 Back
BackMicrobial Regulatory Systems, Biofilm Formation, and Genetics of Bacteria
Study Guide - Smart Notes

Cell-to-Cell Signaling and Quorum Sensing
Quorum Sensing Mechanism
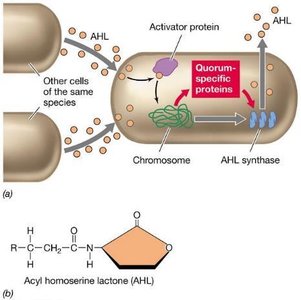
Quorum sensing is a regulatory mechanism by which bacteria and some archaea assess their population density and coordinate group behaviors. This process relies on the production and accumulation of small extracellular molecules called autoinducers, which trigger the transcription of specific genes when a threshold concentration is reached.
Autoinducers: Small peptides or nonpeptide organics produced by each species; include acyl homoserine lactone (AHL), autoinducer 2 (AI-2), and short peptides.
Mechanism: Autoinducers diffuse freely across the cell envelope. When many cells are nearby, the concentration rises, binding to activator proteins or sensor kinases and triggering gene transcription.
Coordinated Behaviors: Includes biofilm formation, toxin production, and bioluminescence.
Example: Aliivibrio fischeri uses quorum sensing to regulate bioluminescence via the Lux operon.
Quorum Sensing and Virulence
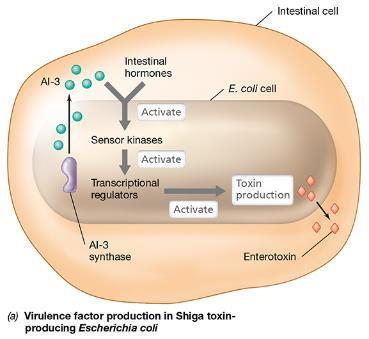
Quorum sensing is critical for the regulation of virulence factors in pathogenic bacteria. For example, Escherichia coli O157:H7 produces AHL AI-3, which, together with host hormones, activates sensor kinases and transcriptional regulators, leading to toxin production and pathogenicity.
Virulence Regulation: Only when a sufficient number of cells are present do virulence genes activate, making bacteria effective pathogens.
Drug Design: Antagonists can block bacterial communication, offering potential therapeutic strategies.
Biofilm Formation
Stages of Biofilm Development
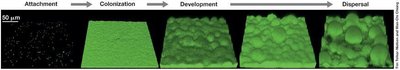
Biofilms are structured communities of microbial cells enmeshed in a polysaccharide matrix and attached to surfaces. Biofilm formation occurs in distinct stages:
Attachment: Planktonic cells attach to surfaces using flagella, fimbriae, or pili.
Colonization: Cells grow and produce extracellular polysaccharides (EPS).
Development: Metabolic changes occur, and microcolonies form.
Dispersal: Cells leave the biofilm to colonize new sites.

Examples of Biofilms
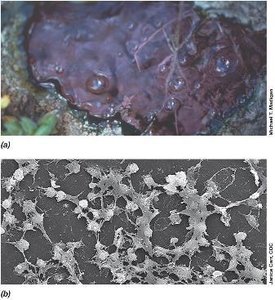
Biofilms are found in diverse environments, from natural settings to medical devices. For instance, microbial mats in hot springs and biofilms on catheters are common examples.
Medical Relevance: Biofilms on indwelling devices can cause persistent infections.
Environmental Relevance: Microbial mats contribute to nutrient cycling in ecosystems.

Cyclic Di-GMP Regulation of Biofilm Formation
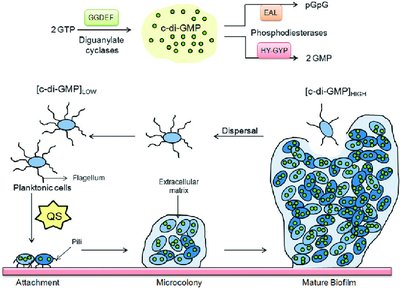
Cyclic di-guanosine monophosphate (c-di-GMP) is a regulatory molecule that guides bacteria in transitioning from planktonic to biofilm growth. Elevated c-di-GMP levels trigger physiological events such as reduced flagellar activity, increased attachment protein expression, and extracellular matrix biosynthesis.
Distribution: c-di-GMP is widely distributed in bacteria.
Regulation: Synthesis and degradation depend on environmental and cellular cues.
Genetics of Bacteria and Archaea
Mutation and Genetic Diversity
Mutations are heritable changes in the genome that can alter the properties of an organism. Bacteria accumulate mutations rapidly due to exponential growth, and horizontal gene transfer generates larger genetic changes, fueling evolution.
Types of Mutations: Base-pair substitutions (missense, nonsense, silent), frameshifts, insertions, and deletions.
Horizontal Gene Transfer: Includes transformation, transduction, and conjugation.
Molecular Basis of Mutation
Base-pair substitutions can result in missense, nonsense, or silent mutations, affecting protein function to varying degrees. Silent mutations often occur at the third base of a codon and do not alter the encoded polypeptide.
Missense Mutation: Changes amino acid sequence; may affect protein function.
Nonsense Mutation: Introduces a stop codon, resulting in truncated proteins.
Silent Mutation: No effect on protein sequence or phenotype.
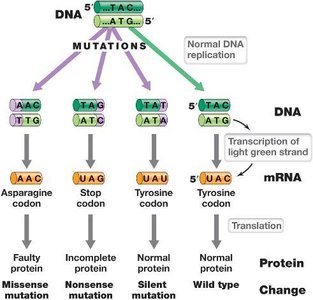
Frameshift Mutations
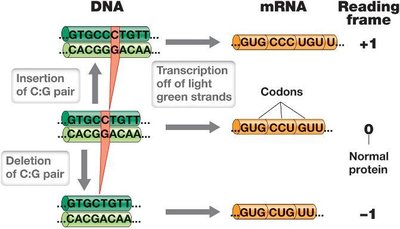
Frameshift mutations result from insertions or deletions of base pairs, causing a shift in the reading frame and scrambling the downstream polypeptide sequence. These mutations often lead to loss of gene function and can be lethal.
Insertion/Deletion: Addition or removal of base pairs alters the reading frame.
Impact: Usually results in nonfunctional proteins.
Mutagenesis and DNA Repair
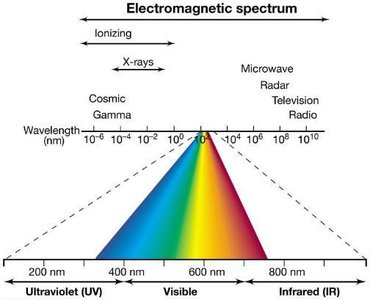
Mutagenesis can be induced by radiation, such as ultraviolet (UV) or ionizing radiation. UV primarily forms pyrimidine dimers, while ionizing radiation generates free radicals that damage DNA. Bacteria possess repair systems, including the SOS response, to correct DNA damage.
Nonionizing Radiation: UV causes pyrimidine dimers.
Ionizing Radiation: X-rays, gamma rays cause breaks and rearrangements.
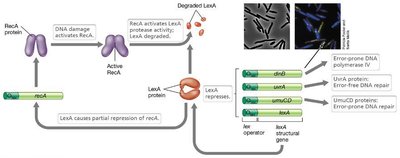
SOS System: Activated by DNA damage; involves LexA and RecA proteins.
Gene Transfer in Bacteria
Mechanisms of Horizontal Gene Transfer
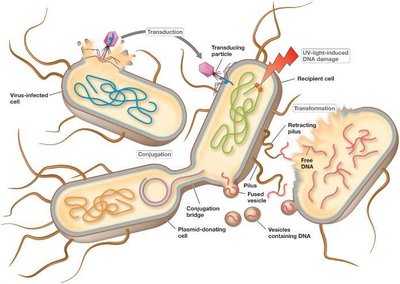
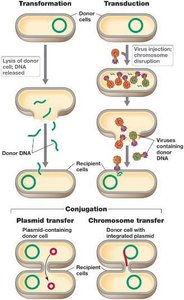
Bacteria can exchange genes through three main mechanisms: transformation, transduction, and conjugation. These processes allow rapid acquisition of new traits and contribute to genetic diversity.
Transformation: Uptake of free DNA from the environment.
Transduction: Transfer of DNA via bacteriophages.
Conjugation: Direct cell-to-cell transfer of plasmids or chromosomal DNA.
Homologous Recombination
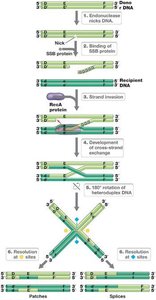
Homologous recombination is the physical exchange of DNA between genetic elements with similar sequences. The RecA protein is essential for this process, which can result in new genotypes and phenotypes.
Mechanism: Involves strand invasion, formation of heteroduplex regions, and resolution into patches or splices.
Detection: Recombinant cells are identified by selectable characteristics.
Transformation
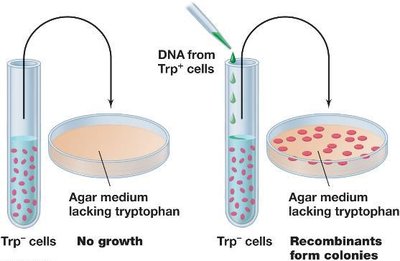
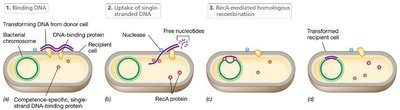
Transformation involves the uptake and integration of DNA from the environment. Competent cells bind and internalize DNA, which is then integrated into the chromosome by RecA-mediated recombination.
Natural Transformation: Begins with reversible DNA binding, followed by irreversible uptake.
Competence: Cells capable of transformation bind much more DNA than noncompetent cells.
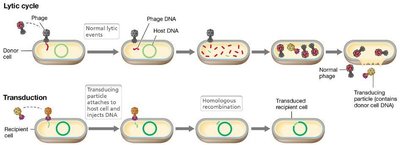
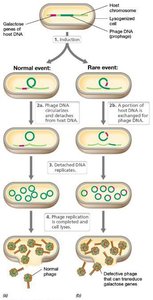
Transduction
Transduction is the transfer of DNA from one cell to another by a bacteriophage. It occurs in two modes: generalized and specialized transduction.
Generalized Transduction: Any gene can be transferred; host DNA is accidentally packaged into phage particles.
Specialized Transduction: Only specific genes adjacent to the phage integration site are transferred.
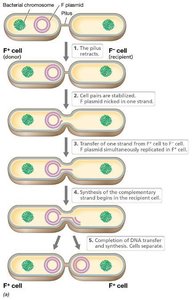
Conjugation
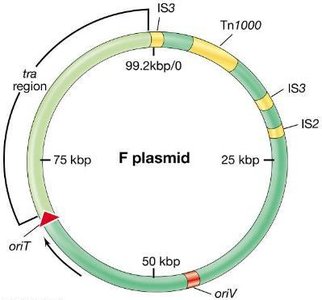
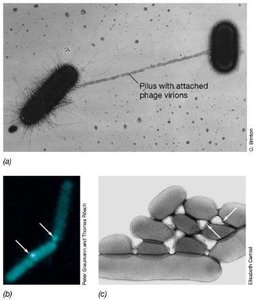
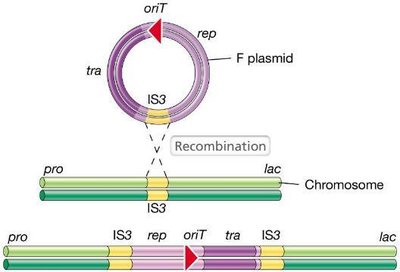
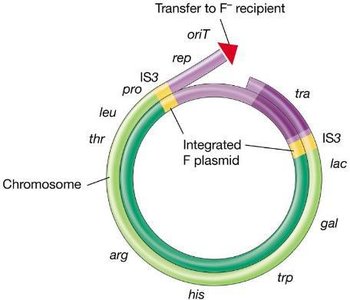
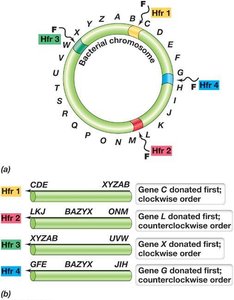
Conjugation is a plasmid-encoded process requiring cell-to-cell contact. The F (fertility) plasmid in E. coli encodes genes for pilus formation and DNA transfer. Conjugation can mobilize chromosomal genes, especially when the F plasmid integrates into the chromosome, forming Hfr strains.
F Plasmid: Contains replication, transfer, and integration genes.
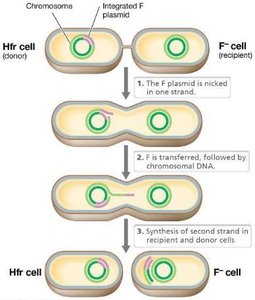
Hfr Strains: High frequency recombination strains transfer chromosomal genes to recipients.
Mobile DNA and CRISPR Systems
Transposable Elements
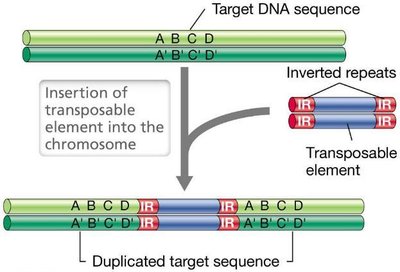
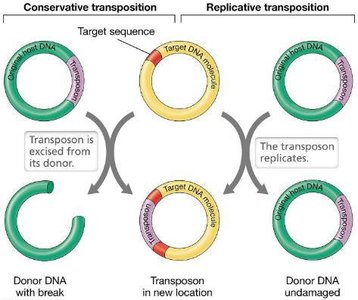
Transposable elements are discrete DNA segments that move within genomes. They include insertion sequences (ISs) and transposons, which encode transposase and have inverted repeats at their ends. Transposition can occur via conservative or replicative mechanisms.
Conservative Transposition: Transposon excised and reinserted elsewhere; copy number remains constant.
Replicative Transposition: New copy produced and inserted; copy number doubles.
Transposon Mutagenesis
Transposon mutagenesis is a powerful tool for creating mutants. Transposons carrying antibiotic resistance genes are introduced into cells, and mutants are selected for loss of function or altered phenotypes.
Screening: Mutants are screened for specific traits, such as biofilm formation.
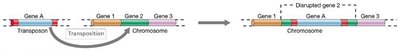
CRISPR Adaptive Immunity
CRISPR (Clustered Regulatory Interspaced Short Palindromic Repeats) is a major antiviral defense system in bacteria and archaea. It consists of short repeats and spacers derived from previous invaders, providing genetic memory and specificity analogous to adaptive immunity.
Cas Proteins: Mediate defense and incorporate new spacers into CRISPR regions.
Immunization: Cas proteins recognize and excise protospacer regions from viral DNA, inserting them as spacers.
Interference: crRNAs guide Cas proteins to complementary viral DNA, triggering degradation.
Distribution: CRISPR systems are found in ~90% of archaea and 70% of bacteria. Viruses may evade CRISPR by mutating PAM regions or producing Cas inhibitors.
