 Back
Back6: Molecular Information Flow and Protein Processing in Microorganisms
Study Guide - Smart Notes

Molecular Information Flow and Protein Processing
DNA and Genetic Information Flow
The flow of genetic information in microorganisms is governed by the central dogma, which describes the transfer of information from DNA to RNA to protein. Genes, the functional units of genetic information, are part of larger genetic elements such as chromosomes and plasmids. The genome encompasses all genetic elements within a cell.
DNA: Serves as the genetic blueprint for cellular functions.
RNA: Transcription product of DNA, includes messenger RNA (mRNA), transfer RNA (tRNA), and ribosomal RNA (rRNA).
Informational macromolecules: Nucleic acids (DNA and RNA) and proteins.
Central Dogma: Information flows from DNA (replication) to RNA (transcription) to protein (translation).
Genome: The complete set of genetic elements in a cell.
Gene Expression: The process by which information from a gene is used to synthesize RNA and protein.
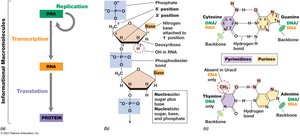
Nucleic Acid Structure and Composition
Nucleic acids are polymers of nucleotides, each consisting of a pentose sugar, a nitrogenous base, and a phosphate group. DNA and RNA differ in their sugar and base composition.
Nucleotide: Monomer of nucleic acids; contains pentose sugar (ribose in RNA, deoxyribose in DNA), nitrogenous base, and phosphate.
Nucleoside: Contains pentose sugar and nitrogenous base, but no phosphate.
Nitrogenous Bases: Purines (adenine, guanine) and pyrimidines (cytosine, thymine in DNA, uracil in RNA).
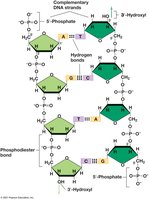
Properties of the DNA Double Helix
DNA is a double-stranded molecule held together by hydrogen bonds between complementary bases. The strands are antiparallel and form a double helix with major and minor grooves, which are important for protein binding.
Phosphodiester Bonds: Connect the 3′-carbon of one sugar to the 5′-carbon of the adjacent sugar.
Base Pairing: Adenine pairs with thymine; guanine pairs with cytosine.
Antiparallel Strands: One strand runs 5′ to 3′, the other 3′ to 5′.
Grooves: Major groove (protein binding site) and minor groove.
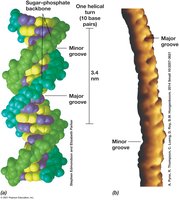
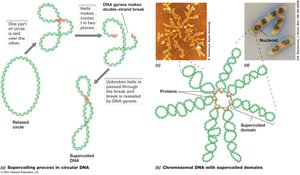
Size, Shape, and Supercoiling of DNA
DNA size is measured in base pairs, and its length is much greater than the cell, requiring supercoiling for compaction. Topoisomerases, including DNA gyrase, regulate supercoiling.
Supercoiling: Compacts DNA to fit within the cell.
Topoisomerases: Enzymes that insert or remove supercoils.
Negative Supercoiling: Common in most cells; positive supercoiling prevents DNA melting at high temperatures.
Genes and Steps in Biological Information Flow
Gene expression involves replication, transcription, and translation. Replication duplicates DNA, transcription synthesizes RNA from DNA, and translation builds proteins from mRNA.
Replication: DNA is duplicated by DNA polymerase.
Transcription: DNA information is transferred to RNA by RNA polymerase.
Translation: mRNA is used to build polypeptides on ribosomes.
RNA Classes: mRNA (carries information), tRNA (translates information), rRNA (structural and catalytic ribosome component).
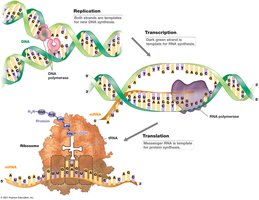
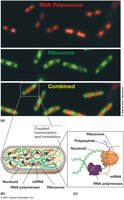
Coupled Transcription and Translation in Prokaryotes
In prokaryotes, transcription and translation are often coupled, allowing rapid protein synthesis. Multiple genes may be transcribed into a single mRNA.
Coupled Process: Transcription and translation occur simultaneously.
Polycistronic mRNA: Single mRNA encoding multiple proteins.
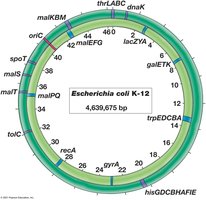
Genetic Elements: Chromosomes and Plasmids
Chromosomes
The chromosome is the main genetic element in prokaryotes, typically a single circular DNA molecule. Eukaryotes have multiple linear chromosomes.
Chromosome: Main genetic element; carries most genes.
Other Elements: Virus genomes, plasmids, organellar genomes, transposable elements.
Plasmids
Plasmids are extrachromosomal DNA elements that replicate independently. They often carry genes for antibiotic resistance, virulence, or metabolic functions.
Plasmids: Circular or linear double-stranded DNA; nonessential but may confer advantages.
R Plasmids: Resistance plasmids; encode antibiotic resistance genes.
Virulence Factors: Some plasmids encode factors for pathogenicity.
Bacteriocins: Proteins encoded by plasmids that inhibit or kill related species.
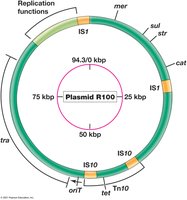
Transposable Elements
Transposable elements are DNA segments that can move within and between DNA molecules, contributing to genetic diversity and evolution.
Transposable Elements: Found in prokaryotes and eukaryotes; can move between chromosomes, plasmids, and viral genomes.
DNA Replication: Templates, Enzymes, and the Replication Fork
Semiconservative Replication
DNA replication is semiconservative, meaning each new double helix contains one parental and one new strand. Replication proceeds from the 5′ to 3′ end.
Semiconservative: Each daughter DNA contains one original and one new strand.
Deoxynucleoside 5′-triphosphate: Precursor for nucleotide addition.
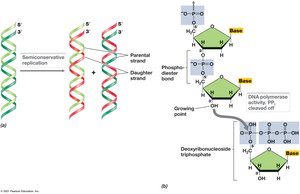
Replication Enzymes and Priming
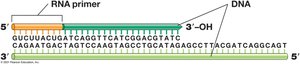
DNA polymerases catalyze the polymerization of deoxynucleotides. Primase synthesizes RNA primers required for DNA polymerase activity.
DNA Polymerases: Multiple types; DNA Pol III is primary for chromosomal replication.
Primer: Short RNA stretch synthesized by primase; required for DNA polymerase.
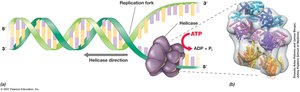
Initiation of DNA Synthesis
Replication begins at the origin of replication, where helicase unwinds the double helix. Loader proteins and primase are recruited to start DNA synthesis.
Origin of Replication (oriC): Site where replication begins.
Helicase: Unwinds DNA helix.
Loader Proteins: Assist helicase binding.
Leading and Lagging Strands
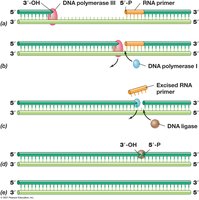
DNA synthesis is continuous on the leading strand and discontinuous on the lagging strand, forming Okazaki fragments. DNA ligase seals nicks between fragments.
Leading Strand: Synthesized continuously.
Lagging Strand: Synthesized discontinuously; requires multiple primers.
DNA Ligase: Seals nicks in DNA.
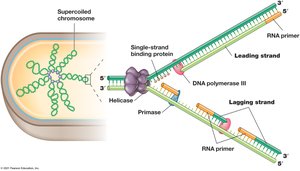
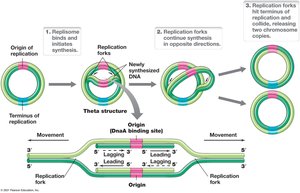
Bidirectional Replication and the Replisome
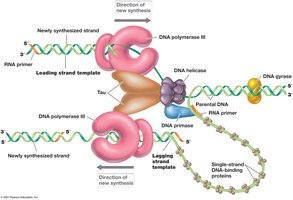
Replication in prokaryotes is bidirectional due to the circular chromosome, with two replication forks moving in opposite directions. The replisome is a large protein complex responsible for DNA synthesis.
Bidirectional Replication: Two forks move away from the origin.
Replisome: Complex of proteins including primosome (helicase and primase).
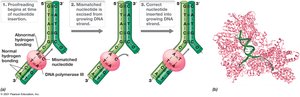
Proofreading and Fidelity
DNA replication is highly accurate due to proofreading by DNA polymerase, which detects and corrects mismatches using exonuclease activity.
Proofreading: Ensures high fidelity; removes mismatched bases.
Exonuclease Activity: Removes incorrect nucleotides.
RNA Synthesis: Transcription
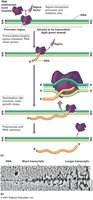
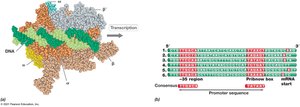
Transcription in Bacteria
Transcription is the synthesis of RNA from a DNA template, catalyzed by RNA polymerase. RNA is typically single-stranded and folds into secondary structures.
RNA Polymerase: Catalyzes RNA synthesis; consists of multiple subunits.
Promoters: DNA sequences recognized by sigma factors for transcription initiation.
Termination: Specific DNA sequences or proteins (Rho) end transcription.
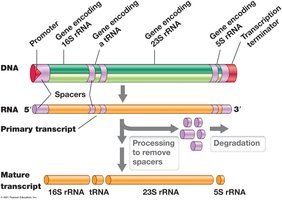
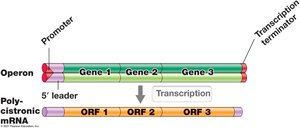
Transcriptional Units and Polycistronic mRNA
Transcriptional units may encode one or multiple genes. Operons are transcribed into polycistronic mRNA, which contains multiple open reading frames.
Transcriptional Unit: DNA segment transcribed into one RNA molecule.
Operon: Cluster of genes transcribed as a single mRNA.
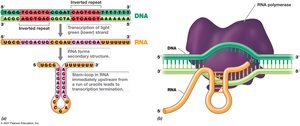
Termination of Transcription
Termination is governed by DNA sequences such as inverted repeats, which cause stem-loop formation in RNA, or by Rho-dependent mechanisms.
Stem-Loop: Formed by intra-strand base pairing; causes RNA polymerase to pause.
Rho Protein: Recognizes specific sequences and releases RNA polymerase.
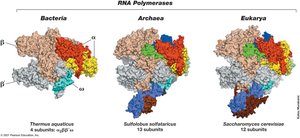
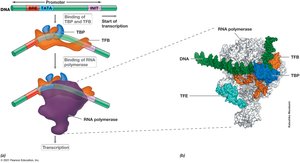
Transcription in Archaea and Eukarya
Archaeal transcription is more similar to eukaryotic transcription than bacterial. Eukaryotes have three RNA polymerases, and transcription initiation involves the TATA box and transcription factors.
TATA Box: Recognition sequence for transcription initiation.
Transcription Factors: TBP and TFB enable RNA polymerase binding.
RNA Processing in Eukaryotes and Archaea
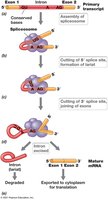
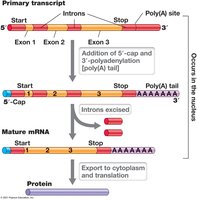
Eukaryotic genes contain exons and introns. RNA processing includes splicing, capping, and polyadenylation to produce mature mRNA.
Splicing: Removal of introns and joining of exons.
Capping: Addition of methylated guanine to 5′ end.
Polyadenylation: Addition of poly(A) tail to 3′ end.
Protein Synthesis: Translation
Amino Acids, Polypeptides, and Proteins
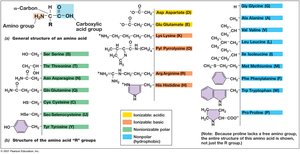
Proteins are polymers of amino acids linked by peptide bonds. They serve catalytic, structural, and regulatory functions in cells.
Amino Acids: Organic compounds with amino and carboxylic acid groups attached to α-carbon.
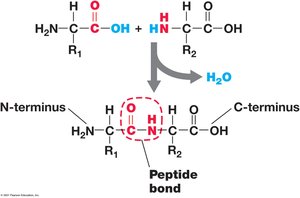
Peptide Bond: Formed between carboxyl group of one amino acid and amino group of another.
Polypeptide: Chain of amino acids; proteins may consist of one or more polypeptides.
Protein Diversity and Structure
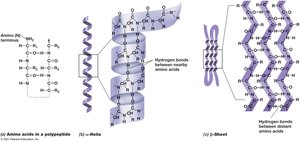
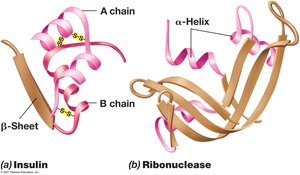
Proteins exhibit primary, secondary, tertiary, and quaternary structures, which determine their function and stability.
Primary Structure: Linear sequence of amino acids.
Secondary Structure: Alpha-helix or beta-sheet formed by hydrogen bonding.
Tertiary Structure: Three-dimensional folding due to hydrophobic interactions.
Quaternary Structure: Arrangement of multiple polypeptide subunits.
Denaturation: Loss of structure and function.
Transfer RNA (tRNA)
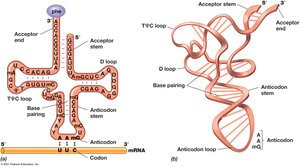
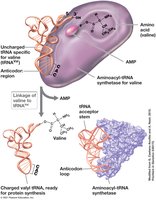
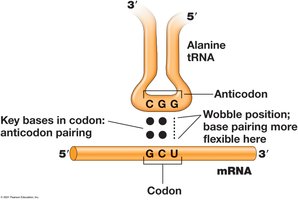
tRNAs carry amino acids to the ribosome during translation. Each tRNA contains an anticodon that recognizes the corresponding codon on mRNA.
Anticodon: Three bases that recognize mRNA codon.
Aminoacyl-tRNA Synthetase: Enzyme that attaches the correct amino acid to tRNA.
Cloverleaf Structure: Extensive secondary structure formed by internal base pairing.
Translation and the Genetic Code
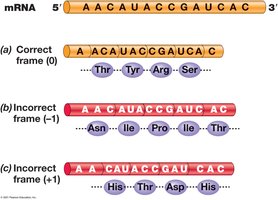
The genetic code is a triplet code, with 64 possible codons encoding 22 amino acids. Codon recognition occurs via base pairing with tRNA anticodons.
Degenerate Code: Multiple codons encode the same amino acid.
Wobble: Irregular base pairing at the third position allows flexibility.
Codon Bias: Unequal usage of synonymous codons.
Start Codon: AUG; encodes N-formylmethionine in Bacteria, methionine in Archaea/Eukarya.
Stop Codons: UAA, UAG, UGA; terminate translation.
Open Reading Frame (ORF): Sequence from start codon to stop codon.
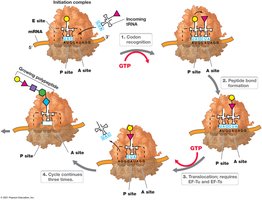
The Mechanism of Protein Synthesis
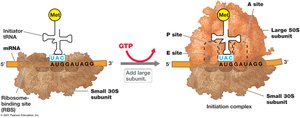
Protein synthesis involves initiation, elongation, and termination steps, requiring mRNA, tRNA, ribosomes, and various protein factors.
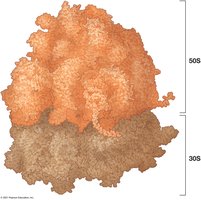
Ribosomes: Complexes of proteins and RNA; site of protein synthesis.
Initiation: Assembly of ribosomal subunits, mRNA, and initiator tRNA.
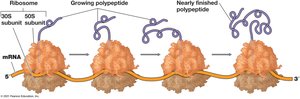
Elongation: Addition of amino acids to the growing polypeptide chain.
Termination: Release factors recognize stop codon and release polypeptide.
Polysomes: Multiple ribosomes translating a single mRNA.
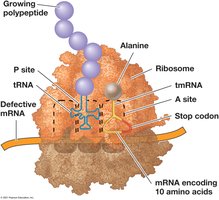
Trans-translation: tmRNA frees stalled ribosomes.
Protein Processing, Secretion, and Targeting
Assisted Protein Folding and Chaperones
Chaperones are proteins that assist in the folding of other proteins, refolding denatured proteins, and assembling protein complexes. They are found in all domains of life and are essential for proper protein function.
Chaperones: Catalyze folding events; include DnaK, DnaJ, GroEL, GroES in E. coli.
Heat Shock Proteins: Chaperones that respond to stress by refolding proteins.
Cold Shock Proteins: Prevent secondary structure formation in RNA.
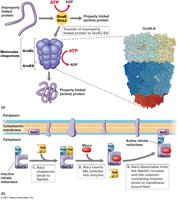
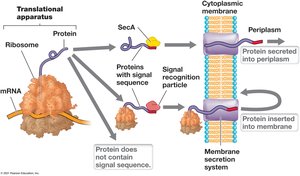
Protein Secretion: The Sec and Tat Systems
Some proteins must be transported outside the cytoplasmic membrane or inserted into membranes. The Sec and Tat systems are responsible for protein export in bacteria and archaea.
Sec System: Exports unfolded proteins; inserts integral membrane proteins.
Tat System: Transports folded proteins with cofactors.
Signal Sequence: N-terminal sequence directs proteins to translocase.
Protein Secretion: Gram-Negative Systems
Gram-negative bacteria possess complex secretion systems to transport proteins and effectors across membranes. These systems facilitate symbiosis, biofilm formation, enzyme secretion, DNA transfer, and more.
Secretion Systems: Large protein complexes forming channels through membranes.
Type I-VI Systems: Vary in mechanism; some require Sec/Tat, others are one-step translocases.
Type III: Injects toxins into eukaryotic host cells.
Type IV: Transfers DNA through conjugation.
Additional info: These notes expand upon the original content by providing definitions, examples, and context for each topic, ensuring completeness and academic quality for microbiology students.
