 Back
BackMicrobial Regulatory Systems: Mechanisms of Transcriptional Control in Bacteria and Archaea
Study Guide - Smart Notes

Regulation of Transcription in Microorganisms
Overview of Transcriptional Regulation
Transcriptional regulation is a fundamental process in bacteria and archaea, allowing cells to adapt gene expression in response to environmental and metabolic signals. While the core mechanisms are similar in both domains, eukaryotes possess additional regulatory layers. Key regulatory strategies include the use of repressor and activator proteins, two-component regulatory systems, anti-sigma factor interactions, and multicomponent phosphorelay systems.
Negative control: Prevents transcription, often through enzyme repression or induction.
Positive control: Activates transcription, typically via activator proteins and inducers.
Negative Control of Transcription
Enzyme Repression: The arg Operon
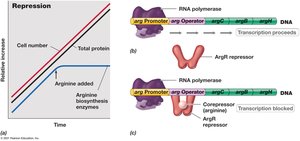
Enzyme repression occurs when the end product of a biosynthetic pathway is abundant, leading to the inhibition of enzymes involved in its synthesis. The arg operon in Escherichia coli is a classic example, regulated by the availability of arginine.
Mechanism: In the absence of arginine, the repressor protein cannot bind to the operator, allowing transcription of genes for arginine biosynthesis.
When arginine is present, it acts as a corepressor, binding to the repressor protein. This complex binds to the operator, blocking RNA polymerase and halting transcription.
Result: Synthesis of arginine biosynthetic enzymes is repressed when arginine is abundant.
Enzyme Induction: The lac Operon
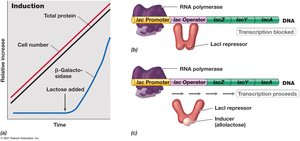
Enzyme induction is the process by which the presence of a substrate induces the synthesis of enzymes required for its metabolism. The lac operon is a well-studied example, induced by lactose.
Mechanism: In the absence of lactose, the LacI repressor binds to the operator, blocking transcription of the lac genes.
When lactose (or its isomer allolactose) is present, it binds to the repressor, causing it to release from the operator. This allows RNA polymerase to transcribe the genes necessary for lactose metabolism.
Result: β-galactosidase and other enzymes are produced only when lactose is available.
Positive Control of Transcription
Activation of the mal Operon
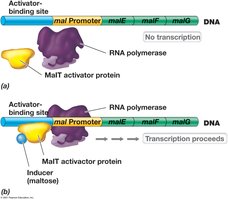
Positive control involves activator proteins that enhance transcription when bound to specific DNA sites. The mal operon, responsible for maltose utilization, is regulated by the MalT activator protein.
Mechanism: In the absence of maltose, the MalT activator cannot bind to the activator-binding site, and transcription does not occur.
When maltose is present, it binds to MalT, enabling it to bind DNA and recruit RNA polymerase, thus activating transcription of maltose utilization genes.
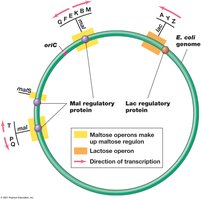
Regulons
A regulon is a group of operons or genes regulated by a single regulatory protein, allowing coordinated control of multiple pathways. For example, the maltose regulon in E. coli includes several operons under the control of MalT.
Catabolite Repression and the Glucose Effect
Catabolite Repression
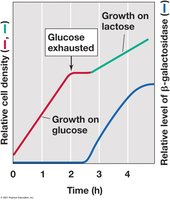
Catabolite repression is a global regulatory mechanism that allows bacteria to preferentially use certain carbon sources (like glucose) over others. This is often referred to as the "glucose effect." When glucose is present, the synthesis of enzymes for the metabolism of alternative sugars (e.g., lactose) is repressed.
Mechanism: Glucose inhibits the synthesis of cyclic AMP (cAMP). Low cAMP levels prevent the cAMP receptor protein (CRP) from binding to DNA, which is necessary for the activation of the lac operon.
Only when glucose is depleted does cAMP accumulate, allowing CRP to bind DNA and activate transcription of alternative sugar operons.
Two-Component Regulatory Systems
Mechanism and Examples
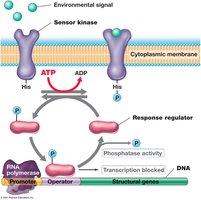
Two-component systems are widespread in bacteria and archaea, enabling cells to sense and respond to environmental changes. These systems typically consist of a sensor kinase and a response regulator.
Sensor kinase: Located in the cytoplasmic membrane, it detects environmental signals and autophosphorylates a conserved histidine residue.
Response regulator: Receives the phosphate group from the sensor kinase and mediates changes in gene expression, often by acting as a transcriptional activator or repressor.
Global Regulatory Mechanisms
The Stringent Response
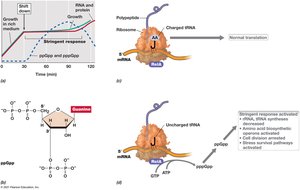
The stringent response is a global regulatory mechanism that allows bacteria to adapt to amino acid starvation and other stresses. It is mediated by alarmones such as ppGpp, which inhibit rRNA and tRNA synthesis, slowing cell growth and redirecting resources.
Trigger: Amino acid limitation leads to accumulation of uncharged tRNAs, activating RelA to synthesize ppGpp.
Effect: ppGpp binds to RNA polymerase, reducing transcription of rRNA and tRNA genes, and altering expression of numerous other genes.
Attenuation: Regulation of the trp Operon
Mechanism of Attenuation
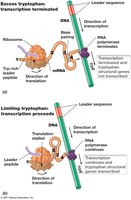
Attenuation is a regulatory mechanism that fine-tunes gene expression in response to amino acid availability. In the trp operon, attenuation controls the completion of transcription based on tryptophan levels.
Leader peptide: The trpL leader sequence encodes a short peptide with multiple tryptophan codons.
High tryptophan: Ribosomes quickly translate the leader peptide, allowing formation of a transcription terminator structure in the mRNA, halting transcription.
Low tryptophan: Ribosomes stall at the tryptophan codons, preventing terminator formation and allowing transcription of the full operon.
Summary Table: Key Regulatory Mechanisms
Mechanism | Key Feature | Example |
|---|---|---|
Negative Control | Repressor blocks transcription | arg operon, lac operon |
Positive Control | Activator enhances transcription | mal operon |
Catabolite Repression | Global repression by preferred carbon source | lac operon (glucose effect) |
Two-Component System | Sensor kinase and response regulator | Nitrogen assimilation, chemotaxis |
Attenuation | Transcription termination based on metabolite levels | trp operon |
