 Back
BackMutations and Genetic Exchange in Bacteria: Study Notes
Study Guide - Smart Notes

Mutations in Bacterial Genomes
Definition and Importance
Mutations are heritable changes in the DNA sequence of an organism. In bacteria, mutations are a primary source of genetic variation and can affect cellular functions, adaptation, and evolution. They may arise spontaneously or be induced by environmental factors.
Types of Mutations
Mutations can be classified based on their nature and effect on the DNA sequence:
Point Mutation: A change in a single nucleotide base pair.
Transition: Substitution of a purine for another purine (A ↔ G) or a pyrimidine for another pyrimidine (C ↔ T).
Transversion: Substitution of a purine for a pyrimidine or vice versa (A or G ↔ C or T).
Insertion: Addition of one or more nucleotide base pairs.
Deletion: Removal of one or more nucleotide base pairs.
Inversion: A segment of DNA is reversed within the chromosome.
Reversion: A mutation that restores the original sequence or function.
Classes of Mutations by Information Effect
Silent Mutation: Alters a codon but does not change the encoded amino acid due to the redundancy of the genetic code (wobble effect).
Missense Mutation: Changes a codon to encode a different amino acid, potentially altering protein function.
Nonsense Mutation: Converts a codon to a stop codon, resulting in premature termination of translation.
Frame-shift Mutation: Insertion or deletion of nucleotides that alters the reading frame, often leading to extensive changes in the protein sequence.
Examples of Mutation Effects
Mutations can have diverse effects on protein synthesis and function. The following examples illustrate key mutation types:
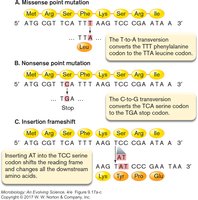
Missense Mutation: A single base change results in a different amino acid (e.g., TTT to TTA changes phenylalanine to leucine).
Nonsense Mutation: A base change creates a stop codon, truncating the protein (e.g., TCG to TGA).
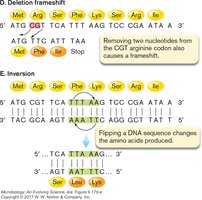
Insertion Frame-shift: Addition of bases shifts the reading frame, altering downstream amino acids.
Deletion Frame-shift: Removal of bases shifts the reading frame, often resulting in a nonfunctional protein.
Inversion: Reversal of a DNA segment changes the amino acid sequence produced.

Genetic Exchange in Bacteria
Horizontal Gene Transfer
Bacteria can acquire new genetic material through horizontal gene transfer (HGT), which is distinct from vertical inheritance. HGT mechanisms include transformation, transduction, and conjugation. These processes contribute to genetic diversity and adaptation, such as the spread of antibiotic resistance.
Conjugation: Mechanism and Significance
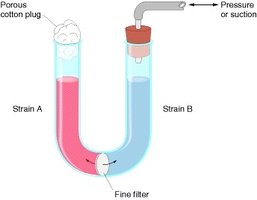
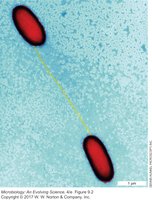
Conjugation is a process where DNA is transferred from one bacterium (donor) to another (recipient) via direct cell-to-cell contact. It is often referred to as "bacterial sex" and is a major mechanism for the spread of plasmids, including those conferring antibiotic resistance.
Requires physical contact between cells, typically mediated by a conjugation pilus.
Transfer is unidirectional: donor to recipient.
Facilitates exchange of plasmids and, occasionally, chromosomal genes.
Important in bacterial evolution and adaptation.
Fertility Factor (F Factor) and Surface Exclusion
The Fertility factor (F factor) is a plasmid that enables conjugation in Escherichia coli. Cells with the F factor (F+) can donate DNA to F– cells. Surface exclusion prevents F+ cells from conjugating with other F+ cells.
F+ × F–: Fertile, DNA transfer occurs.
F+ × F+: Much less fertile due to surface exclusion.
F– × F–: Infertile, no DNA transfer.
Mechanism of Conjugation
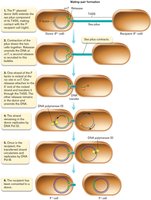
Conjugation involves the transfer of plasmid DNA from the donor to the recipient. The F factor contains two replication origins:
oriV: Used in nonconjugating cells.
oriT: Used during DNA transfer.
The process begins with pilus formation, followed by DNA transfer and replication in both cells.
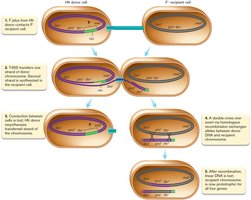
Transfer of Chromosomal Genes: HFR Strains
High-Frequency Recombination (HFR) Strains
HFR strains are bacteria in which the F factor is integrated into the chromosome. During conjugation, chromosomal genes adjacent to the F factor can be transferred to the recipient cell, leading to recombination and genetic diversity.
Transfer begins at the origin of transfer (oriT) and proceeds linearly.
Only part of the chromosome is usually transferred before the cells separate.
Transferred genes can recombine with the recipient's chromosome.
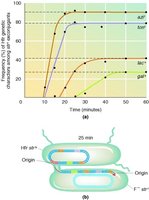
Mapping Bacterial Genes Using HFR Conjugation
Gene mapping can be performed by measuring the time required for different genes to transfer during conjugation. The order and timing of gene transfer reflect their positions on the chromosome.
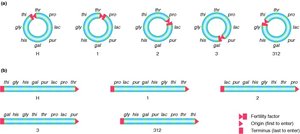
HFR Strain Genotypes and Chromosomal Maps
Different HFR strains have unique integration sites and transfer origins, resulting in distinct gene transfer patterns. The following table summarizes the genotypes of several HFR strains:
Hfr Strain | Transferred Genes |
|---|---|
H | O thr pro lac pur gal his gly thi F |
1 | O thr thi gly his gal pur lac pro F |
2 | O pro thr thi gly his gal pur lac F |
3 | O pur lac pro thr thi gly his gal F |
AB 312 | O thi thr pro lac pur gal his gly F |

The following diagram shows the chromosomal maps and transfer origins for these strains:
Summary Table: Mutation Types and Effects
Mutation Type | Definition | Effect |
|---|---|---|
Silent | Base change, same amino acid | No effect on protein |
Missense | Base change, different amino acid | Altered protein function |
Nonsense | Base change, stop codon | Truncated protein |
Frame-shift | Insertion/deletion, reading frame shift | Extensive protein changes |
Inversion | DNA segment reversed | Changed amino acid sequence |
Key Equations and Concepts
Central Dogma:
Mutation Rate:
Gene Transfer Frequency:
Additional info: Academic context was added to clarify mutation types, conjugation mechanisms, and HFR mapping, ensuring completeness and self-contained explanations for exam preparation.
