 Back
BackRegulation of Bacterial Cellular Processes: Mechanisms and Examples
Study Guide - Smart Notes

Regulation of Bacterial Cellular Processes
Common Regulatory Mechanisms in Bacteria
Bacteria employ a variety of mechanisms to regulate cellular processes, ensuring efficient adaptation to environmental changes. These mechanisms control gene expression at multiple levels, including transcription initiation, transcription elongation, translation, and posttranslational modification. The three domains of life (Bacteria, Archaea, Eukarya) differ in their genome structure and regulatory strategies.
Regulation of gene expression: Modulation of when and how genes are transcribed and translated.
Transcription initiation: Control of the start of mRNA synthesis.
Transcription elongation: Regulation of the continuation of mRNA synthesis.
Translation: Control of protein synthesis from mRNA.
Posttranslational regulation: Modification of proteins after synthesis to alter activity.
Regulation of Transcription Initiation
Transcription initiation is a critical control point for gene expression. Genes can be classified based on their regulation:
Constitutive genes: Housekeeping genes expressed continuously to maintain basic cellular functions.
Inducible genes: Genes expressed only under specific conditions, such as the presence of certain substrates (e.g., β-Galactosidase).
Repressible genes: Genes involved in biosynthetic pathways, typically expressed unless the end product is available.
Positive Control: Activation
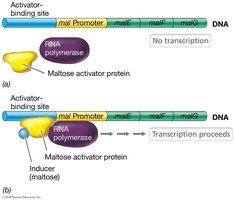
Positive control involves activator proteins that enhance the binding of RNA polymerase to DNA, promoting transcription. A classic example is maltose catabolism in Escherichia coli, where the maltose activator protein must bind maltose (the inducer) before it can bind DNA and activate transcription.
Activator proteins: Bind to specific DNA sequences (activator-binding sites) to facilitate transcription.
Inducer: A molecule (e.g., maltose) that enables the activator protein to bind DNA.
Example: Maltose activator protein in E. coli requires maltose to bind DNA and activate transcription of maltose catabolism genes.
Positive Control: DNA Looping
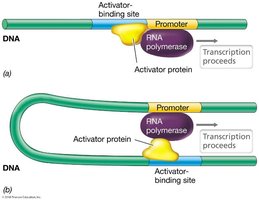
Activator proteins can also function by DNA looping, bringing distant regulatory sites into proximity with the promoter to facilitate transcription.
DNA looping: Allows activator proteins bound at distant sites to interact with RNA polymerase at the promoter.
Example: DNA looping enables activator proteins to regulate transcription even when their binding sites are far from the promoter.
Negative Control: Repression and Induction
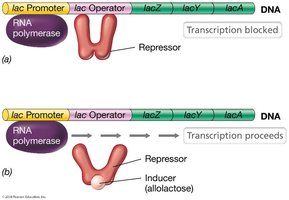
Negative control involves repressor proteins that inhibit transcription by binding to operator sequences. Operons are clusters of genes regulated together by a single operator.
Operon: A group of genes transcribed as a single mRNA, regulated by a common operator.
Repressor: Protein that binds to the operator, blocking RNA polymerase and preventing transcription.
Inducer: Molecule that inactivates the repressor, allowing transcription to proceed.
Example: The lac operon in E. coli is regulated by a repressor that blocks transcription unless lactose (inducer) is present.
Global Control and the lac Operon
Global regulatory mechanisms allow bacteria to prioritize energy sources. The lac operon is regulated by catabolite activator protein (CAP), which responds to glucose availability, enabling preferential use of glucose over lactose.
Diauxic growth: Two-phase growth when two energy sources are available; the preferred source is used first.
CAP: Activates transcription of the lac operon when glucose is absent.
Regulation of Transcription Elongation
Transcription elongation can be regulated by mechanisms such as attenuation and riboswitches. Attenuation involves premature termination of transcription, as seen in the trp operon.
Attenuation: Termination of transcription within the leader region, controlled by mRNA secondary structures.
Riboswitches: mRNA elements that change folding in response to effector molecules, affecting transcription or translation.
Example: The trp operon uses attenuation to regulate transcription based on tryptophan levels.
Regulation of Translation
Translation can be regulated by riboswitches and small RNA molecules. In Gram-negative bacteria, riboswitches at the 5’ end of mRNA alter folding patterns to control translation initiation.
Small RNAs: Can bind mRNA and affect translation initiation.
Quorum Sensing
Quorum sensing is a cell-to-cell communication mechanism mediated by small signaling molecules such as N-acyl-homoserine lactone (AHL). It couples cell density to gene regulation, playing a key role in virulence, symbiosis, biofilm formation, and differentiation.
AHL: Signaling molecule used in quorum sensing.
LuxR: Transcriptional regulator activated by AHL in Vibrio fischeri.
Example: High cell density in V. fischeri leads to AHL accumulation, activation of LuxR, and bioluminescence gene expression.
Preserving Genome Integrity: CRISPR Interference
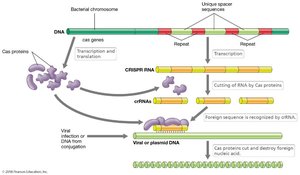
CRISPR (Clustered Regulatory Interspaced Short Palindromic Repeats) is a prokaryotic immune system that protects against viral infection and maintains genome stability. It consists of repeated DNA sequences interspaced with segments of foreign DNA (spacers). Cas proteins process CRISPR RNAs (crRNAs) that guide the destruction of invading nucleic acids.
CRISPR mechanism: Transcription of CRISPR array produces long RNA, which is cleaved by Cas proteins to generate crRNAs.
crRNAs: Base-pair with invading nucleic acids, leading to their destruction.
Distribution: CRISPR systems are found in ~90% of Archaea and 70% of Bacteria.
Example: CRISPR-Cas system provides adaptive immunity against bacteriophages and plasmids.
Regulatory Mechanism | Key Proteins/Elements | Example |
|---|---|---|
Positive Control | Activator protein, Inducer | Maltose activator protein in E. coli |
Negative Control | Repressor, Operator, Inducer | Lac operon in E. coli |
Attenuation | Leader peptide, Stem-loop structures | trp operon |
Riboswitches | mRNA leader sequence, Effector molecule | Transcription/translation regulation |
Quorum Sensing | AHL, LuxR | Bioluminescence in V. fischeri |
CRISPR Interference | CRISPR array, Cas proteins, crRNAs | Immunity against phages |
Additional info: Expanded explanations and examples were added for clarity and completeness. The table summarizes key regulatory mechanisms, proteins, and examples for quick review.
