 Back
BackChromosomal Variations, Mutations, and DNA Repair Mechanisms
Study Guide - Smart Notes

Chromosomal Variation
Types of Inversions
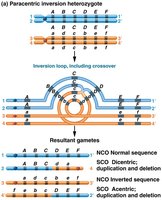
Inversions are chromosomal rearrangements in which a segment of a chromosome is reversed end to end. They can significantly impact genetic function and inheritance patterns.
Paracentric Inversion: The centromere is not part of the inverted segment. The lengths of the chromosome arms remain unchanged.
Pericentric Inversion: The centromere is included in the inverted segment, resulting in changes to the lengths of the chromosome arms.
Mechanism: Inversions occur when a chromosome segment is removed, flipped, and reinserted in the opposite orientation.
Genetic Consequences: In homozygotes, recombination produces gametes with the correct gene copy number. In heterozygotes, recombination can result in gametes with duplications or deletions.
Translocations
Translocations involve the transfer of a chromosome segment to a nonhomologous chromosome. They can be classified as:
Reciprocal Translocation: Two chromosomes exchange segments.
Intrachromosomal: Occurs within a homologous pair.
Interchromosomal: Occurs between nonhomologous chromosomes.
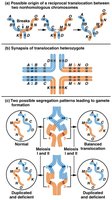
Robertsonian Translocation
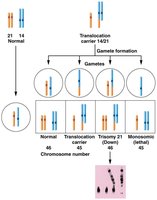
Robertsonian translocation (centric fusion) involves breaks at the ends of two nonhomologous acrocentric chromosomes, resulting in the fusion of the long arms and loss of the short arms. This can lead to genetic disorders such as familial Down syndrome (14/21 translocation).
Mutation, Repair, and Recombination
Gene Mutations
Mutations are alterations in the DNA sequence. They can occur in somatic or germ cells and may affect coding or noncoding regions.
Point Mutation: Change from one base pair to another.
Missense Mutation: New triplet codes for a different amino acid.
Nonsense Mutation: New triplet codes for a stop codon, terminating translation prematurely.
Silent Mutation: New triplet still codes for the same amino acid.
Frameshift Mutation: Insertions or deletions that shift the reading frame, altering downstream amino acid sequence.
Classification by Phenotype
Loss-of-function: Reduces or eliminates gene product function.
Gain-of-function: Enhances or creates new gene function.
Null Mutation: Complete loss of function.
Lethal Mutation: Interrupts essential processes, causing death.
Conditional Mutation: Expressed only under certain environmental conditions (e.g., temperature-sensitive mutations).
Neutral Mutation: No effect on organism's fitness.
Mutation Rates and Causes
Spontaneous Mutations: Occur naturally due to errors in replication or normal cellular processes.
Induced Mutations: Result from exposure to mutagens such as radiation, chemicals, or UV light.

Mechanisms of Spontaneous Mutation
Replication Slippage: DNA polymerase slips on repeat sequences, causing insertions or deletions.
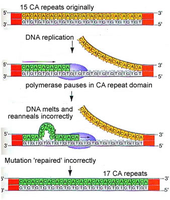
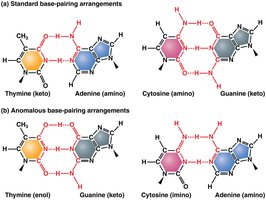
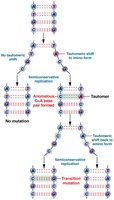
Tautomeric Shifts: Temporary changes in base structure increase mispairing during replication, leading to point mutations.
Depurination: Loss of a purine base, creating an apurinic site.
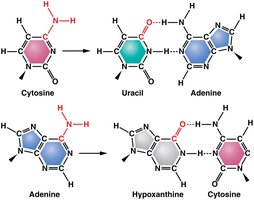
Deamination: Removal of an amino group from cytosine or adenine, altering base-pairing properties.
Oxidative Damage and Free Radicals
Reactive oxygen species generated during cellular metabolism can damage DNA, causing base modifications, strand breaks, and chromosomal rearrangements.
Induced Mutations
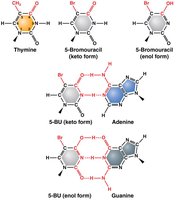
Base Analogs: Chemicals that mimic DNA bases and are incorporated during replication, increasing mutation rates (e.g., 5-bromouracil).
Alkylating Agents: Add alkyl groups to bases, altering base-pairing and causing transitions (e.g., ethylmethanesulfonate).
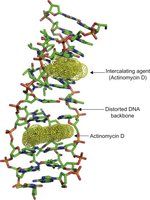
Intercalating Agents: Insert between DNA bases, causing frameshift mutations (e.g., ethidium bromide).
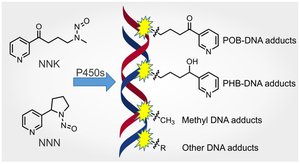
Adduct-Forming Agents: Covalently bind to DNA, distorting its structure and interfering with replication (e.g., acetaldehyde, HCAs).
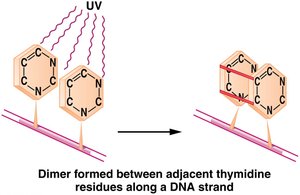
UV Light: Induces pyrimidine dimers, distorting DNA and blocking replication.
Ionizing Radiation: Causes single- and double-strand breaks, base damage, and chromosomal fragmentation.
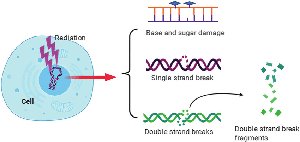
DNA Repair Mechanisms
Proofreading and Mismatch Repair
DNA polymerase proofreads newly synthesized DNA, removing and replacing incorrect nucleotides. If errors escape proofreading, mismatch repair systems detect and correct them, using strand discrimination based on methylation patterns.
Excision Repair
Base Excision Repair (BER): Removes and replaces damaged bases.
Nucleotide Excision Repair (NER): Removes bulky lesions that distort the double helix.
Double-Strand Break Repair
Homologous Recombination Repair: Uses a homologous sequence as a template for accurate repair.
Nonhomologous End Joining: Directly ligates broken DNA ends, often resulting in mutations.
Transposable Elements
Definition and Effects
Transposable elements (TEs) are DNA sequences that can move within or between genomes. They can disrupt gene function, alter gene expression, and cause chromosomal rearrangements.
Autonomous Elements: Can transpose independently.
Nonautonomous Elements: Require the presence of autonomous elements for transposition.
TEs are found in all organisms and contribute to genetic diversity and evolution. In humans, LINEs and SINEs are common TEs, and their movement can cause mutations.
