 Back
BackMolecular Biology: DNA, RNA, and Gene Regulation in Microbiology
Study Guide - Smart Notes

Genotype, Phenotype, and the Central Dogma
Definitions and Concepts
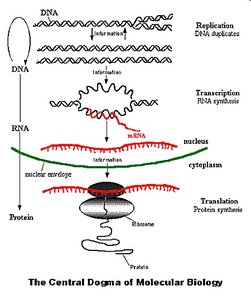
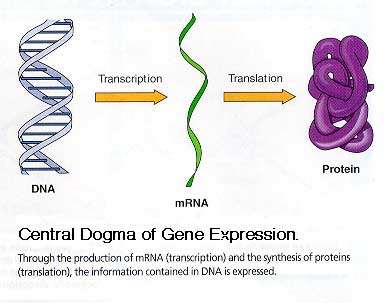
The genotype refers to the complete set of genes in an organism, representing its genetic potential. The phenotype is the observable expression of these genes, typically as proteins and traits. The Central Dogma of Molecular Biology describes the flow of genetic information: DNA is replicated, transcribed into RNA, and then translated into protein.
Genotype: All genetic material (DNA) of an organism.
Phenotype: Expressed traits (proteins) resulting from gene expression.
Central Dogma: DNA → RNA → Protein
DNA: Structure and Organization
Nucleotides and DNA Structure
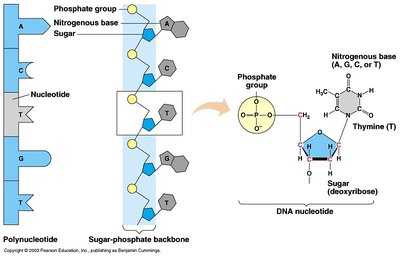
DNA is a polymer made of nucleotide monomers. Each nucleotide consists of a phosphate group, a deoxyribose sugar, and a nitrogenous base (A, T, G, C). Genes are stretches of DNA coding for functional products, and chromosomes are large DNA molecules containing many genes. The genome is the total genetic material in a cell.
Nucleotides: Monomers of DNA and RNA; composed of phosphate, sugar, and base.
Chromosome: Large DNA molecule with many genes.
Genome: All genetic material in a cell.
Nitrogenous Bases and Base Pairing
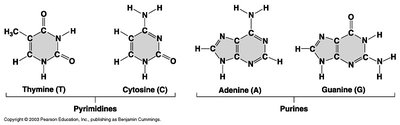
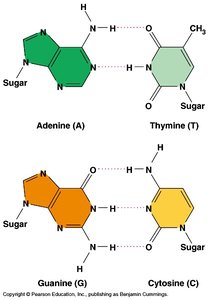
DNA contains four nitrogenous bases: adenine (A), thymine (T), guanine (G), and cytosine (C). Bases pair specifically: A with T, and G with C. These pairs are stabilized by hydrogen bonds.
Pyrimidines: Thymine (T), Cytosine (C)
Purines: Adenine (A), Guanine (G)
Base Pairing: A:T (2 hydrogen bonds), G:C (3 hydrogen bonds)
Antiparallel Strands and Double Helix
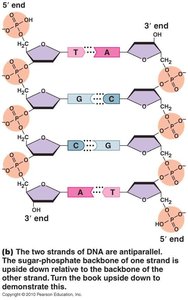
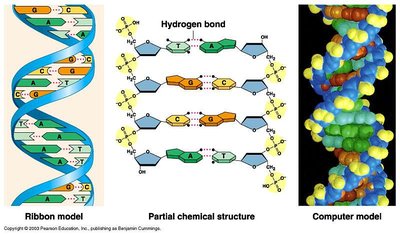
DNA is a double-stranded helix with antiparallel strands, meaning the two strands run in opposite directions (5' to 3' and 3' to 5'). The sugar-phosphate backbone is held together by covalent bonds, while the two strands are joined by hydrogen bonds between bases.
Antiparallel: Strands run in opposite directions.
Double Helix: Twisted ladder structure of DNA.
DNA Replication
Semi-Conservative Replication
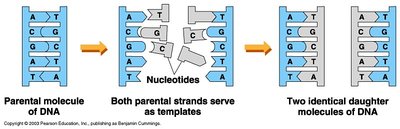
DNA replication is semi-conservative: each new DNA molecule contains one original (parental) strand and one newly synthesized strand. This ensures genetic continuity during cell division.
Template: Each strand serves as a template for synthesis of a new strand.
Accuracy: Proofreading and repair mechanisms ensure high fidelity.
Origin of Replication and Bidirectional Replication
Replication begins at specific sites called origins of replication. In prokaryotes, replication is bidirectional from a single origin on a circular chromosome.
Origin of Replication: Specific DNA sequence where replication starts.
Bidirectional: Replication proceeds in both directions from the origin.
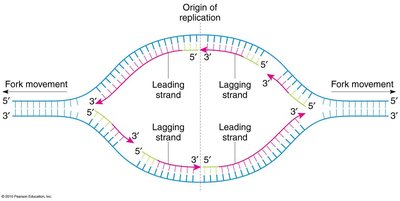
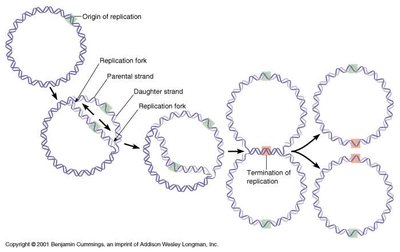
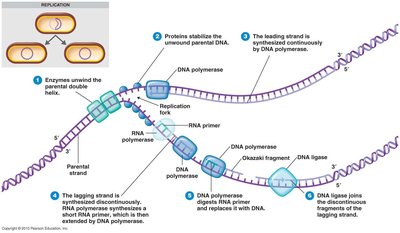
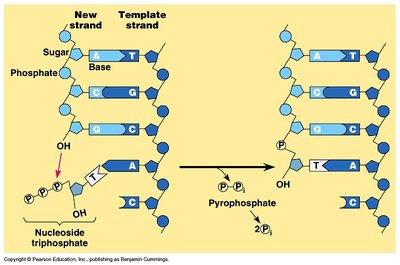
Enzymes and Steps in DNA Replication
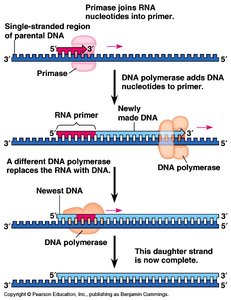
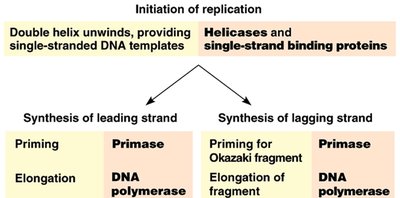
Multiple enzymes coordinate DNA replication. Primase synthesizes RNA primers, DNA polymerase extends the DNA chain, and DNA ligase joins fragments. Leading strand synthesis is continuous, while lagging strand synthesis is discontinuous (Okazaki fragments).
Primase: Synthesizes RNA primers.
DNA Polymerase: Extends DNA in 5' to 3' direction.
DNA Ligase: Joins Okazaki fragments on lagging strand.
Gyrase (Topoisomerase): Relaxes DNA ahead of replication fork.
Helicase: Unwinds double-stranded DNA.
Single-strand binding proteins: Stabilize unwound DNA.

Gene Regulation and Expression
RNA Structure and Types
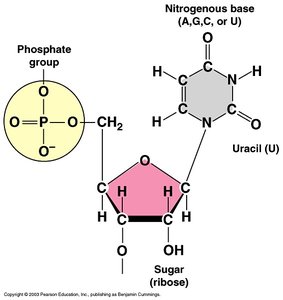
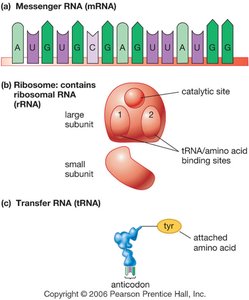
RNA is single-stranded and contains ribose sugar and uracil (U) instead of thymine (T). There are three main types of RNA: messenger RNA (mRNA), ribosomal RNA (rRNA), and transfer RNA (tRNA).
mRNA: Carries genetic code from DNA to ribosome.
rRNA: Combines with proteins to form ribosomes.
tRNA: Brings amino acids to ribosome during translation.

Transcription: DNA to RNA
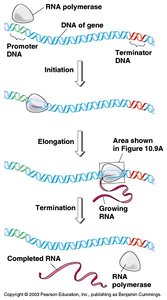
Transcription is the process of synthesizing RNA from a DNA template. RNA polymerase binds to the promoter, synthesizes RNA, and detaches at the terminator sequence.
Initiation: RNA polymerase binds to promoter.
Elongation: RNA polymerase synthesizes RNA.
Termination: RNA polymerase detaches at terminator.
Prokaryotic vs. Eukaryotic Gene Expression
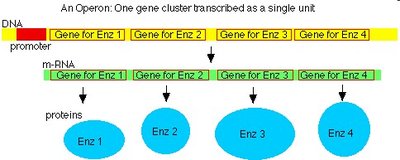
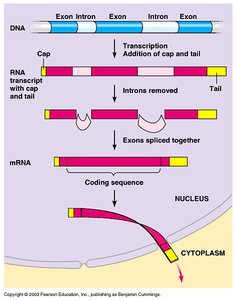
Prokaryotic genes are organized in operons and lack introns. Eukaryotic genes contain exons (coding) and introns (noncoding), and pre-mRNA must be processed (splicing, capping, tailing) before export from the nucleus.
Prokaryotes: No introns, operons, simultaneous transcription and translation.
Eukaryotes: Introns and exons, mRNA processing, transcription in nucleus, translation in cytoplasm.
The Genetic Code and Translation
The Genetic Code
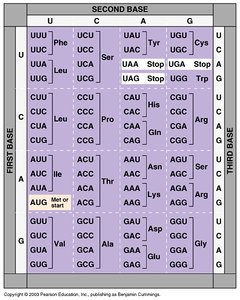
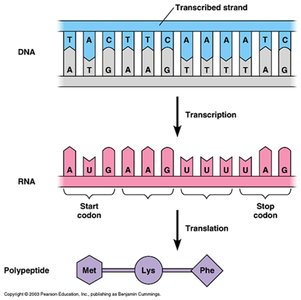
The genetic code translates nucleotide sequences into amino acids. Codons are three-nucleotide sequences in mRNA that specify amino acids. The code is redundant; multiple codons can code for the same amino acid.
Codon: Three-nucleotide sequence coding for an amino acid.
Redundancy: Multiple codons for most amino acids.
Translation: RNA to Protein
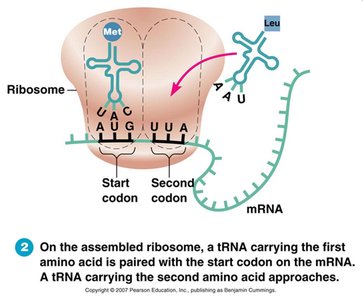
Translation is the synthesis of proteins from mRNA. It occurs in three stages: initiation, elongation, and termination. Ribosomes facilitate the process, and tRNA brings amino acids to the growing polypeptide chain.
Initiation: Ribosome assembles at start codon (AUG).
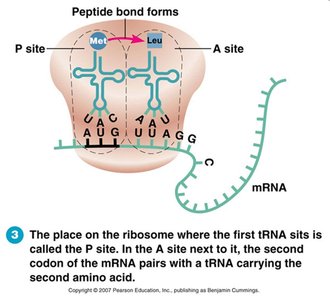
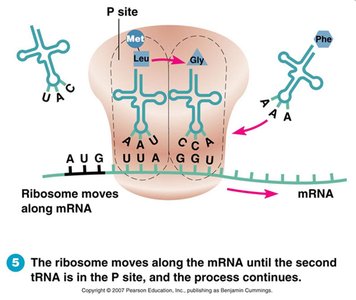
Elongation: tRNA brings amino acids, peptide bonds form.
Termination: Ribosome reaches stop codon, polypeptide released.
Prokaryotic Gene Regulation
Operons and Regulation Types
Gene regulation in prokaryotes involves operons, which are clusters of genes controlled by a single promoter and operator. Genes can be constitutive (always on), inducible (activated by substrate), or repressible (turned off by product).
Constitutive: Always expressed (e.g., rRNA genes).
Inducible: Off until substrate is present (e.g., lac operon).
Repressible: On until product accumulates (e.g., biosynthetic operons).
Catabolite Repression
Catabolite repression is a regulatory mechanism where the presence of glucose inhibits the transcription of other operons (such as the lac operon). When glucose is absent, cAMP levels rise, activating CAP, which promotes transcription.
Glucose present: Low cAMP, CAP inactive, operon off.
Glucose absent: High cAMP, CAP active, operon on.
Summary Table: Enzymes of DNA Replication
Enzyme | Function |
|---|---|
Gyrase (Topoisomerase) | Relaxes DNA ahead of replication fork |
Helicase | Unwinds double-stranded DNA |
Primase | Synthesizes RNA primers |
Single-strand binding proteins | Stabilize unwound DNA |
DNA polymerase | Synthesizes new DNA strand |
DNA ligase | Joins Okazaki fragments |
Key Equations
Base Pairing Rule:
DNA Replication: (semi-conservative)
Transcription:
Translation:
----------------------------------------
