 Back
BackBiotechnology and Recombinant DNA Technology: Principles and Applications 21
Study Guide - Smart Notes

Biotechnology and Genetic Engineering
Introduction to Biotechnology
Biotechnology is the field of applied biology in which living organisms, or processes derived from them, are used—often with human modification—to serve scientific, medical, or industrial purposes. Modern biotechnology, also known as genetic engineering or molecular biology, relies heavily on knowledge from genetics and molecular biology, especially from model organisms.
Genetic engineering involves the precise modification of an organism's genome.
Applications span medicine, agriculture, environmental science, and industry.
Major Areas of Biotechnology
Medical biotechnology: Drug and vaccine production, genetic testing, pharmacogenomics, gene therapy, stem cell technology, cloning.
Agricultural biotechnology: Genetically modified (GM) animals and plants for improved traits.
Environmental biotechnology: Bioremediation using organisms to clean up pollutants.
Genetic modification of humans: Emerging area with ethical considerations.
The biotechnology industry employs a diverse workforce, including biologists, chemists, computer scientists, engineers, lawyers, and business professionals.
Genetically Modified Organisms (GMOs)
Definition and Historical Context
A genetically modified organism (GMO) is any organism whose genome has been altered by human intervention. While selective breeding has been practiced for thousands of years (e.g., domestication of corn, wheat, cows, sheep, and dogs), modern GMOs are created through precise molecular techniques.

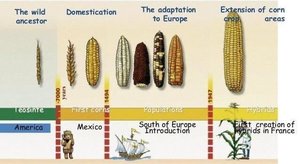
Modern genetic modification includes gene deletion, modification, replacement, or addition (from the same or different species—transgenic organisms).
Purposes of Creating GMOs
Production of large quantities of proteins or drugs (e.g., insulin in bacteria).
Modification of plant or animal phenotypes for agriculture.
Treatment of human diseases (gene therapy).
Functional genomics: Knocking out or modifying genes to study their roles.
Recombinant DNA Technology
Principles of Recombinant DNA
Recombinant DNA technology involves combining DNA from different sources to create new genetic combinations. This is foundational for producing proteins such as human insulin in bacteria.
First successful recombinant DNA drug: Humulin (human insulin), produced by Genentech in 1982.
Key concept: Insert the gene of interest downstream of a strong, inducible bacterial promoter (e.g., lac promoter).
Plasmids as Cloning Vectors
Plasmids are small, circular, extrachromosomal DNA molecules found naturally in bacteria. They can replicate independently and often carry antibiotic resistance genes, making them useful as vectors for gene cloning.
Essential features of plasmid vectors:
Origin of replication (ori)
Selectable marker (e.g., antibiotic resistance gene)
Multiple cloning site (MCS) with unique restriction sites
Inducible promoter for gene expression
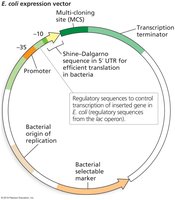
Restriction Enzymes and DNA Ligation
Restriction enzymes are bacterial proteins that cut DNA at specific palindromic sequences (usually 4–8 base pairs). They are named after the organisms from which they were isolated (e.g., EcoRI from E. coli).
Restriction enzymes can create 'blunt' or 'sticky' ends, facilitating the joining of DNA fragments from different sources.
DNA ligase is used to seal the sugar-phosphate backbone, forming recombinant DNA molecules.
Example recognition sequence for EcoRI:
Cloning a Gene into a Plasmid
To clone a gene (e.g., human insulin) into a plasmid:
Obtain the gene of interest, often as cDNA (complementary DNA) to avoid introns.
Add restriction sites to the ends of the cDNA using linker DNA.
Digest both the plasmid and the cDNA with the same restriction enzyme.
Ligate the fragments together with DNA ligase.
Transform the recombinant plasmid into bacteria.
Select for bacteria containing the plasmid using antibiotic resistance.
Transformation and Selection
Bacteria can naturally take up DNA from their environment—a process called transformation. In the lab, this is used to introduce recombinant plasmids into bacterial cells. Only cells that successfully take up the plasmid (and thus the antibiotic resistance gene) will grow on selective media.
Screening for Recombinant Clones
Blue-white screening is a common method for identifying recombinant bacteria. In the presence of X-gal, bacteria with an intact lacZ gene produce blue colonies, while those with disrupted lacZ (due to insertion of foreign DNA) produce white colonies.
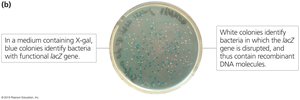
Restriction Mapping and Gel Electrophoresis
Restriction mapping is used to confirm the orientation and presence of the insert. Digestion with specific enzymes followed by agarose gel electrophoresis separates DNA fragments by size, allowing visualization of expected band patterns.
Key points:
DNA is negatively charged and migrates toward the positive electrode.
Smaller fragments move faster through the gel matrix.
DNA bands are visualized using intercalating dyes under UV light.
Applications of Recombinant DNA Technology
Production of Therapeutic Proteins
Genetically modified bacteria are used to produce large quantities of medically important proteins, including:
Human insulin (for diabetes)
Human growth hormone (for dwarfism)
Blood clotting factors (for hemophilia)
Follicle-stimulating hormone (for fertility treatments)
Tissue plasminogen activator (for dissolving blood clots)
Other applications include recombinant vaccines, industrial enzymes, and bioremediation.
Antibiotics and Antibiotic Resistance
Antibiotics: Discovery and Mechanisms
Antibiotics are natural or synthetic compounds that inhibit the growth of bacteria and fungi. The first antibiotic, penicillin, was discovered by Alexander Fleming in 1928. Antibiotics target various bacterial structures and processes, such as cell wall synthesis, protein synthesis, DNA replication, and metabolic pathways.
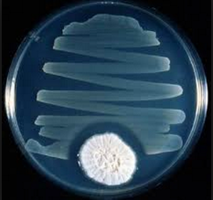
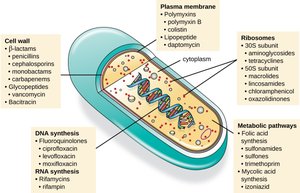
Antibiotic Resistance
Antibiotic resistance arises rapidly due to natural selection, overuse of antibiotics, incomplete treatments, poor hygiene, and prophylactic use in agriculture. Bacteria can acquire resistance by mutation or by obtaining resistance genes from other bacteria via plasmids.
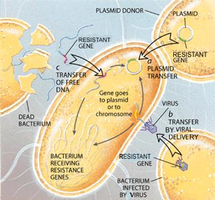
Multi-drug resistant (MDR) bacteria are a growing global health threat.
Plasmids play a key role in spreading resistance genes.
Plasmids in Antibiotic Resistance and Biotechnology
Plasmids are 'selfish' DNA molecules that often carry antibiotic resistance genes, ensuring their retention in bacterial populations. These properties make them valuable tools in recombinant DNA technology.
Summary Table: Common Type II Restriction Enzymes
Enzyme | Microorganism Source | Recognition Sequence | Type of Fragment End Produced |
|---|---|---|---|
BamHI | Bacillus amyloliquefaciens | 5'-G^GATCC-3' | Sticky |
EcoRI | Escherichia coli | 5'-G^AATTC-3' | Sticky |
HindIII | Haemophilus influenzae | 5'-A^AGCTT-3' | Sticky |
PvuII | Proteus vulgaris | 5'-CAG^CTG-3' | Blunt |
SmaI | Serratia marcescens | 5'-CCC^GGG-3' | Blunt |
Conclusion
Biotechnology and recombinant DNA technology have revolutionized genetics, medicine, and agriculture. Understanding the principles of gene cloning, plasmid vectors, restriction enzymes, and antibiotic resistance is essential for modern genetic analysis and biotechnological applications.
