 Back
BackChapter 13: The Genetic Code and Transcription – Study Notes
Study Guide - Smart Notes

Chapter 13: The Genetic Code and Transcription
Introduction to Transcription
Transcription is the process by which genetic information encoded in DNA is transferred to an RNA molecule. This process is a key step in the central dogma of molecular genetics, which describes the directional flow of genetic information from DNA to RNA to protein. Transcription serves as the first step in gene expression, ultimately leading to the synthesis of proteins.
Central Dogma of Molecular Genetics
Overview
The central dogma outlines the flow of genetic information:
DNA is transcribed into RNA.
RNA is translated into protein.
Gene expression refers to the process by which information from a gene is used to synthesize functional gene products (RNA and proteins).
Transcription: Basic Concepts
Transcription Overview
Transcription (txn) is the synthesis of RNA from a DNA template. The genetic information stored in DNA is transferred to RNA, which serves as an intermediate molecule between DNA and proteins. Each triplet codon in mRNA is complementary to the anticodon of tRNA during translation.
Open Reading Frames (ORFs) and Overlapping Genes
An open reading frame (ORF) is a DNA sequence that produces RNA with defined start and stop signals for transcription. In bacteria and viruses, overlapping genes can occur, where a single mRNA contains multiple initiation points, resulting in different ORFs and more than one protein product.
Differences Between DNA and RNA
Key Differences
Strandedness: DNA is double-stranded; RNA is single-stranded.
Sugar: DNA contains deoxyribose; RNA contains ribose.
Bases: DNA contains thymine (T); RNA contains uracil (U), which pairs with adenine (A).
RNA Polymerase and RNA Synthesis
RNA Polymerase Structure and Function
RNA polymerase is the enzyme responsible for synthesizing RNA from a DNA template. In bacteria, the RNA polymerase holoenzyme consists of multiple subunits, including the sigma (σ) subunit, which is essential for promoter recognition and initiation of transcription. No primer is required for initiation, and the RNA produced is single-stranded.
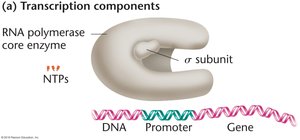
Stages of Transcription in Bacteria
Overview of Steps
Promoter binding
Initiation of transcription
Chain elongation
Termination
Step 1: Promoter Binding
Transcription begins with the binding of RNA polymerase to the promoter region of DNA, a process called template binding. The promoter is a specific DNA sequence located upstream (5’ direction) of the transcription initiation point. The sigma subunit (σ) is responsible for recognizing the promoter. In E. coli, two key promoter sequences are TTGACA and TATAAT.
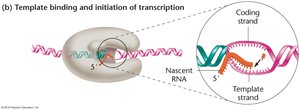
Step 2: Initiation of Transcription
During initiation, the DNA double helix is unwound to make the template strand accessible for RNA polymerase. The interaction between promoters and RNA polymerase regulates the efficiency of transcription. Both cis-acting elements (DNA sequences) and trans-acting factors (proteins) can influence initiation.
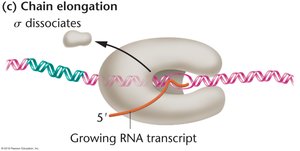
Step 3: Chain Elongation
After initiation, the sigma subunit dissociates, and the core enzyme continues RNA synthesis. Elongation proceeds in the 5’ to 3’ direction, with RNA polymerase adding ribonucleotides complementary to the DNA template strand. The enzyme can also proofread and replace mismatched bases. In bacteria, ribosomes can bind to the RNA transcript during elongation, which differs from eukaryotic transcription.
Coding and Template Strands
During transcription, one DNA strand serves as the template (antisense, -) strand, directing RNA synthesis. The other strand is the coding (sense, +) strand, which has the same sequence as the RNA transcript (except T is replaced by U in RNA). RNA polymerase synthesizes RNA in the 5’ to 3’ direction.
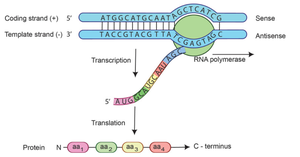
Step 4: Termination
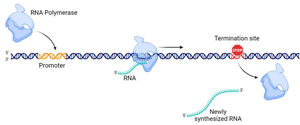
Termination occurs when RNA polymerase encounters a termination sequence in the DNA. In bacteria, intrinsic termination is common and involves the formation of a hairpin structure at the 3’ end of the mRNA transcript, causing RNA polymerase to stall and the transcript to dissociate from the DNA template.
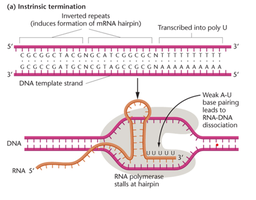
Transcription in Eukaryotes
Key Differences from Prokaryotic Transcription
Occurs in the nucleus (not the cytoplasm).
mRNA must be processed and exported from the nucleus for translation.
Chromatin remodeling is required to make DNA accessible.
Multiple RNA polymerases exist, each with specific functions.
Transcription factors, enhancers, and silencers regulate transcription.
Termination mechanisms differ from those in bacteria.
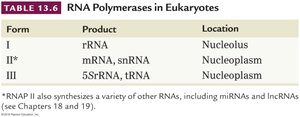
RNA Polymerases in Eukaryotes
Eukaryotes possess three main RNA polymerases, each responsible for transcribing different types of genes:
Form | Product | Location |
|---|---|---|
I | rRNA | Nucleolus |
II | mRNA, snRNA | Nucleoplasm |
III | 5S rRNA, tRNA | Nucleoplasm |
Additional info: RNA Pol II also synthesizes other RNAs, including miRNAs and lncRNAs.
RNA Polymerase II and Transcription Initiation
RNA Polymerase II (RNAPII) transcribes protein-coding genes and requires general transcription factors and core promoter elements, such as the TATA box. The TATA-binding protein (TBP) of transcription factor TFIID binds to the TATA box, determining the transcription start site. Enhancers and silencers modulate the efficiency of transcription initiation.
mRNA Processing in Eukaryotes
Before mRNA can be translated, it undergoes several post-transcriptional modifications:
Addition of a 5’ cap (7-methylguanosine) to protect the mRNA and aid in export and translation initiation.
Addition of a 3’ poly-A tail to prevent degradation and assist in export.
Excision (splicing) of introns to produce mature mRNA.
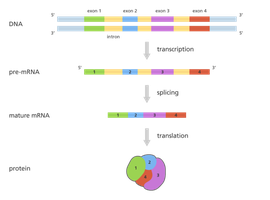
Introns and Exons in Eukaryotic Genes
Introns and Exons
Introns are non-coding sequences in the initial RNA transcript (pre-mRNA) that are not represented in the final mRNA product. Exons are the coding sequences that remain and are expressed in the protein. Prokaryotes do not have introns; only exons are present in their genes.
Splicing of Introns
Splicing is the process by which introns are removed from pre-mRNA, and exons are joined together to form mature mRNA. This process occurs in the spliceosome, a large complex composed of small nuclear RNAs (snRNAs) and proteins (snRNPs or "snurps").
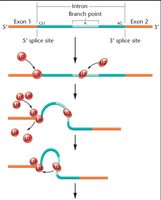
Steps of the Splicing Reaction
U1 snRNP binds to the 5’ splice site of the intron.
U2 snRNP binds to the branch point at the 3’ end of the intron.
U4/U5/U6 complex is recruited to U1.
U1 and U4 are released; U5/U6 bind to U2.
Catalysis occurs, forming a lariat structure and excising the intron.
Exons are ligated together, and the spliceosome is recycled.
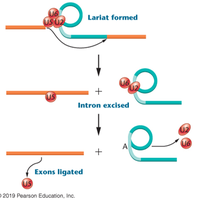
Additional info: The spliceosome is a ribozyme (RNA with catalytic activity). Thomas Cech and Sidney Altman were awarded the Nobel Prize in 1989 for their work on catalytic RNA.
Visualization of Transcription
Electron Microscopy of Transcription
Electron microscopy has allowed scientists to visualize transcription in bacteria, showing RNA strands emanating from DNA templates. Multiple transcription events can occur simultaneously, and in prokaryotes, ribosomes can bind to mRNA while it is still being transcribed (polyribosomes).
