 Back
BackRegulation of Transcription in Microorganisms: Mechanisms and Examples
Study Guide - Smart Notes

Regulation of Transcription in Microorganisms
Overview of Transcriptional Regulation
Transcriptional regulation is a fundamental process in bacteria, archaea, and eukarya, controlling gene expression in response to environmental and cellular signals. While bacteria and archaea share similar regulatory mechanisms, eukarya possess additional layers of control. Key regulatory elements include repressor and activator proteins, two-component regulatory systems, and anti-sigma factor interactions.
Repressor Proteins: Bind to operator regions to block transcription.
Activator Proteins: Enhance transcription by binding to activator sites.
Two-Component Systems: Involve sensor kinases and response regulators for environmental sensing.
Anti-Sigma Factor Interactions: Modulate sigma factor activity, affecting RNA polymerase binding.
Negative Control of Transcription
Enzyme Repression: The arg Operon
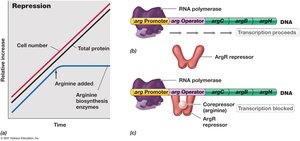
Enzyme repression occurs when the end product of a biosynthetic pathway accumulates, inhibiting further synthesis of enzymes involved in that pathway. The arg operon is a classic example, regulated by the presence of arginine.
Mechanism: When arginine is absent, the repressor protein does not bind to the operator, allowing transcription to proceed.
In the presence of arginine: Arginine binds to the repressor, enabling it to bind to the operator and block transcription.
Result: Synthesis of arginine biosynthetic enzymes is halted when arginine is abundant.
Example: Regulation of the arg operon in Escherichia coli.
Enzyme Induction: The lac Operon
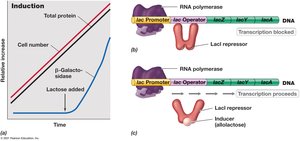
Enzyme induction is triggered by the presence of a substrate, leading to the synthesis of enzymes required for its metabolism. The lac operon is induced by lactose.
Mechanism: In the absence of lactose, the LacI repressor binds to the operator, blocking transcription.
In the presence of lactose: Lactose (or allolactose) binds to the repressor, causing its release from the operator and allowing transcription.
Result: Synthesis of β-galactosidase and other enzymes for lactose metabolism.
Example: Regulation of the lac operon in Escherichia coli.
Positive Control of Transcription
Activation of Transcription: The mal Operon
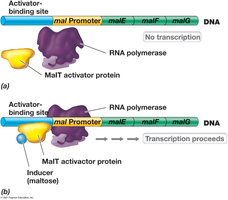
Positive control involves activator proteins that enhance transcription when an inducer is present. The mal operon is activated by maltose.
Mechanism: In the absence of maltose, the MalT activator protein does not bind to the activator site, and transcription does not occur.
In the presence of maltose: Maltose binds to MalT, enabling it to bind to the activator site and activate transcription.
Result: Synthesis of enzymes for maltose metabolism.
Example: Regulation of the mal operon in Escherichia coli.
Regulons: Coordinated Control of Multiple Operons
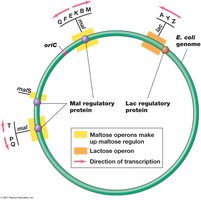
A regulon is a group of operons controlled by a single regulatory protein, allowing coordinated expression of functionally related genes.
Example: The maltose regulon in Escherichia coli includes several operons regulated by MalT.
Catabolite Repression and Diauxic Growth
Catabolite Repression: The Glucose Effect
Catabolite repression is a global regulatory mechanism where the presence of a preferred carbon source (e.g., glucose) inhibits the expression of genes for the metabolism of other substrates.
Mechanism: Glucose inhibits cAMP synthesis, preventing cAMP-CRP complex formation and binding to the CRP site, thus blocking transcription of the lac operon.
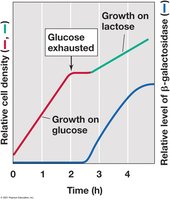
Diauxic Growth: Cells first consume glucose, then switch to lactose after glucose is exhausted, resulting in two distinct growth phases.
Transcriptional Regulation in Archaea
Dual Functionality of TrmBL1 Regulator
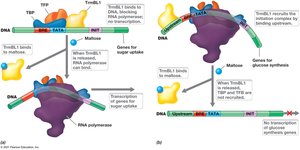
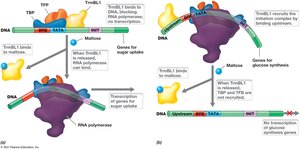
Archaea utilize unique regulatory proteins such as TrmBL1, which can act as both a repressor and an activator depending on the presence of specific sugars.
Repression: TrmBL1 represses maltose uptake genes in the absence of maltose.
Activation: TrmBL1 activates genes for glucose synthesis in the absence of maltose.
Release: Presence of maltose releases TrmBL1, allowing transcription of maltose uptake genes.
Repression of Nitrogen Metabolism Genes by NrpR
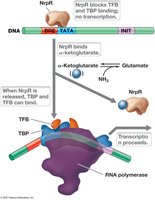
NrpR is a repressor protein in archaea that regulates genes involved in nitrogen metabolism.
Mechanism: NrpR represses transcription in the presence of ammonia.
Release: Accumulation of α-ketoglutarate binds to NrpR, releasing it and allowing transcription of nitrate assimilation genes.
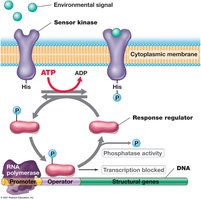
Two-Component Regulatory Systems
Mechanism of Two-Component Systems
Two-component systems are widespread in bacteria and archaea, enabling cells to sense and respond to environmental changes.
Components: Sensor kinase (detects signal) and response regulator (modulates gene expression).
Process: Sensor kinase autophosphorylates in response to a signal, then transfers the phosphate to the response regulator, which activates or represses transcription.
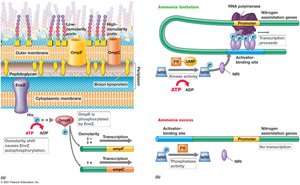
Example: Regulation of nitrogen assimilation proteins in Escherichia coli.
Global Control via Alarmones: The Stringent Response
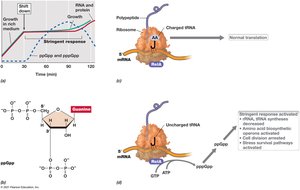
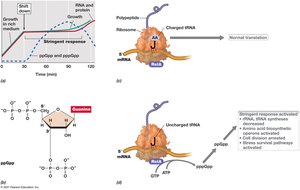
The Stringent Response in Escherichia coli
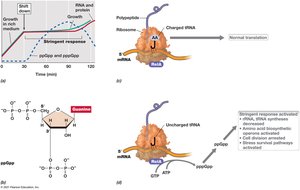
The stringent response is a global regulatory mechanism activated during amino acid starvation or other stresses, mediated by alarmones such as ppGpp.
Mechanism: ppGpp binds to RNA polymerase, inhibiting rRNA and tRNA synthesis.
Effect: Ribosome activity decreases, cell division is arrested, and stress survival pathways are activated.
Example: Shift from amino acid excess to limitation triggers the stringent response in Escherichia coli.
Summary Table: Types of Transcriptional Regulation
Type | Mechanism | Example |
|---|---|---|
Negative Control | Repressor binds operator to block transcription | arg operon, lac operon |
Positive Control | Activator binds activator site to enhance transcription | mal operon |
Catabolite Repression | Preferred substrate inhibits expression of other metabolic genes | lac operon (glucose effect) |
Two-Component System | Sensor kinase and response regulator modulate gene expression | Nitrogen assimilation in E. coli |
Stringent Response | Alarmones inhibit rRNA/tRNA synthesis under stress | ppGpp in E. coli |
Additional info: These mechanisms are central to microbial regulatory systems, as covered in Chapter 7 of a typical microbiology curriculum. Understanding these systems is essential for grasping microbial adaptation, metabolism, and pathogenesis.
