 Back
BackChromosome Mapping and Linkage in Eukaryotes: Study Guide
Study Guide - Smart Notes

Chromosome Mapping in Eukaryotes
Introduction
Chromosome mapping is a fundamental technique in genetics that allows researchers to determine the relative positions of genes on chromosomes. This process is essential for understanding gene linkage, recombination, and inheritance patterns in eukaryotic organisms. The following study notes cover key concepts from Chapter 5, including gene linkage, crossing over, and mapping principles, with a focus on Drosophila genetics and Huntington's disease as examples.
Gene Linkage and Independent Assortment
Genes Linked on the Same Chromosome Segregate Together
Genes located on the same chromosome are said to be linked and tend to be inherited together. This is in contrast to genes on different chromosomes, which assort independently during meiosis.
Independent Assortment: Genes on different chromosomes sort independently, resulting in equal probability of each allele combination in gametes.
Linked Genes: Genes on the same chromosome do not assort independently and are transmitted together unless crossing over occurs.
Complete Linkage: No crossing over occurs between linked genes, so only parental (non-crossover) gametes are produced.
'
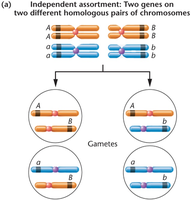
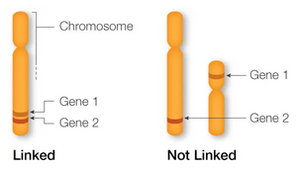
Meiotic Consequences of Linkage
During meiosis, the arrangement of genes on chromosomes determines the types of gametes produced. Linked genes can result in parental and recombinant gametes depending on whether crossing over occurs.
Parental Gametes: Produced when no crossing over occurs between linked genes.
Recombinant Gametes: Produced when crossing over occurs between non-sister chromatids, resulting in new allele combinations.
Chiasmata: Physical sites of crossing over, observed during Prophase I of meiosis.
Principles of Linkage, Crossing-Over, and Mapping
Framework and Mapping Principles
Chromosomes are the units of transmission in meiosis, and linked genes cannot undergo independent assortment. The frequency of crossing over between genes is proportional to their distance apart, which forms the basis for chromosome mapping.
Chromosome Maps: Indicate the relative locations of genes based on recombination frequencies.
Recombination Frequency: Used to estimate the distance between genes; higher frequency indicates greater distance.
Gene Representation: Dominant alleles are indicated by italic uppercase letters, recessive alleles by italic lowercase letters, and wild-type alleles by a plus sign (e.g., e+).
Drosophila Crosses and Linkage Analysis
Example: Complete Linkage in Drosophila
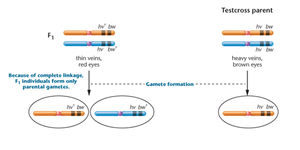
Drosophila melanogaster is a model organism for studying gene linkage. In crosses involving two recessive mutations (heavy wing vein and brown eye), complete linkage results in only parental gametes and characteristic F2 ratios.
Parental Cross: hv+/hv+ bw/bw × hv/hv bw+/bw+
F1 Generation: All flies are heterozygous for both gene pairs and exhibit dominant traits.
F2 Generation: 1:2:1 ratio (thin vein/brown eye : thin vein/red eye : heavy vein/red eye) is observed, characteristic of complete linkage.

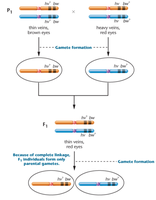
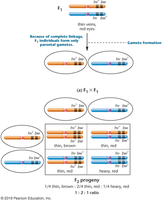
Test Crosses and Linkage Ratios
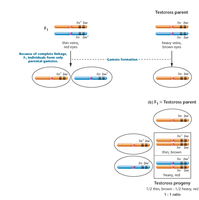
Test crosses are used to determine linkage by mating F1 individuals with homozygous mutants. Complete linkage produces a 1:1 ratio in the F2 generation, while independent assortment would produce a 1:1:1:1 ratio.
Test Cross: F1 (heterozygous) × homozygous mutant
Complete Linkage: Only parental gametes are produced, resulting in a 1:1 ratio.
Independent Assortment: Four phenotypes are produced in a 1:1:1:1 ratio.
Huntington's Disease and Genetic Anticipation
Genetic Anticipation in Huntington's Disease
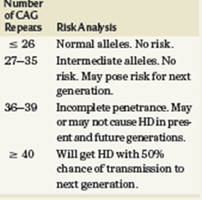
Huntington's disease (HD) is caused by an expansion of CAG repeats in the HTT gene. The number of repeats determines the risk and penetrance of the disease, and repeats may increase in subsequent generations, a phenomenon known as genetic anticipation.
Normal Alleles: ≤ 26 CAG repeats; no risk.
Intermediate Alleles: 27–35 repeats; no risk, but may pose risk for next generation.
Incomplete Penetrance: 36–39 repeats; may or may not cause HD in present and future generations.
Pathogenic Alleles: ≥ 40 repeats; will cause HD with 50% chance of transmission to next generation.
Genetic Anticipation: CAG repeats may expand during gamete formation, increasing risk in offspring.
Number of CAG Repeats | Risk Analysis |
|---|---|
≤ 26 | Normal alleles. No risk. |
27–35 | Intermediate alleles. No risk. May pose risk for next generation. |
36–39 | Incomplete penetrance. May or may not cause HD in present and future generations. |
≥ 40 | Will get HD with 50% chance of transmission to next generation. |

Summary Table: Key Concepts in Chromosome Mapping
Concept | Description |
|---|---|
Gene Linkage | Genes on the same chromosome are inherited together unless crossing over occurs. |
Independent Assortment | Genes on different chromosomes assort independently during meiosis. |
Crossing Over | Physical exchange between non-sister chromatids during Prophase I, resulting in recombinant gametes. |
Chromosome Mapping | Determining the relative positions of genes based on recombination frequency. |
Genetic Anticipation | Expansion of repeat sequences (e.g., CAG) in successive generations, increasing disease risk. |
Key Equations
Recombination Frequency
The recombination frequency (RF) is used to estimate the distance between genes:
One map unit (centimorgan, cM) corresponds to 1% recombination frequency.
Conclusion
Understanding gene linkage, crossing over, and chromosome mapping is essential for analyzing inheritance patterns and genetic diseases. Drosophila crosses and Huntington's disease provide practical examples of these principles in action.
