 Back
BackChpt 8 Study guide Microbiology Study Guide: Recombinant DNA Technology, PCR, and Related Concepts
Study Guide - Smart Notes

Q1. Thoroughly explain how real time PCR works and how it can be used to identify the presence of covid 19 virus in a nasal sample.
Background
Topic: Real-Time PCR (qPCR) in Viral Detection
This question tests your understanding of how real-time PCR is used to detect and quantify viral genetic material, specifically for diagnosing COVID-19.
Key Terms and Concepts:
PCR (Polymerase Chain Reaction): A technique to amplify specific DNA sequences.
Real-Time PCR (qPCR): PCR with fluorescent markers to monitor DNA amplification as it happens.
Denaturation, Annealing, Extension: The three main steps in each PCR cycle.
Fluorescent Dye: Binds to double-stranded DNA, allowing quantification.
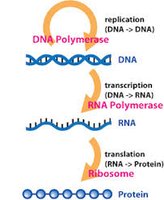
Step-by-Step Guidance
Start by extracting RNA from the nasal swab sample, since SARS-CoV-2 is an RNA virus.
Use reverse transcriptase to convert viral RNA into complementary DNA (cDNA).
Set up the PCR reaction with primers specific to SARS-CoV-2 genes, DNA polymerase, nucleotides, and fluorescent dye.
Run the PCR cycles:
Denaturation: Heat to separate DNA strands.
Annealing: Cool so primers bind to target sequences.
Extension: DNA polymerase synthesizes new DNA strands.
During each cycle, the fluorescent dye binds to double-stranded DNA. The increase in fluorescence is measured in real time, indicating the amount of DNA present.
Try solving on your own before revealing the answer!
Final Answer:
Real-time PCR amplifies and detects viral genetic material by measuring fluorescence during each cycle. If the sample contains SARS-CoV-2 RNA, the fluorescence will increase as the target DNA is amplified, confirming infection. This method is sensitive and allows for quantification of viral load.
Q2. Carlos has 2 symptoms that match with covid 19 symptoms. Would you be reasonably convinced he has a covid 19 infection? Explain.
Background
Topic: Symptom-Based Diagnosis vs. Laboratory Confirmation
This question tests your understanding of the limitations of symptom-based diagnosis and the importance of confirmatory testing in infectious diseases.
Key Terms:
Symptoms: Observable effects of disease (e.g., cough, fever).
Diagnostic Test: Laboratory method to confirm infection (e.g., PCR test).
Step-by-Step Guidance
List the symptoms Carlos is experiencing (cough and fever) and recognize that these are common to many respiratory illnesses.
Consider the overlap of COVID-19 symptoms with other diseases like the flu or common cold.
Explain why laboratory confirmation (such as PCR testing) is necessary to definitively diagnose COVID-19.
Try solving on your own before revealing the answer!
Final Answer:
Having two symptoms alone is not enough to confirm COVID-19, as these symptoms are not unique to the virus. A diagnostic test is required to detect the presence of viral genetic material and confirm infection.
Q3. What is a “false negative” test? How does it happen?
Background
Topic: Diagnostic Test Accuracy
This question tests your understanding of test reliability and the factors that can lead to incorrect results in medical diagnostics.
Key Terms:
False Negative: A test result that incorrectly indicates absence of a condition when it is actually present.
Sensitivity: The ability of a test to correctly identify those with the disease.
Step-by-Step Guidance
Define what a false negative result is in the context of diagnostic testing.
List possible reasons for a false negative, such as low viral load, improper sample collection, or testing at the wrong time during infection.
Discuss how variations in viral presence in different body sites or technical errors can contribute to false negatives.
Try solving on your own before revealing the answer!
Final Answer:
A false negative occurs when a test fails to detect a virus that is actually present. This can happen due to low viral levels, poor sample collection, or testing too early or late in the infection.
Q4. The CDC also determines the particular strain of dengue in each sample using genetic analysis. How do they do this?
Background
Topic: Viral Strain Identification by Genetic Sequencing
This question tests your understanding of how genetic sequencing is used to differentiate between viral strains.
Key Terms and Concepts:
Genetic Sequencing: Determining the order of nucleotides in DNA or RNA.
Serotype: Distinct variations within a species of virus, classified by genetic differences.
Step-by-Step Guidance
After confirming the presence of dengue virus RNA using real-time PCR, the viral RNA is sequenced to determine the nucleotide order.
The obtained sequence is compared to reference sequences in a genetic database to identify the specific serotype.
This comparison allows the CDC to track which strains are circulating and monitor for new variants.
Try solving on your own before revealing the answer!
Final Answer:
The CDC sequences the viral RNA and compares it to known dengue virus sequences in a database. This identifies the specific serotype and helps track the spread of different strains.
Q5. Why would the Centers for Disease Control and Prevention (CDC) be so interested in confirming grandma’s diagnosis?
Background
Topic: Public Health Surveillance and Disease Reporting
This question tests your understanding of the importance of disease confirmation for public health monitoring and outbreak prevention.
Key Terms:
Reportable Disease: Diseases that must be reported to public health authorities.
Outbreak Surveillance: Monitoring and tracking disease cases to prevent spread.
Step-by-Step Guidance
Explain that dengue is a reportable disease and must be tracked by health authorities.
Discuss how confirming cases helps monitor outbreaks, especially in travelers from endemic regions.
Describe how tracking strains and cases informs public health responses and prevention strategies.
Try solving on your own before revealing the answer!
Final Answer:
The CDC confirms diagnoses to monitor outbreaks, track disease spread, and guide public health interventions. This helps prevent larger outbreaks and informs mosquito control and public health warnings.
Vocabulary Chart Guidance
Background
Topic: Key Terms in Recombinant DNA Technology and Molecular Biology
Understanding vocabulary is essential for mastering microbiology concepts. Here are some terms with explanations and relevant images to reinforce your learning.
Recombinant DNA Technology: Combining DNA from different organisms to create new genetic combinations.
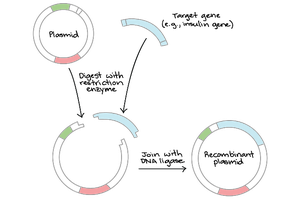
Plasmid: Small, circular DNA molecule used as a vector in genetic engineering.
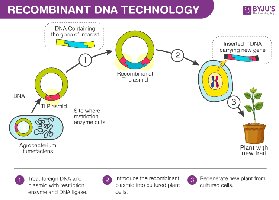
PCR (Polymerase Chain Reaction): Technique to amplify DNA.

Restriction Enzymes: Enzymes that cut DNA at specific sequences.
Sticky Ends: Overhanging DNA sequences that help fragments join together.

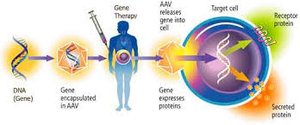
Gene Therapy: Treating diseases by introducing or altering genetic material.

Try filling out the rest of the vocabulary chart on your own before checking your answers!
