 Back
BackChapter 17: Recombinant DNA Technology – Study Notes
Study Guide - Smart Notes

Recombinant DNA Technology
Introduction to Recombinant DNA Technology
Recombinant DNA technology, also known as gene splicing, involves the creation of DNA molecules by joining together genetic material from different sources. This technology allows scientists to isolate, manipulate, and study specific DNA sequences, enabling advances in genetics, biotechnology, and medicine.
Recombinant DNA: DNA formed by combining DNA from different organisms.
Clones: Identical copies of recombinant DNA molecules, used to study gene structure and function.
Tools of Recombinant DNA Technology
Restriction Enzymes
Restriction enzymes are essential tools for cutting DNA at specific sequences, enabling the manipulation and recombination of genetic material.
Produced by bacteria as a defense against bacteriophage infection.
Recognize specific DNA sequences (restriction sites) and cleave both DNA strands, generating restriction fragments.
Cleavage can produce sticky ends (overhangs) or blunt ends (no overhangs).
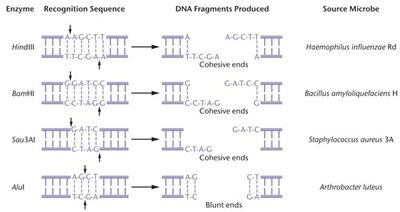
Palindromic Sequences and DNA Ligation
Restriction sites are often palindromic, meaning the sequence reads the same on both DNA strands. Sticky ends facilitate the joining of DNA fragments from different sources.
Sticky ends: Single-stranded overhangs that can anneal with complementary sequences.
DNA ligase: Enzyme that covalently joins DNA fragments by sealing the phosphodiester backbone.
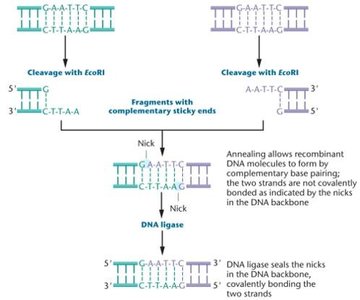
Cloning Vectors
Properties of Cloning Vectors
Cloning vectors are DNA molecules that can carry foreign DNA fragments and replicate within a host cell. They are engineered to facilitate the insertion and selection of recombinant DNA.
Must replicate independently of the host chromosome.
Contain multiple restriction sites (multiple cloning site, MCS).
Carry selectable marker genes (e.g., antibiotic resistance) to identify host cells with the vector.
Bacterial Plasmid Vectors
Plasmids are circular DNA molecules used as vectors in bacterial cloning. They are engineered with features such as MCS and selectable markers.
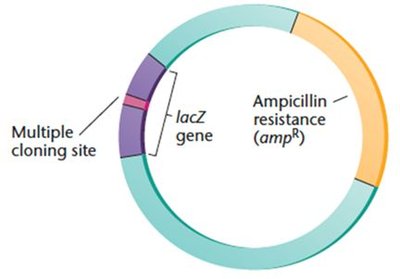
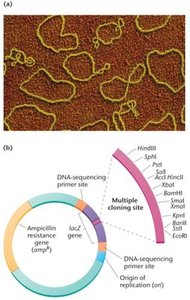
Transformation and Cloning Process
Transformation is the process of introducing recombinant plasmids into bacterial cells. Two main techniques are used: calcium ion/heat shock and electroporation.
Plasmid and target DNA are cut with the same restriction enzyme.
Fragments are ligated to form recombinant DNA.
Recombinant plasmids are introduced into bacteria, which are then selected for further study.
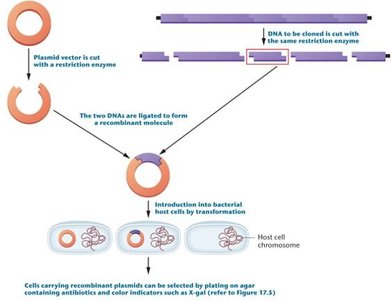
Blue-White Screening
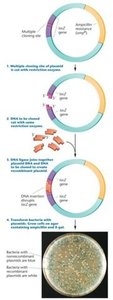
Blue-white screening is a method to distinguish recombinant from nonrecombinant plasmids using the lacZ gene and X-gal substrate.
lacZ gene: Encodes β-galactosidase, which cleaves X-gal to produce a blue color.
Bacteria with recombinant plasmids (disrupted lacZ) form white colonies; nonrecombinant form blue colonies.
Other Types of Cloning Vectors
For larger DNA fragments, alternative vectors are used:
Phage vectors: Modified bacteriophage genomes, can carry up to 45 kb of DNA.
Bacterial artificial chromosomes (BACs): Large plasmids for 100–300 kb inserts.
Yeast artificial chromosomes (YACs): Can carry up to 1000 kb, contain telomeres, centromere, and origin of replication.
Expression Vectors
Expression vectors are designed to ensure transcription and translation of cloned genes, enabling protein production in host cells. They are available for both prokaryotic and eukaryotic systems.
Genomic and cDNA Libraries
Genomic Libraries
A genomic library is a collection of DNA fragments representing the entire genome of an organism, constructed by cloning overlapping DNA fragments into vectors.
Contains both coding and noncoding sequences (e.g., introns).
cDNA Libraries
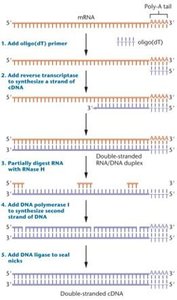
cDNA libraries are made from mRNA and represent only the expressed genes at the time of library construction. Useful for studying gene expression and identifying genes involved in specific processes.
Constructed by reverse transcription of mRNA to cDNA, followed by cloning into vectors.
Library Screening
Library screening uses labeled DNA or RNA probes to identify and isolate specific genes from a library. Probes must be complementary to the target sequence and labeled for detection.
Polymerase Chain Reaction (PCR)
PCR Principles and Requirements
PCR is a rapid, in vitro method for amplifying specific DNA sequences without the need for cloning in host cells.
Requires double-stranded target DNA, DNA polymerase, primers, four dNTPs, and Mg2+ as a cofactor.
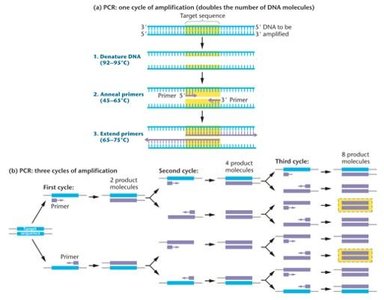
PCR Process
Denaturation: DNA strands are separated by heating.
Annealing: Primers bind to complementary sequences on single-stranded DNA.
Extension: DNA polymerase synthesizes new DNA strands from primers.
Each cycle doubles the amount of target DNA, leading to exponential amplification.
Limitations and Applications of PCR
Requires prior knowledge of target sequence for primer design.
Highly sensitive to contamination.
Cannot efficiently amplify very long DNA segments.
Widely used in genetic testing, forensics, and molecular diagnostics.
RT-PCR and qPCR
RT-PCR: Reverse transcription PCR, used to study gene expression by converting mRNA to cDNA before amplification.
qPCR: Quantitative real-time PCR, allows real-time monitoring and quantification of DNA amplification.
Agarose Gel Electrophoresis
Principle and Application
Agarose gel electrophoresis separates DNA fragments by size. Smaller fragments migrate farther through the gel matrix. DNA is visualized using stains and UV illumination.
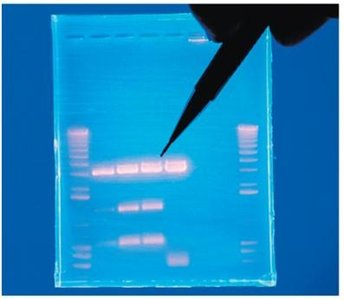
Blotting Techniques
Southern Blot
Southern blotting identifies specific DNA sequences within a complex mixture. It involves gel electrophoresis, transfer to a membrane, hybridization with a labeled probe, and detection.
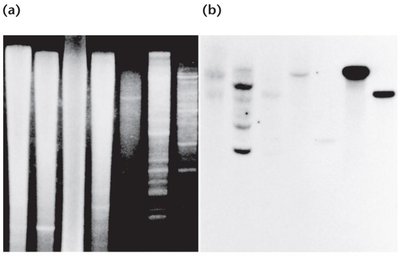
Northern and Western Blotting
Northern blot: Used to study RNA expression patterns (gene transcription).
Western blot: Used to detect and analyze proteins.
Fluorescent in Situ Hybridization (FISH)
FISH uses fluorescently labeled probes to detect specific DNA or RNA sequences directly in chromosomes or tissues, useful for karyotyping and developmental studies.
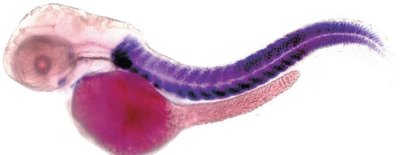
DNA Sequencing
Sanger Sequencing (Dideoxynucleotide Chain-Termination)
Sanger sequencing is the most common method for determining DNA sequences. It uses dideoxynucleotides to terminate DNA synthesis at specific bases, generating fragments that can be separated and read to determine the sequence.
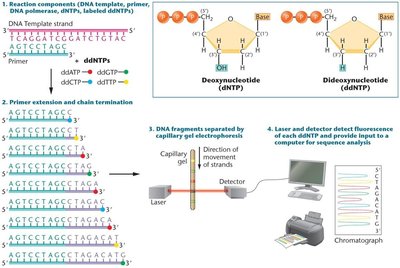
Genome Editing with CRISPR-Cas
CRISPR-Cas System
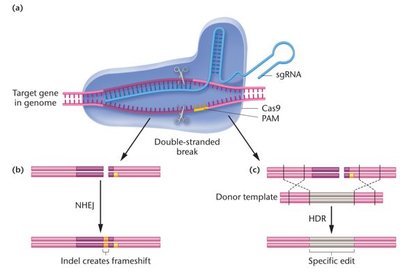
CRISPR-Cas is a revolutionary genome editing tool that allows precise removal, addition, or alteration of DNA sequences in living cells. The Cas9 nuclease, guided by a single guide RNA (sgRNA), introduces double-stranded breaks at specific genomic locations.
Cas9 recognizes a protospacer adjacent motif (PAM) sequence (5'-NGG-3').
Double-stranded breaks are repaired by nonhomologous end-joining (NHEJ) or homology-directed repair (HDR).
Limitations of CRISPR-Cas
Potential for off-target effects due to imperfect sgRNA binding.
Ongoing improvements include engineering Cas9 for higher specificity and optimizing sgRNA design.
