 Back
BackRNA-Based Regulation and Enzyme Activity Modification in Microbial Regulatory Systems
Study Guide - Smart Notes

RNA-Based Regulation of Gene Expression
Noncoding RNAs (ncRNAs)
Noncoding RNAs are RNA molecules that are not translated into proteins but play essential roles in cellular regulation. They include rRNAs, tRNAs, and other RNAs involved in processes such as protein secretion and mRNA splicing. Among these, small RNAs (sRNAs) and microRNAs (miRNAs) are particularly important for gene regulation.
Small RNAs (sRNAs): Typically 40–400 nucleotides long; regulate gene expression in both prokaryotes and eukaryotes.
Functions: Modulate cellular processes like oxidative stress, iron stress, quorum sensing, biofilm formation, and stationary phase events.
Mechanisms of sRNA Regulation
sRNAs exert regulatory effects by base pairing with complementary sequences in mRNAs, affecting translation and mRNA stability.
Translation Inhibition: sRNA blocks the ribosome-binding site (RBS), preventing translation.
Translation Stimulation: sRNA opens a blocked RBS, allowing translation.
RNA Degradation: sRNA binding can recruit ribonucleases, leading to mRNA degradation.
RNA Protection: sRNA can block ribonuclease binding sites, protecting mRNA from degradation.
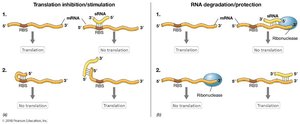
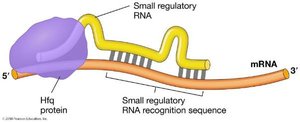
Role of Hfq Protein
The Hfq protein acts as an RNA chaperone, facilitating the binding of sRNAs to their target mRNAs and protecting sRNAs from degradation. sRNAs that are spatially separated from their target mRNAs are called trans-sRNAs.
Riboswitches
Definition and Mechanism
Riboswitches are cis-acting regulatory elements usually located in the 5’ untranslated regions (UTRs) of mRNAs. They regulate gene expression in response to the concentration of specific small molecules, typically at the level of translation.
The 5’ UTR folds into a secondary structure that binds a small metabolite.
Binding causes a conformational change, altering downstream mRNA base-pairing and inhibiting translation initiation.
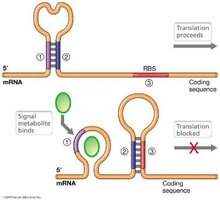
Metabolite Range and Biological Significance
Riboswitches respond to a variety of metabolites, including vitamins, amino acids, and nucleotides. The metabolite bound is often the product of the biosynthetic pathway encoded by the mRNA.
If metabolite levels are high, riboswitch binding prevents further synthesis.
Riboswitches are found in some bacteria, plants, and fungi, and may be remnants of the ancient RNA world.
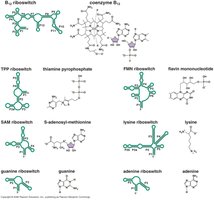
Attenuation
Mechanism and Biological Context
Attenuation is a transcriptional control mechanism that causes premature termination of mRNA synthesis. It is found in prokaryotes (Bacteria and Archaea) but not in eukaryotes due to spatial separation of transcription and translation.
The leader sequence in mRNA can fold into two alternative secondary structures:
One allows continued transcription.
The other causes premature termination.
Attenuation in the Tryptophan Operon
The tryptophan operon contains structural genes for five proteins, a promoter, and regulatory sequences. The leader sequence encodes a short peptide with tandem tryptophan codons.
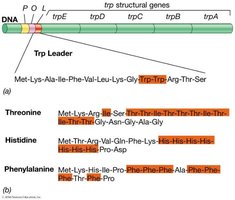
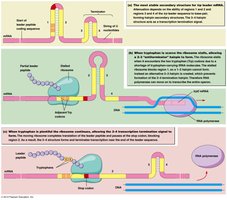
When tryptophan is abundant, ribosomes quickly translate the leader peptide, leading to transcription termination.
When tryptophan is scarce, ribosomes pause, allowing transcription to proceed.
Regulation of Enzyme Activity
Feedback Inhibition
Feedback inhibition is a mechanism where the end product of a biosynthetic pathway inhibits an early enzyme, temporarily shutting off the pathway. This inhibition is reversible.
The inhibited enzyme has two binding sites:
Active site: Binds substrate.
Allosteric site: Binds product (allosteric effector).
Excess product binds the allosteric site, causing a conformational change that prevents substrate binding.
When product concentration decreases, inhibition is lifted and the pathway resumes.
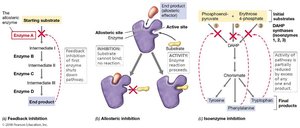
Isoenzymes
Some pathways use isoenzymes, which are different enzymes catalyzing the same reaction but subject to different regulatory controls. This prevents the excess of one or two products from shutting down the entire pathway.
Post-Translational Modifications
Biosynthetic enzymes can be regulated by covalent modifications, such as phosphorylation, methylation, adenylylation, and uridylylation. These modifications change the enzyme’s conformation and activity.
Common modifiers: cyclic AMP, ADP, inorganic phosphate, methyl groups.
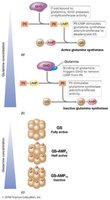
Regulation of PII signal transduction proteins affects nitrogen metabolism in Bacteria, Archaea, and plastids.
Uridylylation of PII stimulates glutamine synthetase adenyltransferase, increasing ammonia assimilation.
Regulation of Sigma Factor Activity
Inactivation of Sigma Factors
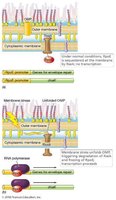
Anti-sigma factors can bind and inactivate sigma factors, a form of post-translational regulation. For example, RpoE (a sigma factor) promotes expression of proteins for outer membrane maintenance. When the membrane is not stressed, RseA (anti-sigma factor) binds RpoE, preventing expression. Under stress, RseA is degraded, allowing RpoE to initiate transcription.
Summary Table: RNA-Based Regulation and Enzyme Activity Modification
Regulatory Mechanism | Key Features | Example |
|---|---|---|
sRNAs | Base pairing with mRNA; translation inhibition/stimulation; mRNA degradation/protection | Hfq-assisted sRNA regulation |
Riboswitches | 5’ UTR secondary structure; metabolite binding; translation control | Thiamine pyrophosphate riboswitch |
Attenuation | Leader sequence; alternative mRNA structures; premature transcription termination | Tryptophan operon |
Feedback Inhibition | End product inhibits early enzyme; reversible; allosteric regulation | Amino acid biosynthesis |
Post-Translational Modification | Covalent modification; changes enzyme activity | Phosphorylation of glutamine synthetase |
Sigma Factor Inhibition | Anti-sigma factor binding; stress response regulation | RpoE/RseA system |
Additional info: The notes expand on brief lecture points to provide academic context, definitions, and examples for each regulatory mechanism. All images included are directly relevant to the adjacent explanations, visually reinforcing key concepts.
