 Back
BackMolecular Aspects of Viruses: Baltimore Classification, Replication, and Viral Ecology
Study Guide - Smart Notes

Virus Classification and Molecular Aspects
Baltimore Virus Classification
The Baltimore classification system organizes viruses based on their genome composition and the mechanisms of replication and mRNA expression. This system defines seven fundamental groups, each with distinct replication strategies and genome types.
Group I: Double-stranded DNA viruses
Group II: Single-stranded DNA viruses
Group III: Double-stranded RNA viruses
Group IV: (+) sense single-stranded RNA viruses
Group V: (–) sense single-stranded RNA viruses
Group VI: Retroviruses (RNA reverse-transcribing viruses)
Group VII: Pararetroviruses (DNA reverse-transcribing viruses)
Each group utilizes unique enzymes and pathways to produce mRNA and replicate their genomes. The primary factor determining the life cycle of an animal virus is the form of its genome.
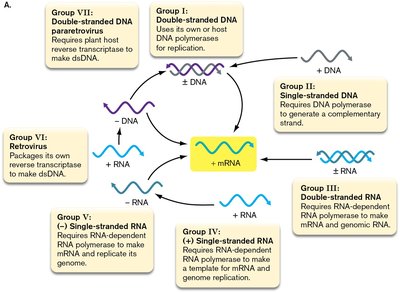
Genome Replication Strategies
Viruses employ different strategies depending on their genome type:
DNA viruses: Can utilize the host replication machinery.
RNA viruses: Use an RNA-dependent RNA polymerase to transcribe their mRNA.
Retroviruses: Use a reverse transcriptase to copy their genomic sequence into DNA for insertion in the host chromosome.
Animal Virus Replication Cycles
Attachment, Entry, and Uncoating
Animal viruses initiate infection by binding to specific host cell receptors, followed by entry and genome uncoating. The mechanism of entry and uncoating varies by virus type and host cell.
Attachment: Viral proteins bind to host cell surface receptors.
Entry: Can occur via direct fusion, endocytosis, or other mechanisms.
Uncoating: The viral genome is released into the cytoplasm or nucleus.
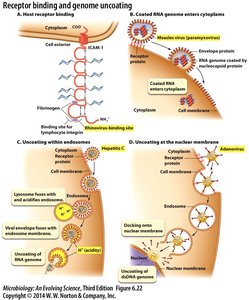
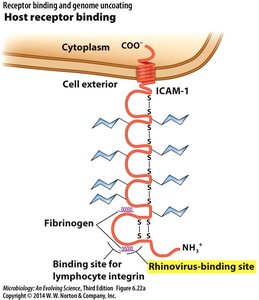
Host Receptor Binding Example
Viruses such as rhinovirus bind to specific sites on host cell receptors like ICAM-1, facilitating entry.
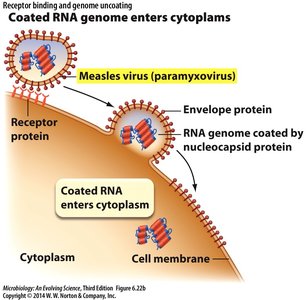
Entry Mechanisms
Coated RNA genome enters cytoplasm: Example: Measles virus (paramyxovirus).
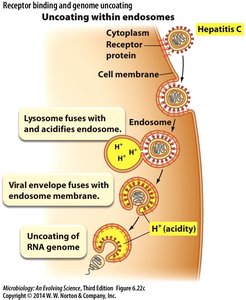
Uncoating within endosomes: Example: Hepatitis C virus, where acidification triggers genome release.
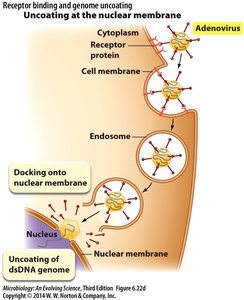
Uncoating at the nuclear membrane: Example: Adenovirus, which docks at the nuclear membrane for genome release.
Viral Genome Expression and Assembly
All animal viruses make proteins using host ribosomes, with translation occurring in the cytoplasm. Assembly of new virions involves the formation of capsids and packaging of the genome, which may occur in the cytoplasm or nucleus. Envelope proteins are inserted into cellular membranes.
Release of Progeny Viruses
Lysis: Host cell is destroyed, releasing virions.
Budding: Virions exit the cell by budding through membranes, acquiring a lipid envelope.
All enveloped viruses bud from a membrane, either the plasma membrane or an organelle membrane.
Specific Virus Life Cycles
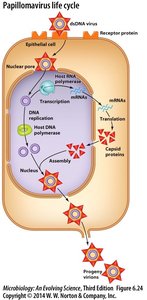
Papillomavirus (DNA Virus) Life Cycle
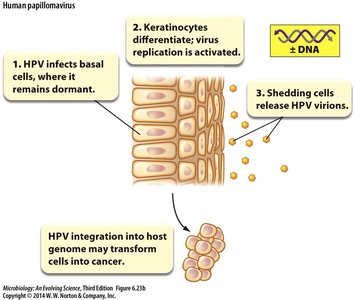
Human papillomavirus (HPV) infects basal cells, remains dormant, and activates replication upon keratinocyte differentiation. Shedding cells release HPV virions, and integration into the host genome may transform cells into cancerous states.
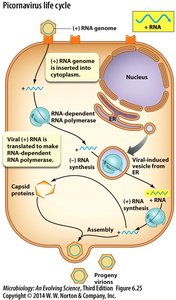
Picornavirus (RNA Virus) Life Cycle
Picornaviruses, such as poliovirus, have a (+) RNA genome that is translated directly in the cytoplasm. Viral RNA-dependent RNA polymerase synthesizes new RNA genomes, and assembly occurs in the cytoplasm.
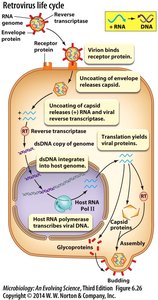
Retrovirus Life Cycle
Retroviruses, such as HIV, use reverse transcriptase to convert their (+) RNA genome into DNA, which integrates into the host genome. Host RNA polymerase transcribes viral DNA, and assembly occurs in the cytoplasm.
Oncogenic Viruses
Mechanisms of Oncogenesis
Oncogenic viruses can transform host cells to become cancerous through several mechanisms:
Insertion of an oncogene into the host genome
Integration of the entire viral genome
Expression of viral proteins that interfere with host cell cycle regulation
Examples include Epstein-Barr virus (lymphomas) and hepatitis C virus (liver cancer).
Plant Virus Replication Cycles
Entry and Transmission
Plant viruses infect cells by mechanisms that do not involve specific membrane receptors. Entry usually requires mechanical transmission, such as contact with damaged tissues, transmission by animal vectors, or through seeds.
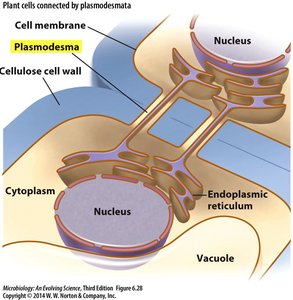
Cell-to-Cell Movement
Plant cells are connected by plasmodesmata, which facilitate the movement of viruses between cells.
Host Defenses Against Viruses
Defense Mechanisms
Genetic resistance: Hosts experience mutations that confer resistance.
Immune system: Includes innate immunity (interferons) and adaptive immunity (antibodies).
RNA interference (RNAi): Widespread among eukaryotes and archaea, silencing viral RNA.
Viral Ecology and Roles in Ecosystems
Persistence and Emergence
Viruses exist within host organisms in complex ecosystems, influencing population dynamics and host diversity. Some viruses persist in the wild with broad host ranges, while new viruses can emerge through human interaction with wildlife or as variants of endemic pathogens.
Ecological Functions
Limiting host population density without causing extinction
Participating in population decline and resurgence
Selecting for host diversity and fostering evolution of distinct species
Influenza Virus: Structure and Replication
Structure of Influenza A Virus
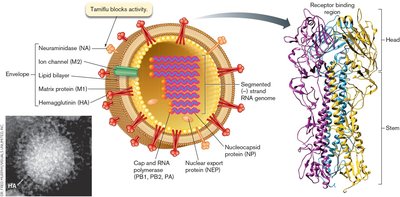
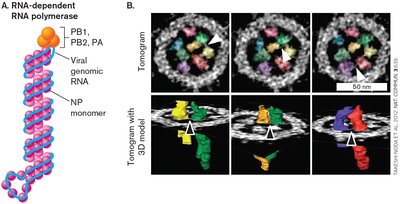
Influenza A is a (–) strand RNA virus with a segmented genome. Its structure includes envelope proteins (hemagglutinin, neuraminidase), matrix proteins, and RNA-dependent RNA polymerase.
Genome Packaging and Replication
The RNA complexes are selectively packaged in the virion, and the genome consists of multiple segments, each encoding different viral proteins.
Attachment and Entry
Influenza virus attaches to host cells via hemagglutinin binding to sialic acid receptors. Entry occurs through endocytosis, followed by acidification-induced fusion and release of viral contents into the cytoplasm.
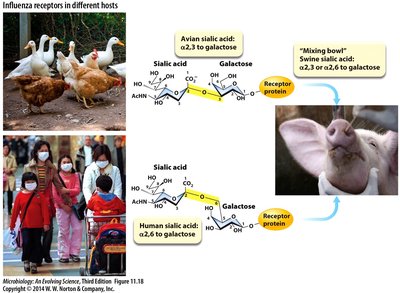
Antigenic Shift and Drift
Influenza viruses undergo antigenic shift (reassortment of genome segments) and antigenic drift (accumulation of mutations), leading to new strains and challenges for vaccine development.
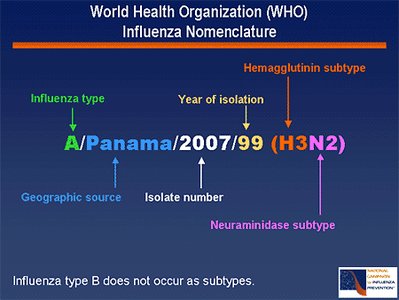
Vaccine Strains
Annual influenza vaccines are formulated based on circulating strains, including H1N1, H3N2, and B/Victoria lineage viruses.
Summary Table: Baltimore Virus Groups
Group | Genome Type | Replication Strategy | Key Enzyme |
|---|---|---|---|
I | dsDNA | Host or viral DNA polymerase | DNA polymerase |
II | ssDNA | DNA polymerase makes complementary strand | DNA polymerase |
III | dsRNA | RNA-dependent RNA polymerase | RNA polymerase |
IV | (+) ssRNA | Direct translation or RNA polymerase for replication | RNA polymerase |
V | (–) ssRNA | RNA-dependent RNA polymerase | RNA polymerase |
VI | (+) ssRNA (retrovirus) | Reverse transcription to DNA | Reverse transcriptase |
VII | dsDNA (pararetrovirus) | Reverse transcription to DNA | Reverse transcriptase |
Additional info: Academic context was added to clarify mechanisms, provide definitions, and expand on brief points for completeness.
