 Back
Backlec 15:Molecular Complexity, Bacterial Defense, and Synthetic Biology: Microbial Arms Race and Biotechnology Applications
Study Guide - Smart Notes

Molecular Complexity and the Microbial Arms Race
Bacteria and Phages: An Evolutionary Arms Race
Bacteria and bacteriophages (phages) have co-evolved for billions of years, engaging in a dynamic evolutionary struggle. This arms race has driven the development of sophisticated molecular mechanisms for both attack and defense, resulting in remarkable biological diversity and innovation.
Bacteriophages are viruses that specifically infect bacteria, injecting their genetic material to hijack the host's cellular machinery.
Evolutionary arms race refers to the continuous cycle of adaptation and counter-adaptation between bacteria and phages.
This dynamic is often described as the "Red Queen" effect, where both sides must constantly evolve to maintain their relative fitness.
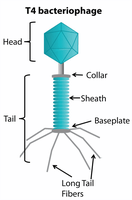
Bacterial Defense Mechanisms Against Phages
CRISPR-Cas Systems: Adaptive Immunity
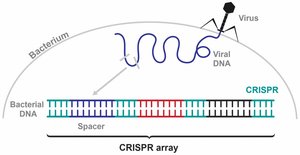
Bacteria have evolved the CRISPR-Cas system as an adaptive immune mechanism to recognize and destroy invading viral DNA. This system provides sequence-specific defense and a form of molecular memory.
CRISPR array: A region in the bacterial genome containing short, repetitive DNA sequences interspaced with "spacers" derived from previous phage infections.
Memory: Surviving a phage attack, bacteria incorporate a fragment of phage DNA into their CRISPR array.
Recognition: Upon reinfection, the bacterium transcribes the spacer into guide RNA, which directs the Cas enzyme to the matching phage DNA.
Destruction: The Cas enzyme (e.g., Cas9) cleaves the invading DNA, neutralizing the threat.
Restriction-Modification (R-M) Systems
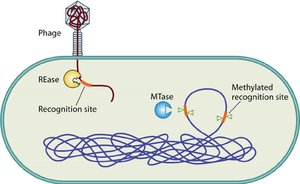
Restriction-modification systems act as a bacterial innate immune system, distinguishing self from non-self DNA through methylation patterns.
Restriction enzymes (REases) cut foreign DNA at specific recognition sites.
Methyltransferases (MTases) add methyl groups to the host's own DNA at these sites, protecting it from cleavage.
Unmethylated phage DNA is recognized as foreign and degraded.
Surface Masking
Bacteria can evade phage infection by altering or hiding the surface receptors that phages use for attachment, effectively preventing viral entry.
Mutations or modifications in cell wall proteins can block phage binding.
This strategy is analogous to "locking the door" against viral invaders.
Abortive Infection (Abi) Systems
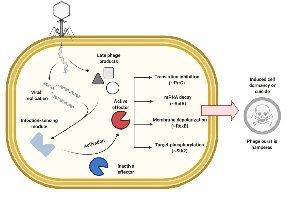
Abortive infection is a self-sacrificial defense where an infected bacterium triggers its own death to prevent phage replication and protect the colony.
Abi systems detect phage infection and activate cell death pathways.
This limits the spread of phages within the bacterial population.
Phage Counter-Defenses
Anti-CRISPR Proteins
Phages have evolved anti-CRISPR (Acr) proteins that inhibit the CRISPR-Cas system, allowing them to evade bacterial immunity.
Acr proteins bind to Cas enzymes, blocking their DNA-cutting activity.
This neutralizes the bacterial adaptive immune response.
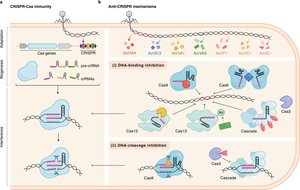
DNA Shielding and Epigenetic Mimicry
Some phages protect their genomes by physically shielding their DNA or by mimicking host methylation patterns.
DNA shielding: Formation of a protein shell around phage DNA to prevent recognition by host defenses.
Epigenetic mimicry: Phage-encoded methyltransferases add methyl groups to phage DNA, disguising it as host DNA.
Hyper-Mutation
Phages can rapidly mutate their genomes, enabling them to escape recognition by CRISPR systems and other host defenses.
High mutation rates generate genetic diversity, increasing the likelihood of evading bacterial immunity.
Biotechnological Applications: Harnessing Bacterial Weapons
CRISPR-Cas9 Technology in Genome Editing
CRISPR-Cas9, originally a bacterial defense system, has been adapted as a powerful tool for precise genome editing in various organisms.
Cas9 enzyme: Acts as molecular scissors to cut DNA at targeted sites.
Guide RNA (gRNA): Designed to match a specific DNA sequence, directing Cas9 to the correct location.
PAM sequence: A short DNA motif required for Cas9 binding and activity (e.g., NGG).
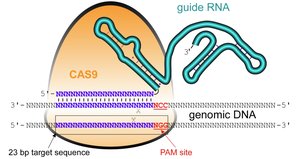
Mechanism of CRISPR-Cas9 Editing
The CRISPR-Cas9 system enables targeted DNA modifications through a three-step process: scanning, matching, and cutting.
Scanning: Cas9-gRNA complex searches for PAM sites in the genome.
Unzipping & Matching: DNA is unwound, and the gRNA checks for sequence complementarity.
Cutting: Upon a perfect match, Cas9 cleaves both DNA strands, creating a double-strand break.
DNA Repair Pathways and Gene Editing Outcomes
Following Cas9-induced DNA breaks, cells repair the damage via two main pathways, leading to different gene editing outcomes.
Non-Homologous End Joining (NHEJ): Error-prone repair that often introduces insertions or deletions, disrupting gene function (gene knockout).
Homology-Directed Repair (HDR): Accurate repair using a provided DNA template, enabling precise gene insertion or correction.
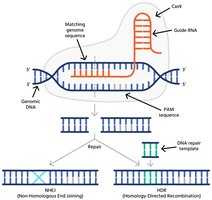
Synthetic Microbiology and Biotechnology
Engineering Bacteria as Living Factories
Synthetic biology enables the design and modification of bacterial genomes to produce valuable products, such as pharmaceuticals, biofuels, and industrial enzymes.
Gene selection: Identify and isolate the gene encoding the desired product (e.g., human insulin).
Recombinant DNA technology: Use restriction enzymes and DNA ligase to insert the gene into a plasmid vector.
Transformation: Introduce the recombinant plasmid into a suitable bacterial host (e.g., E. coli).
Scale-up: Grow engineered bacteria in bioreactors for large-scale production.
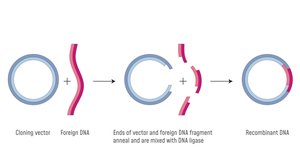
Bacterial Transformation and Selection
Transformation is the process by which bacteria take up external DNA, enabling genetic engineering. Selection markers, such as antibiotic resistance genes, help identify successful transformants.
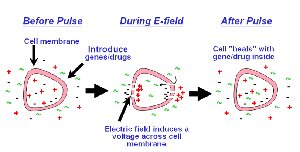
Inducing competence: Methods include chemical treatment (CaCl2 + heat shock) or electroporation.
Selection: Only bacteria that acquire the plasmid survive on antibiotic-containing media.
Scale-Up, Fermentation, and Downstream Processing
Industrial biotechnology involves scaling up bacterial cultures and purifying the desired product through a series of controlled steps.
Fermentation: Large bioreactors maintain optimal conditions for bacterial growth and product synthesis.
Harvesting: Cells are separated from the culture medium by centrifugation or filtration.
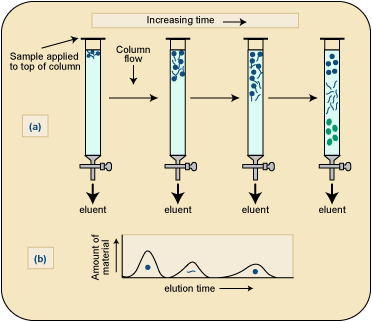
Purification: Chromatography and other techniques isolate the target product.
Quality control: Ensures product purity, activity, and safety, especially for medical applications.

Applications of Engineered Bacteria
Genetically modified bacteria have revolutionized medicine and industry by enabling the production of complex biomolecules and sustainable materials.
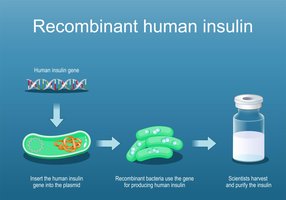
Human insulin production: Recombinant E. coli produce human insulin, replacing animal-derived sources and reducing allergic reactions.
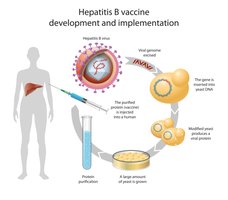
Recombinant vaccines: Bacteria express viral antigens for safe and effective vaccine development (e.g., hepatitis B).
Bioplastics: Engineered bacteria synthesize biodegradable plastics (polyhydroxyalkanoates, PHAs) for eco-friendly materials.
Summary Table: Bacterial Defense and Biotechnological Applications
Defense Mechanism | Description | Biotechnological Application |
|---|---|---|
CRISPR-Cas | Adaptive immunity; sequence-specific DNA cleavage | Genome editing (CRISPR-Cas9) |
Restriction-Modification | Innate immunity; restriction enzymes cut foreign DNA | DNA cloning, molecular biology tools |
Surface Masking | Alteration of phage receptors to prevent infection | Understanding bacterial resistance |
Abortive Infection | Programmed cell death to limit phage spread | Research on population-level immunity |
