 Back
BackTranscription: Mechanisms and Regulation in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

Transcription: The First Step of Gene Expression
Overview of the Central Dogma
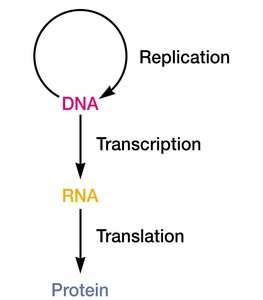
Transcription is the process by which genetic information encoded in DNA is copied into RNA. This is the first step in gene expression, followed by translation, where RNA is used to synthesize proteins. The central dogma of molecular biology describes the flow of genetic information from DNA to RNA to protein.
Steps in Transcription
Transcription occurs in three main stages: initiation, elongation, and termination. Each stage is tightly regulated and involves specific molecular machinery.
Initiation: RNA polymerase binds to the promoter region of DNA, aided by transcription factors or sigma factors, and begins RNA synthesis.
Elongation: RNA polymerase moves along the DNA template, synthesizing RNA in the 5' to 3' direction.
Termination: Transcription ends when RNA polymerase encounters a terminator sequence, releasing the newly synthesized RNA.
Transcription Units and Operons
Monocistronic vs. Polycistronic Transcripts
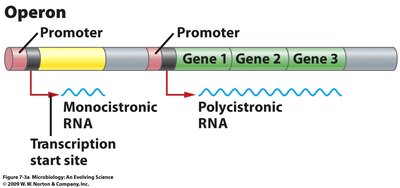
A transcription unit is a segment of DNA transcribed into a single RNA molecule. In prokaryotes, genes can be organized into operons, allowing multiple genes to be transcribed together as a single polycistronic mRNA. In contrast, eukaryotic genes are typically monocistronic, with each gene transcribed separately.
Operon: A cluster of genes under the control of a single promoter, transcribed as one mRNA.
Monocistronic RNA: mRNA that encodes a single protein.
Polycistronic RNA: mRNA that encodes multiple proteins, common in prokaryotes.
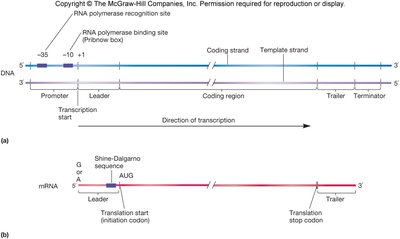
Template and Coding Strands
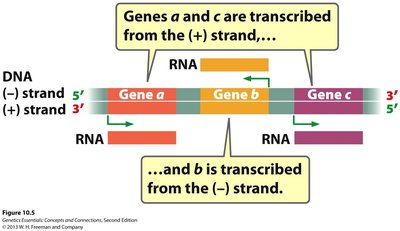
For any given gene, only one of the two DNA strands serves as the template for RNA synthesis. The template strand is read by RNA polymerase to produce a complementary RNA molecule. The other strand, called the coding (or sense) strand, has the same sequence as the RNA (except T is replaced by U).
Both DNA strands can serve as templates, but for different genes.
Directionality is important: RNA is always synthesized 5' to 3'.
RNA Polymerase: Structure and Function
Prokaryotic vs. Eukaryotic RNA Polymerases
RNA polymerase (RNAP) is the enzyme responsible for synthesizing RNA from a DNA template. Prokaryotes have a single RNA polymerase that transcribes all gene types, while eukaryotes have three main RNA polymerases:
RNA Pol I: Transcribes rRNA genes (non-coding RNAs).
RNA Pol II: Transcribes protein-coding genes (mRNA).
RNA Pol III: Transcribes tRNA and other small RNAs (non-coding).
The prokaryotic RNA polymerase core enzyme (α2ββ'ω) can elongate RNA but requires a sigma (σ) factor to initiate transcription at specific promoters. The holoenzyme is the core enzyme plus sigma factor.
Transcription Initiation in Prokaryotes
Promoters and Sigma Factors
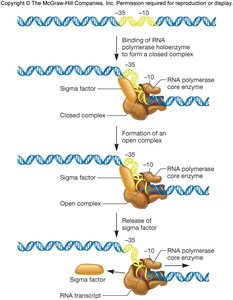
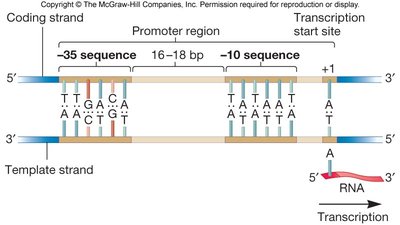
Initiation is highly regulated and involves the recognition of promoter sequences by sigma factors. The promoter contains conserved regions, typically at -35 and -10 positions relative to the transcription start site (+1).
Sigma factors: Direct RNA polymerase to specific promoters. E. coli has a housekeeping sigma factor (σ70) and several specialized sigma factors for stress responses.
Consensus sequence: The most common nucleotide sequence found at each position in a set of promoters. Real promoters often deviate from the consensus, affecting binding strength and gene expression levels.
Transcription Elongation
Mechanism of RNA Synthesis
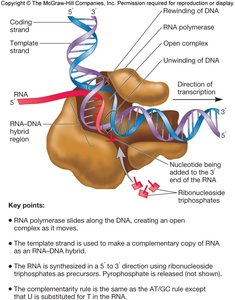
During elongation, the core RNA polymerase adds ribonucleotides (NTPs) to the 3' end of the growing RNA chain. The enzyme moves along the DNA, unwinding it and synthesizing RNA complementary to the template strand. The process is rapid and highly processive, with an error rate of about 1 in 104 nucleotides, which is tolerable for RNA.
Direction: RNA is synthesized 5' to 3'.
Template: Only one DNA strand is used for each gene.
Accuracy: Lower than DNA polymerase, but sufficient for gene expression.
Transcription Termination
Intrinsic (Rho-independent) Termination
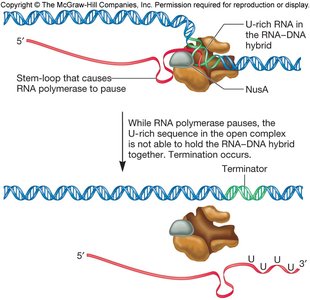
Intrinsic termination relies on specific sequences in the RNA that form a stable hairpin structure followed by a stretch of uracils (U). This causes RNA polymerase to pause and dissociate from the DNA, releasing the RNA transcript.
GC-rich region: Forms a hairpin loop in the RNA.
U-rich sequence: Weakens the RNA-DNA hybrid, facilitating release.
Rho-dependent Termination
Rho-dependent termination requires the Rho protein, an ATP-dependent helicase. Rho binds to the RNA and moves toward the RNA polymerase, causing dissociation when it catches up. This mechanism is used for genes lacking strong intrinsic terminators.
Rho factor: Recognizes specific sequences on the RNA and terminates transcription by disrupting the RNA-DNA hybrid.
Experimental evidence: In vitro transcription of a gene requiring Rho results in longer RNA than in vivo, indicating the necessity of Rho for proper termination.
Comparison: DNA Polymerase vs. RNA Polymerase
Feature | DNA Polymerase | RNA Polymerase |
|---|---|---|
Template | DNA | DNA |
Direction of Synthesis | 5' to 3' | 5' to 3' |
Primer Requirement | Yes | No (de novo synthesis) |
Error Rate | 1/107 - 1/109 | 1/104 |
Strands Copied | Both | One per gene |
Key Terms and Concepts
Promoter: DNA sequence where RNA polymerase binds to initiate transcription.
Terminator: DNA sequence signaling the end of transcription.
Sigma factor: Protein that guides RNA polymerase to specific promoters in prokaryotes.
Consensus sequence: The most common sequence found at a particular position in a set of DNA elements.
Operon: Cluster of genes transcribed as a single mRNA in prokaryotes.
Monocistronic/Polycistronic: Refers to mRNA encoding one or multiple proteins, respectively.
