrifying Bacteria: Chemolithotrophic, oxidize ammonia to nitrate, important in soil and wastewater treatment.
Sulfur-Oxidizing Bacteria: Chemolithotrophic, use reduced sulfur compounds, found in sulfur-rich habitats.
Hydrogen-Oxidizing Bacteria: Use H2 as electron donor, aerobic, autotrophic growth.
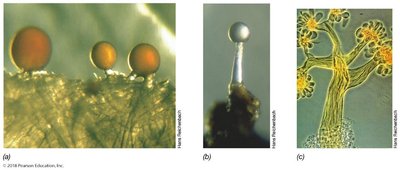
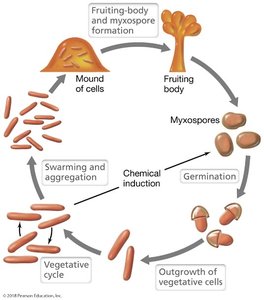
Microbial Predators (Myxobacteria): Complex life cycle, formation of fruiting bodies, predatory behavior.



Gram-Negative and Gram-Positive Bacteria
Gram-Negative Bacteria
Gram-negative bacteria include diverse groups such as Pseudomonads, Acetic Acid Bacteria, Neisseria, Enterobacteriaceae, and Vibrio. They exhibit a range of metabolic strategies, pathogenicity, and ecological roles.
Pseudomonas: Aerobic, motile, oxidase positive, cannot ferment, versatile in organic compound degradation.
Acetic Acid Bacteria: Incomplete oxidation of alcohols, production of acetic acid, used in vinegar production.
Neisseria: Gram-negative cocci, some pathogenic (e.g., N. gonorrhoeae).
Enterobacteriaceae: Facultative, fermentative, motile/non-motile rods, includes E. coli, Salmonella, Shigella, Proteus.
Vibrio: Motile, oxidase positive, some pathogenic (e.g., V. cholerae).
Gram-Positive Bacteria
Gram-positive bacteria include Firmicutes and Actinobacteria, with nonsporulating, endospore-forming, and cell-wall-less groups. They play important roles in fermentation, antibiotic production, and pathogenicity.
Nonsporulating: Staphylococcus, Streptococcus, Lactobacillus, Listeria; lactic acid bacteria, food production, pathogenic species.
Endospore-Forming: Bacillus, Clostridium; soil microorganisms, antibiotic production, pathogenic species (anthrax, botulism, tetanus).
Cell-Wall-Less: Mycoplasma, Spiroplasma; small genomes, parasitic, lack peptidoglycan.
Actinobacteria: Corynebacterium, Mycobacterium, Propionibacterium, Streptomyces; antibiotic production, acid-fastness, filamentous growth.
Spirochetes and Chlamydia
Spirochetes
Spirochetes are Gram-negative, motile, tightly coiled bacteria with endoflagella. They are found in aquatic environments and animals, with several pathogenic genera (e.g., Treponema, Borrelia, Leptospira).
Treponema: Causes syphilis.
Borrelia: Causes Lyme disease.
Leptospira: Causes leptospirosis, transmitted via rodents.
Chlamydia
Chlamydia are obligate intracellular parasites with limited metabolic capacities. They cause diseases such as trachoma and chlamydia STD.
Summary Table: Functional and Taxonomic Groups
Functional Group | Taxonomic Group | Representative Organisms | Key Features |
|---|---|---|---|
Oxygenic Phototrophs | Cyanobacteria | Prochlorococcus, Synechococcus, Anabaena | Oxygenic photosynthesis, nitrogen fixation, thylakoids |
Anoxygenic Phototrophs | Purple, Green Bacteria | Rhodospirillum, Chlorobium | Anoxygenic photosynthesis, diverse pigments |
Nitrifiers | Proteobacteria | Nitrosococcus, Nitrobacter | Ammonia/nitrite oxidation, chemolithotrophy |
Sulfur Oxidizers | Proteobacteria | Thiobacillus, Beggiatoa | Sulfur compound oxidation, acid tolerance |
Microbial Predators | Myxobacteria | Myxococcus | Fruiting bodies, predatory behavior |
Lactic Acid Bacteria | Firmicutes | Streptococcus, Lactobacillus | Lactic acid fermentation, food production |
Endospore Formers | Firmicutes | Bacillus, Clostridium | Endospore formation, soil, pathogenic species |
Actinobacteria | Actinobacteria | Streptomyces, Mycobacterium | Antibiotic production, acid-fastness, filamentous growth |
Spirochetes | Spirochaetes | Treponema, Borrelia, Leptospira | Motility, pathogenicity, endoflagella |
Chlamydia | Chlamydiae | Chlamydia trachomatis | Obligate intracellular, limited metabolism |
