 Back
BackChromosome Mapping and Gene Linkage: Principles and Applications
Study Guide - Smart Notes

Chromosome Mapping in Eukaryotes
Gene Linkage and Recombination
Chromosome mapping is a fundamental technique in genetics used to determine the relative positions of genes on a chromosome. Linked genes are inherited together unless separated by recombination events during meiosis.
Single crossover: Used to determine the distance between two linked genes. The frequency of recombination is proportional to the physical distance between genes.
Double crossover: Involves two independent exchanges of genetic material and is used to map three linked genes. The probability of double crossovers is calculated using the product law.
Multiple crossovers: While possible, their probability decreases as genes are closer together.
Map unit (centimorgan, cM): One map unit equals one percent recombination between two genes.
Example: If the distance between gene A and B is 10 cM, and between B and C is 20 cM, the distance between A and C is 30 cM.
Equation:
(2%)
Three-Point Mapping Criteria
Three-point mapping is used to determine the order and distance of three genes on a chromosome. The following criteria must be met:
The parent must be heterozygous for all three genes under consideration.
The phenotypic class must reflect the genotype of gametes of parents.
A sufficient number of offspring must be produced for a representative sample.
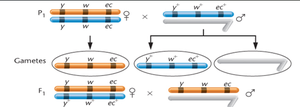
Analysis of Offspring Phenotypes
In three-point mapping experiments, offspring are categorized based on the type of crossover event:
Parental (non-crossover) gametes: Produce the largest percentage of offspring.
Double crossover (DCO) gametes: Occur in the least percentage of offspring.
Single crossover (SCO) gametes: Represent intermediate percentages.
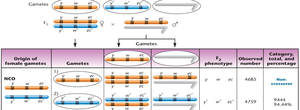
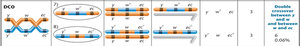
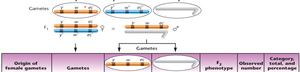
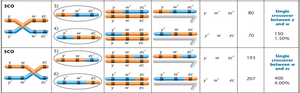
Calculating Map Distances
Map distances are calculated using recombination frequencies. DCOs are included because they represent two simultaneous SCOs.
Formula:
Example: If w and ec = 4% SCO + 0.06% DCO = 4.06 cM
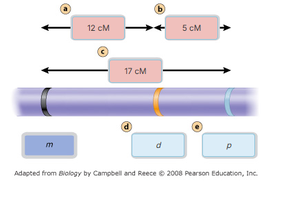
Practical Example: Tomato Gene Mapping
Gene mapping can be applied to various organisms. The following table shows the results of a cross in tomatoes, used to determine recombination frequencies and map distances for M, D, and P genes.
Phenotype | Genotype | Number of Individuals |
|---|---|---|
solid, normal, smooth | MmDdPp | 420 |
solid, normal, peach | MmDdpp | 21 |
solid, dwarf, smooth | MmddPp | 52 |
solid, dwarf, peach | Mmddpp | 62 |
mottled, normal, smooth | mmDdPp | 43 |
mottled, normal, peach | mmDdpp | 23 |
mottled, dwarf, smooth | mmddPp | 46 |
mottled, dwarf, peach | mmddpp | 333 |

Mapping Accuracy and Interference
Limitations of Mapping
As the distance between two genes increases, mapping estimates become less accurate due to undetected crossover events. This leads to underestimation of actual distances.
Genes farther apart increase the probability of undetected crossovers.
Mapping accuracy is highest when genes are close together.
Interference and Coefficient of Coincidence
Interference is the inhibition of further crossover events by another nearby crossover. It reduces the expected number of multiple crossovers and is quantified by the coefficient of coincidence (C).
Interference (I):
Positive interference: Fewer double-crossover events than expected (I positive).
Negative interference: More double-crossover events than expected (I negative).
Applications and Historical Methods
Drosophila and Other Model Organisms
Extensive chromosome mapping has been performed in organisms such as Drosophila, maize, and mice, due to the large number of mutants available.
Human Chromosome Mapping
Historically, the lod score method and somatic cell hybridization were used to map human chromosomes. The lod score (logarithm of the odds) relies on probability calculations and pedigree analysis, as controlled crosses are not possible in humans.

Modern Mapping with DNA Markers
DNA markers such as RFLPs (Restriction Fragment Length Polymorphisms) and SNPs (Single Nucleotide Polymorphisms) are used to map genes. These polymorphisms can "sort" with other genes, allowing scientists to map disease genes to specific chromosomes.
RFLPs and SNPs: Variations in the genetic sequence that are used as markers for gene mapping.
Applications: Mapping disease genes, ancestry testing, and population genetics.
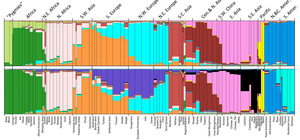
Example: SNPs can be used to determine geographic origins and ethnic background by detecting large numbers of SNPs frequent in certain regions.
Summary Table: Types of Crossover Events
Type | Description | Frequency |
|---|---|---|
Non-crossover (NCO) | No exchange of genetic material | Highest |
Single crossover (SCO) | One exchange between two genes | Intermediate |
Double crossover (DCO) | Two independent exchanges | Lowest |
Additional info: Mapping techniques are continually evolving, with modern approaches relying heavily on DNA sequencing and computational databases for high-resolution mapping.
