 Back
BackMicrobiology Study Notes: Skin and Throat Flora, Staphylococci, and Streptococci
Study Guide - Smart Notes

Skin Flora
Introduction to Skin Microbiota
The human skin is colonized by a diverse microbiota, consisting of approximately 1000 species from 19 phyla. These microorganisms inhabit the superficial layers of the epidermis and hair follicles, playing a crucial role in health and disease.
Microbiota: The community of microorganisms living on the skin.
Non-pathogenic and beneficial: Most skin microbes are harmless and prevent colonization by pathogens by occupying host cell receptors.
Opportunistic pathogens: Resident microbes can cause disease if they gain access to deeper tissues or the bloodstream, leading to conditions such as cellulitis or septicemia.
Examples of Skin Microbiota
The skin hosts a variety of bacteria, with Staphylococcus species being prominent members.
Staphylococcus epidermidis: Common, usually non-pathogenic.
Staphylococcus aureus: Can be pathogenic under certain conditions.
Microbe Transmission and Handwashing
Direct contact with surfaces (animate or inanimate) is a major route for microbe transmission. Proper handwashing is essential for preventing the spread of harmful microbes, especially in healthcare and food service settings.
Handwashing experiment: Culturing finger surfaces before and after washing demonstrates the effectiveness of handwashing.
Handwashing Experiment Results
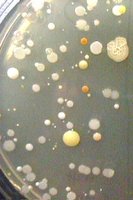
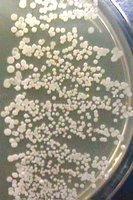
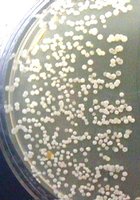
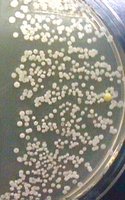
In a typical experiment, finger skin is cultured at four time points: after wetting (Time 0), after 2, 4, and 6 minutes of washing.
Observation: The number of bacterial colonies may increase after washing, but the diversity decreases.
Explanation: Washing removes transient, detergent-susceptible bacteria, leaving tightly adhered resident microbiota.
Staphylococci
Staphylococcus Species and Pathogenicity
Members of the genus Staphylococcus are part of the skin and mucous membrane microbiota.
Opportunistic pathogens: All staphylococci can cause disease under certain conditions.
Staphylococcus aureus: A serious pathogen causing cutaneous and systemic infections (e.g., carbuncles, pneumonia, toxic shock syndrome).
Nosocomial infections: Staphylococci are common in hospital-acquired infections.
Blood Agar and Hemolysis
Staphylococci are cultured on blood agar to assess hemolysis, which helps distinguish pathogenic strains.
β-hemolysis: Complete lysis of red blood cells, characteristic of more pathogenic strains.
Mannitol Salt Agar (MSA)
MSA is a selective-differential medium used to identify Staphylococcus species.
Selective agent: 7.5% NaCl inhibits most bacteria except Staphylococcus spp.
Differential agent: Mannitol; S. aureus ferments mannitol, producing acid and turning phenol red indicator yellow.
MSA Plate Results
Selection: Only Staphylococcus spp. grow.
Differentiation: S. aureus ferments mannitol (yellow), others do not (red).
Coagulase Test
The coagulase test distinguishes S. aureus from other staphylococci.
Coagulase: Enzyme causing blood plasma coagulation.
Free coagulase: Activates prothrombin to thrombin, which converts fibrinogen to fibrin (clot formation).
Bound coagulase: Directly activates fibrinogen.
Tube test: Detects both forms; positive if plasma clots.
Slide test: Detects bound coagulase; positive if agglutination occurs.
Coagulase Reaction Pathway
Latex Agglutination Test
The latex agglutination test uses latex beads coated with proteins that react with bound coagulase and protein A, both associated with S. aureus.
Positive result: Agglutination (clumping) of beads.
Negative result: No agglutination.

Throat Flora
Introduction to Throat Microbiota
The upper respiratory tract (URT) microbiota is diverse, including organisms found on the skin and many others. Most are non-pathogenic and beneficial, but some can cause disease under certain conditions.
Opportunistic pathogens: Resident microbes can cause abscesses or septicemia if mucous membranes are breached.
Transmission: Coughing and sneezing spread microbes.
Key pathogens: Streptococcus pyogenes and Streptococcus pneumoniae.
Streptococci Identification
Streptococci are cultured on blood agar to assess hemolysis and are separated into serological groups (A, B, C, D, etc.) based on cell wall polysaccharides.
Serotyping: Commercial kits are available.
Antibiotic sensitivity: Group A (S. pyogenes) is sensitive to bacitracin; S. pneumoniae is sensitive to optochin.
Throat Swab and Blood Agar
Throat swabs sample the palatine tonsils and are cultured on sheep blood agar to evaluate hemolysis.
Hemolysis types:
α (alpha): Partial hemolysis, green halo
β (beta): Complete hemolysis, clear halo
γ (gamma): No hemolysis
β-hemolytic organism: Streptococcus pyogenes (Group A)
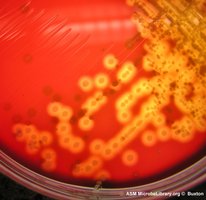
Types of Hemolysis
α-hemolysis: Viridans (green) streptococci
β-hemolysis: Pathogenic streptococci
γ-hemolysis: Non-hemolytic

Streptolysins
Streptolysin S: β-hemolysin, stable in oxygen, detected on surface of blood agar.
Streptolysin O: β-hemolysin, oxygen-labile, detected by stabbing blood agar (less oxygen inside agar).
Group A Streptococcus Antigen Test
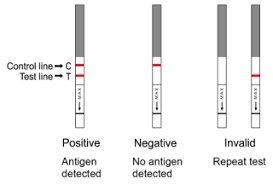
The rapid immunochromatographic assay detects group A streptococcal antigen from throat swabs.
Positive: Two colored lines (control and test) indicate antigen detected.
Negative: One colored line (control) indicates no antigen detected.
Invalid: No control line; test must be repeated.
Additional info
Catalase test: Differentiates staphylococci (catalase positive) from streptococci (catalase negative).
Blood agar precautions: Never perform catalase test directly on blood agar to avoid false positives.
