 Back
BackChromosome Mapping and Linkage in Eukaryotes
Study Guide - Smart Notes

Chromosome Mapping and Linkage in Eukaryotes
Linkage and Mendel’s Postulates
Linkage refers to the phenomenon where genes that are located close together on the same chromosome tend to be inherited together, violating Mendel’s postulate of independent assortment. This affects the expected phenotypic ratios in genetic crosses.
Key Point: Linkage does not affect the number of alleles or dominance relationships, but it does impact the independent assortment of homologous chromosomes during meiosis.
Example: Two genes on the same chromosome that do not undergo crossing over will be inherited as a unit.
f.
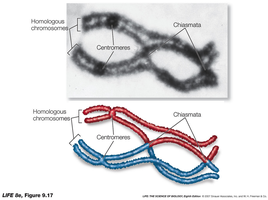
Crossing Over and Chiasmata
Crossing over is the exchange of genetic material between non-sister chromatids during meiosis, occurring at structures called chiasmata. This process produces both parental and recombinant gametes, increasing genetic diversity.
Chiasmata: X-shaped regions where homologous chromosomes exchange genetic material.
Frequency: The likelihood of crossing over between two genes depends on their physical distance on the chromosome.
Example: Genes that are close together have a lower probability of a chiasma forming between them, resulting in fewer recombinant gametes.
Historical Context: Morgan and Sturtevant
Thomas Hunt Morgan and Alfred Sturtevant were pioneers in the study of linkage and chromosome mapping using Drosophila melanogaster (fruit flies). Sturtevant constructed the first genetic map by analyzing recombination frequencies.
Key Point: The frequency of recombination between genes is proportional to their distance apart on the chromosome.
Application: Genetic mapping allows researchers to determine the linear order and relative distances of genes.



Map Units and Recombination Frequency
Sturtevant defined the map unit (mu), also called the centiMorgan (cM), as the distance between genes for which one product of meiosis in 100 is recombinant. Recombination frequencies are additive but may not always sum exactly due to multiple crossovers.
Definition: 1 map unit (1 cM) = 1% recombination frequency.
Calculation: If two genes are 10 mu apart, 10% of gametes are expected to be recombinant for those genes.
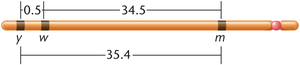
Example: Three genes (y, w, m) on the Drosophila X chromosome have recombination rates that reflect their physical distances.
Single and Double Crossovers
Single crossovers (SCO) involve one exchange between two non-sister chromatids, while double crossovers (DCO) involve two exchanges. SCOs are used to determine distances between two genes, and DCOs are used for mapping three genes.
Single Crossover: Produces two parental and two recombinant gametes.
Double Crossover: The probability is the product of the probabilities of each single crossover event.
Equation:
Example: If A-B is 10 mu and B-C is 20 mu, DCO frequency is or 2%.
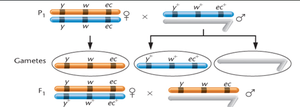
Three-Point Test Crosses and Mapping Criteria
Three-point mapping involves analyzing the offspring of a cross involving three genes to determine their order and distances. The parent must be heterozygous for all three genes, and a large number of offspring is required for statistical accuracy.
Criteria:
Parent must be heterozygous for all three genes.
Phenotypic classes must reflect parental gametes.
Sufficient offspring must be produced for representative sampling.
Interpretation: Parental (non-crossover) gametes produce the most offspring, single crossovers produce intermediate numbers, and double crossovers produce the fewest.
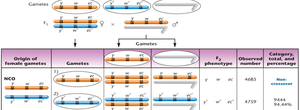
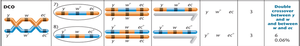
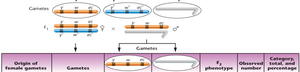
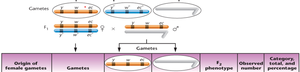
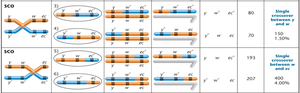
Calculating Map Distances
Map distances are calculated by adding the percentages of single and double crossover events between genes. Double crossovers are included because they represent two simultaneous single crossovers.
Equation:
Example: If the recombination frequency between w and ec is 4% and the DCO frequency is 0.06%, the total map distance is mu.
Summary Table: Types of Gametes and Expected Percentages
Type | Gametes | Expected Percentage | Description |
|---|---|---|---|
Parental (NCO) | Non-crossover | Largest | Most frequent, reflect parental combinations |
Single Crossover (SCO) | Single exchange | Intermediate | Reflect recombination between two genes |
Double Crossover (DCO) | Double exchange | Smallest | Least frequent, require two crossovers |
Additional info: The principles and calculations described here are foundational for understanding chromosome mapping, gene linkage, and recombination in eukaryotes, especially as applied in classical genetics using model organisms like Drosophila melanogaster.
