 Back
BackMicrobial Regulatory Systems: DNA-Binding Proteins and Transcriptional Control
Study Guide - Smart Notes

Microbial Regulatory Systems
Overview of Regulation in Microbial Cells
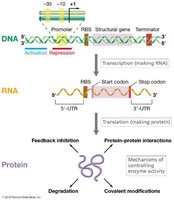
Microbial cells must precisely regulate the types, amounts, and activities of proteins and other macromolecules to orchestrate their diverse biochemical reactions. Regulation of gene expression is essential for conserving energy and resources, ensuring that proteins are produced only when needed.
Constitutive expression: Some proteins and RNA molecules are needed at constant levels under all growth conditions.
Regulated expression: Other proteins are produced only under specific circumstances (e.g., enzymes for lactose catabolism are synthesized only when lactose is present).
Regulation can occur at multiple levels:
Transcriptional regulation: Controls the amount of mRNA produced.
Translational regulation: Controls how much protein is translated from mRNA.
Post-translational regulation: Controls protein activity or stability (e.g., phosphorylation, degradation).
DNA-Binding Proteins
Protein-DNA Interactions
Small molecules influence the binding of regulatory proteins to DNA, thereby turning transcription on or off at specific promoters. Most regulatory proteins interact with DNA in a sequence-specific manner, primarily at the major groove, where amino acid side chains recognize specific base pairs.
Sequence-specific binding: Provided by interactions between protein side chains and DNA bases.
Homodimeric proteins: Many DNA-binding proteins are composed of two identical polypeptides, each binding to an inverted repeat sequence.
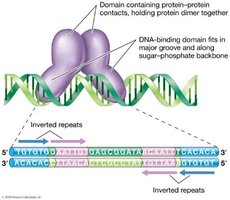
Common DNA-Binding Domains
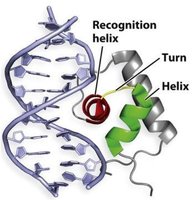
Helix-turn-helix: Two α-helices connected by a short turn; the recognition helix interacts with DNA, while the stabilizing helix supports the structure. Found in many bacterial DNA-binding proteins.
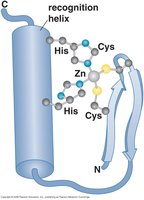
Zinc finger: Found in eukaryotic DNA-binding proteins; a zinc ion stabilizes the domain by interacting with cysteine and histidine residues. Multiple zinc fingers increase binding strength.
Leucine zipper: Contains regularly spaced leucine residues that hold two recognition helices together, forming pincers that grip the DNA.
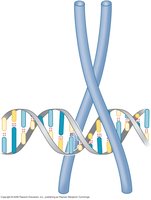
Roles of DNA-Binding Proteins
Catalysis: DNA-binding proteins may catalyze reactions such as transcription (RNA polymerase) or replication (DNA polymerase).
Regulation: DNA binding can block transcription (negative regulation) or activate transcription (positive regulation).
Negative Control: Repression and Induction
Repression
Negative control prevents transcription. Repression mechanisms inhibit gene expression in response to a signal, often the end product of a biosynthetic pathway. This prevents unnecessary synthesis of proteins.
Operon: Cluster of genes transcribed as a polycistronic mRNA under a single promoter.
Operator: Repressor binding site within the operon.
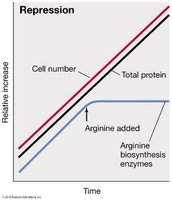
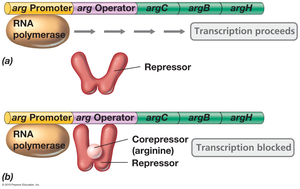
Example: Arginine operon—In the absence of arginine, the repressor cannot bind the operator, and transcription proceeds. When arginine is present, it acts as a corepressor, enabling the repressor to bind the operator and block transcription.
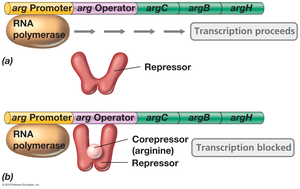
Induction
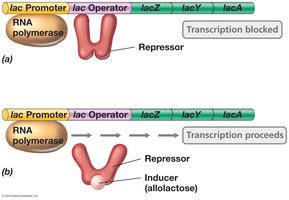
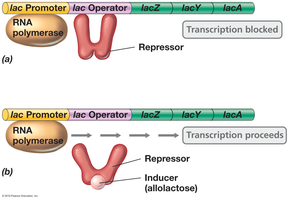
Induction upregulates gene expression in response to a signal, typically affecting catabolic enzymes. The repressor protein prevents transcription unless an inducer is present.
Inducer: Substance that binds the repressor, causing it to release the operator and allowing transcription.
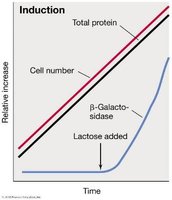
Example: Lac operon—In the absence of lactose, the repressor binds the operator and blocks transcription. When lactose (or its analog allolactose) is present, it binds the repressor, causing it to release the operator and allowing transcription.
Transcriptional Control and Allosteric Regulation
Effectors and Allostery
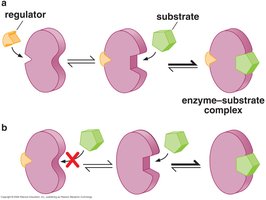
Effectors (inducers and corepressors) are small molecules that regulate DNA-binding proteins by binding and changing their conformation. This is known as allosteric regulation.
Allostery: Regulation of protein function by changing its shape.
Positive allostery: Effector binding enables protein activity.
Negative allostery: Effector binding inhibits protein activity.
Negative Regulation via Allostery
Repressor proteins can be regulated allosterically to control gene expression. Both positive and negative allostery are used in negative regulation, resulting in either repression or induction.
Positive Control: Activation
Mechanism of Positive Control
Positive control uses activator proteins to facilitate RNA polymerase binding to the promoter, often regulated allosterically. Activator proteins may bend DNA or interact directly with RNA polymerase.
Activator: Regulatory protein that activates transcription.
Example: Maltose activator protein in E. coli—Maltose must bind the activator protein for it to bind DNA and help RNA polymerase initiate transcription.
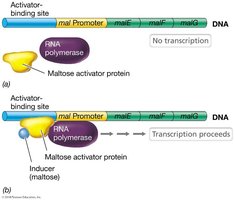
Promoters of positively controlled operons are often weak matches to the consensus sequence, so activator proteins are necessary for efficient transcription.
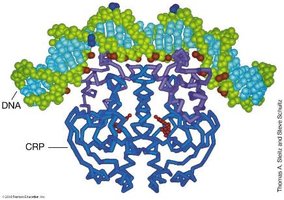
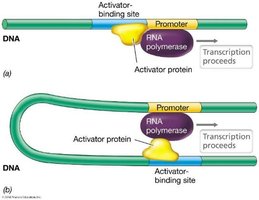
Activator proteins can interact with RNA polymerase either near the promoter (proximal) or at a distance (distal), requiring DNA looping.
Operons vs. Regulons
Definitions and Examples
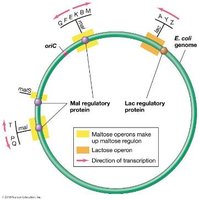
An operon is a cluster of genes under the control of a single promoter. A regulon consists of multiple operons controlled by the same regulatory protein, which can be either an activator or a repressor.
Example: Maltose regulon—Multiple operons for maltose utilization are controlled by the maltose activator protein.
Negative regulon: Arginine regulon—All arginine operons are controlled by the arginine repressor protein.
Transcription Controls in Archaea
Regulatory Mechanisms in Archaea
Transcription regulation in Archaea resembles bacterial mechanisms, using both repressors and activators. Archaeal repressors can block RNA polymerase or prevent transcription factors (TBP and TFB) from assisting RNA polymerase binding. Activator proteins recruit TFB to the promoter, facilitating RNA polymerase recruitment.
Repressors: Block RNA polymerase or transcription factors from binding the promoter.
Activators: Recruit transcription factors to the promoter.
Additional info: In Eukaryotes, transcription regulation is more complex, involving numerous transcription factors and regulatory elements.
