 Back
BackProtein Export and Regulation of Gene Expression in Bacteria
Study Guide - Smart Notes

Protein Export in Bacteria
Overview of Bacterial Protein Localization
Bacterial proteins are distributed throughout various cellular compartments, each serving distinct functions. Understanding protein localization is essential for grasping bacterial physiology and pathogenesis.
Cytoplasmic proteins: Enzymes for metabolism (e.g., pyruvate kinase, phosphofructokinase), ribosomal proteins, chaperones (e.g., GroEL), anabolic and catabolic proteins.
Cytoplasmic membrane proteins: Components of the flagellar motor, ATP synthase, porins.
Periplasmic proteins: Proteins involved in cell wall synthesis and transport.
Exported proteins: Toxins, flagella and pili proteins, coagulase.
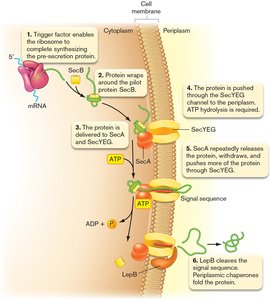
SecA-Dependent General Secretion Pathway
The SecA-dependent pathway is the primary route for exporting proteins from the cytoplasm to the periplasm in Gram-negative bacteria. This process involves several key steps and molecular components.
Step 1: The peptide is fully synthesized in the cytoplasm.
Step 2: The pre-secretion protein is captured by the piloting protein SecB.
Step 3: SecB unfolds and delivers the protein to SecA, which is associated with the SecYEG translocon.
Step 4: SecA uses ATP hydrolysis to push the protein through the SecYEG channel into the periplasm.
Step 5: LepB cleaves the signal sequence, and periplasmic chaperones fold the protein.
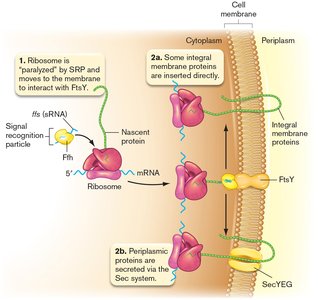
SRP and Cotranslational Export in E. coli
Some integral membrane proteins are inserted directly into the membrane during translation, mediated by the signal recognition particle (SRP) system. This process ensures proper localization and folding of membrane proteins.
SRP: Recognizes signal sequences on nascent proteins and directs ribosomes to the membrane.
Integral membrane proteins: Inserted directly into the membrane via the FtsY receptor and SecYEG translocon.
Periplasmic proteins: Exported via the Sec system after complete synthesis.
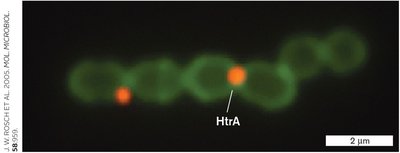
Protein Export in Gram-Positive Bacteria
Gram-positive bacteria also utilize specialized pathways for protein export, often involving similar translocon systems but adapted to their unique cell envelope structure.
Journeys to the Outer Membrane
Gram-negative bacteria possess multiple secretion systems (Type I–VII) to export proteins beyond the periplasm, including digestive enzymes and toxins. These systems vary in complexity and specificity.
Type I–VII secretion systems: Specialized for different substrates and destinations.
Some systems: Deliver proteins to periplasmic transporters; others provide direct export to the extracellular space.
Regulation of Gene Expression in Bacteria
Levels of Gene Expression Control
Bacteria regulate gene expression at multiple levels to adapt to changing environments and conserve resources.
Alteration of DNA sequence: Mutations or rearrangements affecting gene function.
Transcriptional control: Regulation of mRNA synthesis via repressors, activators, and sigma factors.
Control of mRNA stability: Degradation rates influence protein production.
Translational control: Regulation of protein synthesis from mRNA.
Posttranslational control: Modification or degradation of proteins after synthesis.
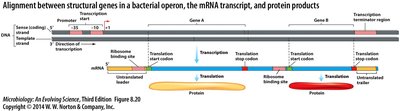
Operons and Transcriptional Regulation
Operons are clusters of genes transcribed together, allowing coordinated regulation. The classic example is the lacZYA operon, which is induced by lactose.
Promoter: Site where RNA polymerase binds to initiate transcription.
Operator: DNA sequence where repressors or activators bind.
Structural genes: Encode proteins with related functions.
Transcription Repressors and Activators
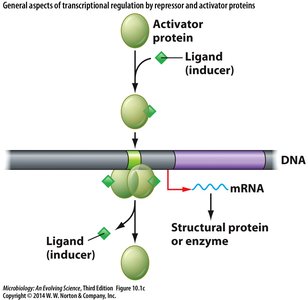
Transcriptional control is mediated by regulatory proteins that bind DNA and influence RNA polymerase activity.
Repressors: Prevent transcription by binding to operators.
Activators: Stimulate transcription by enhancing RNA polymerase binding.
Inducers and corepressors: Small molecules that modulate repressor/activator activity.
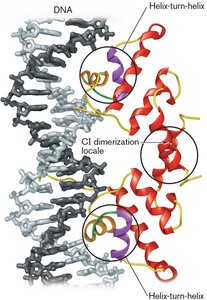
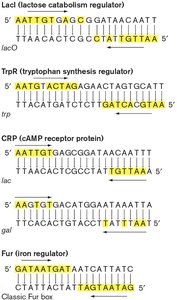
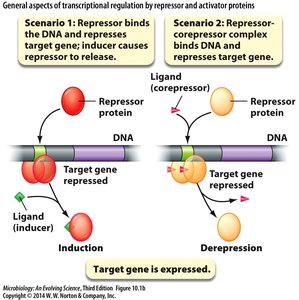
General Concepts of Transcriptional Control
Regulatory proteins interact with specific DNA sequences to control gene expression in response to intracellular and extracellular signals.
Regulatory protein: Encoded by a gene, binds to regulatory sequence.
Ligand: Inducer or corepressor that affects regulatory protein function.
Target gene: Structural gene or enzyme whose expression is regulated.
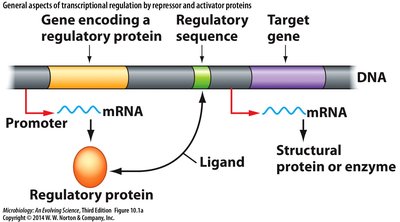
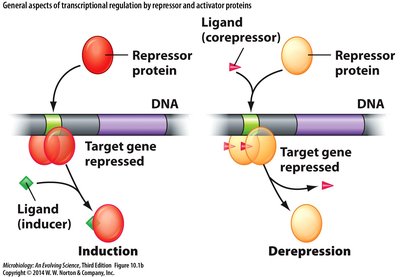
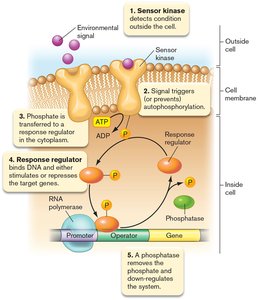
Sensing the Extracellular Environment
Bacteria use two-component systems to sense and respond to environmental changes. These systems consist of a sensor kinase and a response regulator.
Sensor kinase: Detects external signals and autophosphorylates.
Response regulator: Receives phosphate and regulates target gene expression.
Phosphatase: Removes phosphate to reset the system.
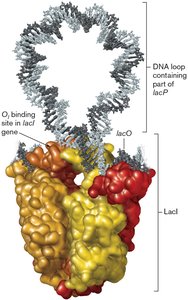
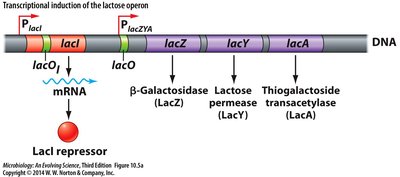
Lactose Operon (lacZYA)
The lacZYA operon is a model for inducible gene expression. It is regulated by the LacI repressor and is induced in the presence of lactose.
LacI repressor: Binds to the operator to block transcription in the absence of lactose.
Lactose (inducer): Binds LacI, causing it to release the operator and allowing transcription.
Gene products: β-Galactosidase (LacZ), lactose permease (LacY), thiogalactoside transacetylase (LacA).
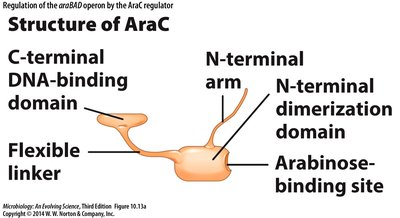
Arabinose Operon (araBAD)
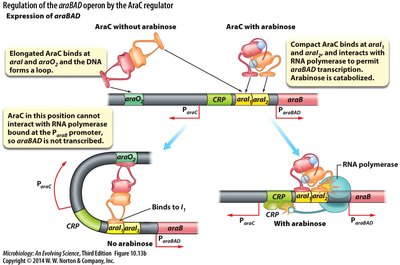
The araBAD operon is regulated by the AraC protein, which acts as both a repressor and activator depending on the presence of arabinose.
AraC: Changes conformation upon binding arabinose, allowing transcription.
Without arabinose: AraC represses transcription by looping DNA.
With arabinose: AraC activates transcription by facilitating RNA polymerase binding.
Repression of Anabolic Pathways
Anabolic pathways are often repressed when their end products are abundant, conserving energy and resources.
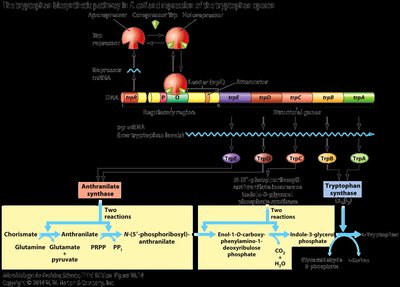
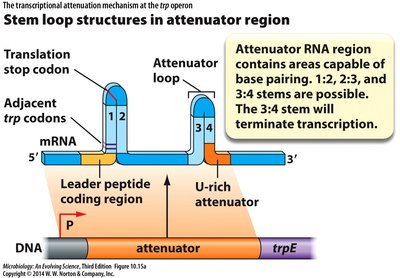
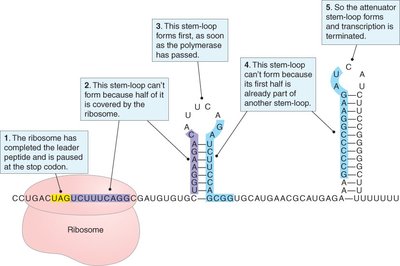
Attenuation of the trp Operon
The trp operon uses attenuation to fine-tune expression based on tryptophan levels. This mechanism involves the formation of stem-loop structures in the mRNA leader region.
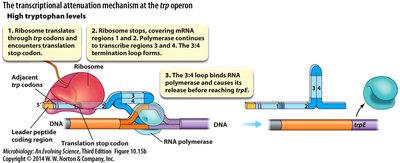
High tryptophan: Ribosome quickly translates leader peptide, allowing formation of terminator stem-loop (3:4), halting transcription.
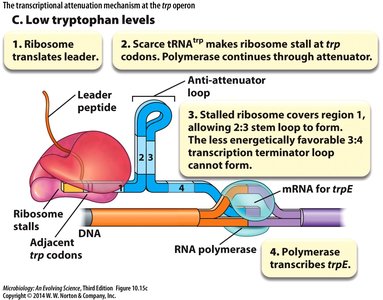
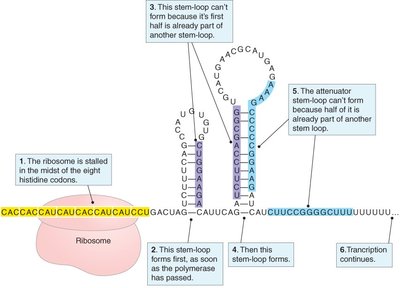
Low tryptophan: Ribosome stalls, allowing anti-terminator stem-loop (2:3) to form, permitting transcription.
Sigma Factors and Regulatory RNAs
Sigma Factors
Sigma factors are proteins that direct RNA polymerase to specific promoters, enabling coordinated expression of gene sets (regulons).
Regulon: Group of genes controlled by a single regulatory protein.
Alternative sigma factors: Allow bacteria to respond to environmental changes by switching gene expression programs.
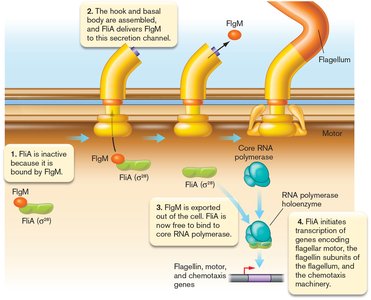
Regulation by Anti-Sigma Factors
Anti-sigma factors inhibit sigma factor activity, while anti-anti-sigma factors can relieve this inhibition, providing additional layers of control.
Summary Table: Protein Localization in Bacteria
Location | Example Proteins | Function |
|---|---|---|
Cytoplasm | Pyruvate kinase, GroEL | Metabolism, protein folding |
Cytoplasmic Membrane | Flagellar motor, ATP synthase | Motility, energy production |
Periplasm | Cell wall synthesis proteins | Cell wall assembly, transport |
Exported/Extracellular | Toxins, pili proteins | Pathogenesis, adhesion |
Additional info: These notes expand on the original lecture content with definitions, examples, and mechanisms to provide a comprehensive study guide for microbiology students.
