 Back
Backlec 20:Molecular Regulation in Microbial Systems: Mechanisms and Examples
Study Guide - Smart Notes

Molecular Regulation in Microbial Systems
Introduction to Molecular Regulation
Molecular regulation refers to the complex processes by which cells control gene expression and internal activities to maintain homeostasis and respond to environmental changes. This regulation involves the interplay of genes, proteins, and other molecules to ensure that biological functions occur at the appropriate time, location, and quantity.
Transcriptional Regulation
Transcription Factors: Activators and Repressors
Transcription factors are proteins that bind to specific DNA sequences to regulate gene expression. They function as molecular switches, either promoting or inhibiting the transcription of target genes by interacting with RNA polymerase.
Activators: Proteins that increase gene transcription by recruiting RNA polymerase to the promoter or by remodeling chromatin to make DNA more accessible.
Repressors: Proteins that decrease or block gene transcription by binding to operator or promoter regions, preventing RNA polymerase from initiating transcription.
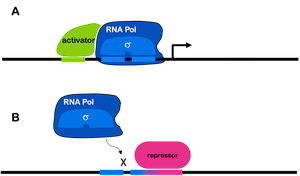
Mechanisms of Transcriptional Regulation
Recruitment: Activators bind to enhancer regions and facilitate the assembly of the transcriptional machinery.
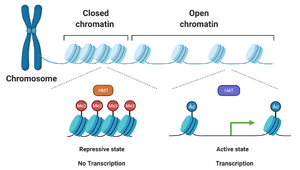
Chromatin Remodeling: Activators or repressors recruit enzymes that modify histones, altering chromatin structure and gene accessibility.
Competitive Binding: Repressors block RNA polymerase binding by occupying promoter or operator sites.
Quenching: Repressors bind to activators, preventing them from functioning.
Silencing: Repressors recruit enzymes that condense chromatin, making genes inaccessible.
Dynamic Regulation: The Rheostat Effect
Gene regulation is often not binary (on/off) but operates as a rheostat, with multiple activators and repressors fine-tuning gene expression levels. The amount of mRNA produced is determined by the balance of these regulatory signals.

Alternative Sigma Factors in Bacteria
Role of Sigma Factors
In bacteria, sigma (σ) factors are essential for directing RNA polymerase to specific promoter sequences. The primary sigma factor (σ70 in E. coli) controls housekeeping genes, while alternative sigma factors enable rapid shifts in gene expression in response to environmental changes.
σ32 (RpoH): Activates heat-shock genes during temperature stress.
σ38 (RpoS): Induces genes for survival during starvation or stationary phase.
Bacillus subtilis: Uses different sigma factors to regulate sporulation stages.
Anti-Sigma and Anti-Anti-Sigma Factors
Anti-sigma factors inhibit sigma factors by direct binding, preventing their association with RNA polymerase. Anti-anti-sigma factors can release sigma factors from inhibition in response to specific signals, adding another regulatory layer.
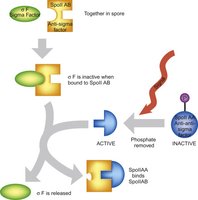

RNA-Based Regulation
Riboswitches
Riboswitches are regulatory segments within mRNA that bind small molecules, causing conformational changes that affect gene expression. They can terminate transcription prematurely or block translation initiation, allowing the mRNA to self-regulate based on metabolite concentrations.
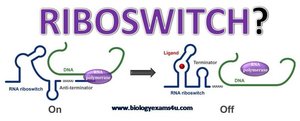
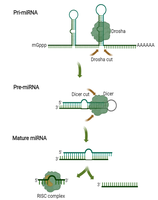
RNA Interference (RNAi)
In eukaryotes, small non-coding RNAs such as miRNAs and siRNAs silence gene expression by targeting mRNAs for degradation or translational repression. This system also serves as a defense against viral RNA.
miRNA: Endogenously encoded, binds complementary mRNA to degrade or inhibit translation.
siRNA: Often exogenous, guides mRNA cleavage and degradation.
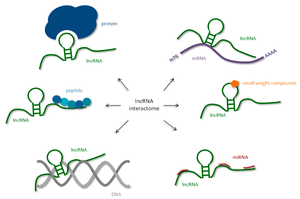
Long Non-coding RNAs (lncRNAs)
lncRNAs are longer than 200 nucleotides and regulate gene expression by recruiting chromatin-modifying proteins or acting as decoys for miRNAs. They play roles in chromatin remodeling and gene silencing, such as X-chromosome inactivation.
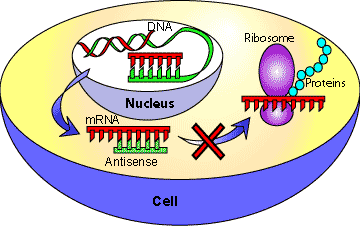
Antisense RNA
Antisense RNAs are complementary to specific mRNAs and can block translation or trigger mRNA degradation, effectively silencing gene expression.
CRISPR-Cas Systems
In bacteria and archaea, CRISPR-Cas systems provide adaptive immunity against viruses. Small CRISPR RNAs (crRNAs) guide Cas enzymes to recognize and cleave foreign DNA, preventing infection and providing a molecular memory of past invaders.
Second Messengers in Signal Transduction
Role and Mechanisms
Second messengers are intracellular molecules that relay signals from cell-surface receptors to internal targets, amplifying and distributing the signal to coordinate cellular responses. Examples include cyclic AMP (cAMP), calcium ions (Ca2+), and inositol triphosphate (IP3).
Signal amplification: One receptor can generate many second messenger molecules, amplifying the response.
Signal distribution: Second messengers activate multiple downstream targets.
Spatiotemporal control: Cells tightly regulate the timing and location of second messenger action.
Cellular Clocks, Thermometers, and Switches
Molecular Clocks
Molecular clocks are internal timing mechanisms based on transcription–translation negative feedback loops (TTFLs). These cycles regulate circadian rhythms and ensure biological processes occur at the correct time.
Feedback loop: A protein inhibits its own gene's expression, creating oscillatory cycles.
Example: The suprachiasmatic nucleus (SCN) in mammals coordinates circadian rhythms.
Molecular Thermometers
Molecular thermometers are RNA or protein structures that change conformation in response to temperature, modulating gene expression. RNA thermometers (RNATs) in bacteria control translation of heat-shock proteins by altering ribosome access to mRNA.
Molecular Switches
Molecular switches enable cells to make binary decisions, such as whether to divide or differentiate. These switches often rely on positive feedback or phosphorylation events.
Bistability: Positive feedback maintains the 'on' state even after the initial signal is gone.
Phospho-switches: Kinases add phosphate groups to activate proteins; phosphatases remove them to deactivate.
G-proteins: Act as on/off switches depending on whether they are bound to GTP (active) or GDP (inactive).
