 Back
BackMolecular Aspects of Viral Replication: HIV, HSV, and SARS-CoV-2
Study Guide - Smart Notes

Molecular Aspects of Viral Replication
Overview of Viral Replication
Viral replication is a fundamental process in microbiology, involving the multiplication of viruses within host cells. Different virus types utilize distinct mechanisms and enzymes for replication, such as RNA-dependent RNA polymerase in RNA viruses and reverse transcriptase in retroviruses.
RNA-dependent RNA polymerase: Enzyme used by single-stranded +RNA, single-stranded -RNA, and double-stranded RNA viruses for genome replication.
Reverse transcriptase: Enzyme used by retroviruses to convert RNA genome into double-stranded DNA.
HIV: Human Immunodeficiency Virus
Classification and Structure
HIV is a retrovirus belonging to the lentivirus group, characterized by a complex genome and slow progression of infection. It causes AIDS and is transmitted through blood and sexual contact. HIV-1 and HIV-2 are the two major types, with HIV-1 being the most prevalent worldwide.
Retrovirus: Virus that reverse-transcribes its RNA genome into DNA.
Lentivirus: Subgroup of retroviruses with complex genomes and slow disease progression.
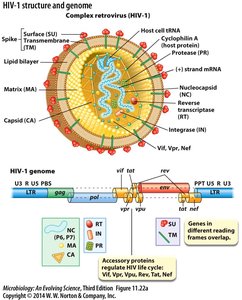
Origin and Evolution
HIV evolved from viruses infecting African monkeys. High mutation rates in infected patients generate multiple virus strains, forming a quasispecies—a dynamic population of diverse mutants with varying replication, tissue tropism, and drug resistance. 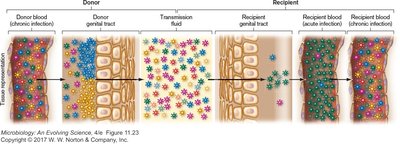
HIV Genome and Accessory Proteins
The HIV genome encodes structural proteins, enzymes, and accessory proteins that regulate viral replication and immune evasion.
Structural proteins: Matrix (MA), Capsid (CA), Nucleocapsid (NC)
Enzymes: Reverse transcriptase (RT), Integrase (IN), Protease (PR)
Accessory proteins: Vif, Vpr, Nef, Vpu, Rev, Tat
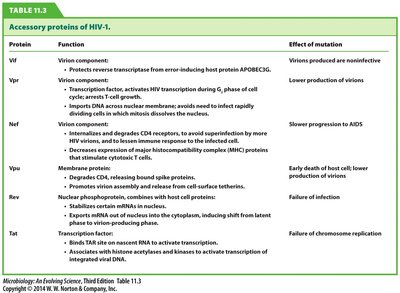
Protein | Function | Effect of Mutation |
|---|---|---|
Vif | Protects reverse transcriptase from host restriction | Virions produced are noninfective |
Vpr | Activates HIV transcription, imports DNA into nucleus | Lower production of virions |
Nef | Downregulates CD4, evades immune response | Slower progression to AIDS |
Vpu | Degrades CD4, releases virions | Early death of host cell, lower production of virions |
Rev | Stabilizes mRNA, exports viral proteins | Failure of infection |
Tat | Activates transcription | Failure of chromosome replication |
HIV Structure: SU-TM Trimer Complex
The HIV envelope contains the SU (gp120) and TM (gp41) proteins, forming a trimer complex essential for host cell attachment and membrane fusion. 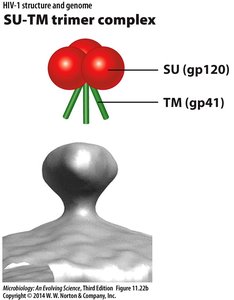
Attachment and Entry into Host Cells
HIV attaches to host cells via the CD4 receptor and CCR5 co-receptor. The gp120 protein binds to CD4, triggering conformational changes that allow gp41 to mediate membrane fusion. Entry inhibitors, such as Maraviroc, block CCR5 binding. 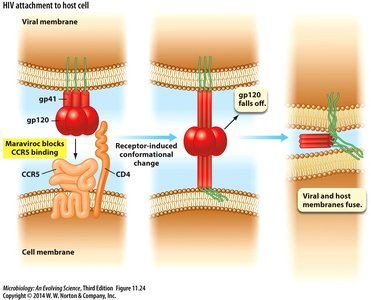
Replication Cycle of HIV
The HIV replication cycle involves several steps:
Attachment to CD4+ T cells
Fusion of viral and host membranes
Reverse transcription of RNA genome into DNA
Integration of viral DNA into host genome
Transcription and translation of viral proteins
Assembly and release of new virions
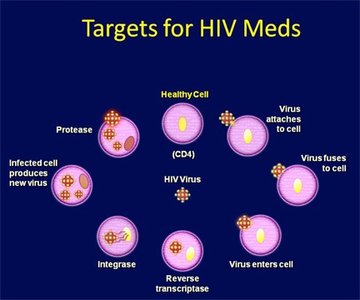
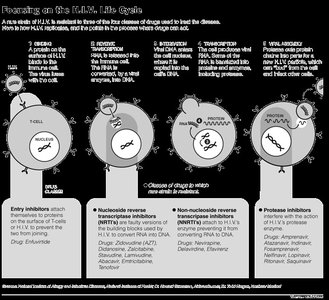
HIV Drug Actions
Nucleoside reverse transcriptase inhibitors: Block reverse transcription (e.g., AZT)
Non-nucleoside reverse transcriptase inhibitors: Inhibit reverse transcriptase by binding to enzyme
Protease inhibitors: Prevent maturation of viral proteins
Fusion inhibitors: Block fusion of viral and host membranes
Entry inhibitors: Block CCR5 co-receptor
Integrase inhibitors: Prevent integration of viral DNA into host genome
Herpes Simplex Virus (HSV)
Classification and Structure
Herpes Simplex Virus (HSV) is a DNA virus with two main strains: HSV-1 (oral lesions) and HSV-2 (genital lesions). Primary infection occurs in epithelial cells, but latent infection can persist in neurons.
HSV-1: Mainly causes oral lesions
HSV-2: Mainly causes genital lesions
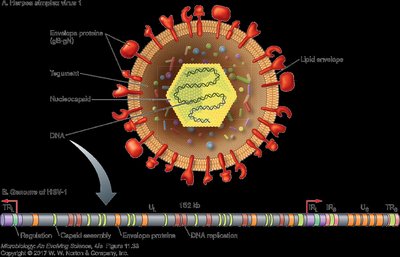
Attachment and Entry
HSV virions use envelope proteins to bind to host cell receptors. Microtubular scaffolds transport virions to the nucleus, where the DNA circularizes for transcription. 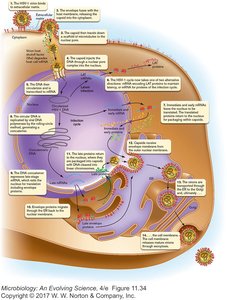
Replication Cycle of HSV
Attachment and entry into epithelial cells
Transport to nucleus and circularization of DNA
Transcription of mRNA for infection cycle or latency
Rolling-circle replication of DNA genome
Production of capsid, matrix, and envelope proteins
Assembly at membrane and release by exocytosis
Persistent Viral Infections
Herpes viruses can establish latency, allowing indefinite infection. Maintenance of latency involves:
Infection of cell types suitable for persistence
Regulation of viral gene expression
Subversion of apoptosis
Evasion of immune responses
SARS-CoV-2: Coronavirus
Replication Cycle
SARS-CoV-2 is a single-stranded RNA virus responsible for COVID-19. Its replication cycle involves entry via endocytosis or membrane fusion, translation of viral proteins, replication of RNA genome, assembly, and release of new virions.
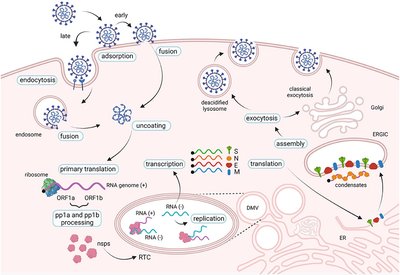
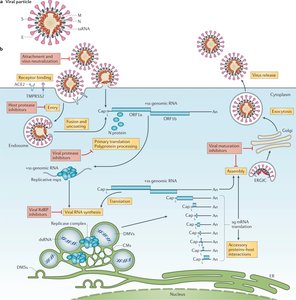
Summary Table: Comparison of HIV, HSV, and SARS-CoV-2
Virus | Genome Type | Replication Enzyme | Latency | Drug Targets |
|---|---|---|---|---|
HIV | ssRNA (retrovirus) | Reverse transcriptase | Yes (integrated DNA) | RT, protease, integrase, entry/fusion |
HSV | dsDNA | DNA polymerase | Yes (neurons) | DNA polymerase, entry |
SARS-CoV-2 | ssRNA (+) | RNA-dependent RNA polymerase | No | RNA polymerase, entry |
Key Equations
Reverse Transcription (HIV):
Rolling-Circle Replication (HSV):
RNA Replication (SARS-CoV-2):
Additional info:
Highly active antiretroviral therapy (HAART) combines multiple drug classes to suppress HIV replication.
Herpes viruses can cause a range of diseases, including chickenpox, mononucleosis, and Kaposi's sarcoma.
SARS-CoV-2 replication involves both cytoplasmic and membrane-associated steps, with unique mechanisms for RNA synthesis and protein translation.
