 Back
BackChromosome Mapping in Eukaryotes: Linkage, Crossing Over, and Genetic Mapping
Study Guide - Smart Notes

Chromosome Mapping in Eukaryotes
Introduction
Chromosome mapping is a fundamental technique in genetics that allows researchers to determine the relative positions of genes on a chromosome. This process relies on the principles of linkage and recombination, which describe how genes are inherited together or separated during meiosis. Understanding these concepts is essential for interpreting genetic crosses and for constructing genetic maps that are crucial in both basic research and applied genetics.
Linkage, Crossing Over, and Mapping
Chromosomes as Units of Transmission
Chromosomes are the primary units of inheritance during meiosis, not individual genes.
Linked genes are genes located on the same chromosome and tend to be inherited together.
The frequency of crossing over between two genes is proportional to the distance between them on the chromosome.
Chromosome maps indicate the relative locations of genes based on recombination frequencies.
5.1 Genes Linked on the Same Chromosome Segregate Together
Meiotic Consequences of Linkage
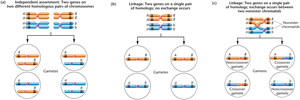
Independent assortment: Genes on different chromosomes assort independently during meiosis.
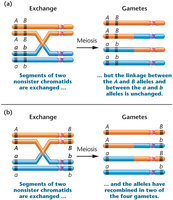
Complete linkage: Genes are so close together that no crossing over occurs; only parental (noncrossover) gametes are produced.
Linkage with crossing over: Crossing over between nonsister chromatids produces both parental and recombinant gametes.
Linkage Ratio and Linkage Groups
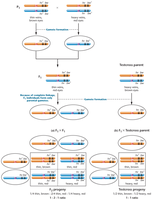
Linkage ratio: The unique phenotypic ratio resulting from complete linkage between two genes.
Linkage group: A set of genes located on the same chromosome; the number of linkage groups corresponds to the haploid number of chromosomes.
5.2 Crossing Over Serves as the Basis for Determining the Distance between Genes in Chromosome Mapping
Chiasmata and Genetic Exchange
Chiasmata are X-shaped structures formed during meiosis where homologous chromosomes exchange genetic material.
The percentage of recombinant offspring depends on the distance between two genes; closer genes are less likely to have a chiasma form between them.
Sturtevant and Genetic Mapping
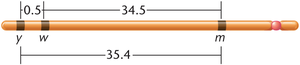
Alfred Sturtevant demonstrated that recombination frequencies between linked genes are additive and can be used to estimate relative distances between genes.
Map unit (m.u.) or centiMorgan (cM): 1% recombination frequency equals 1 map unit.
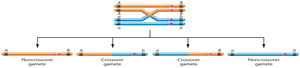
Single Crossovers
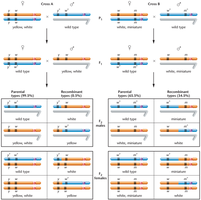
Single crossover (SCO): An exchange between two nonsister chromatids, resulting in 50% recombinant gametes if genes are far apart.
In genes 50 map units apart, crossing over can be expected between 100% of tetrads.
5.3 Determining the Gene Sequence during Mapping Requires the Analysis of Multiple Crossovers
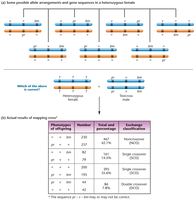
Multiple Crossovers and Three-Point Mapping
Double crossover (DCO): Two exchanges of genetic material, used to determine the order and distance of three linked genes.
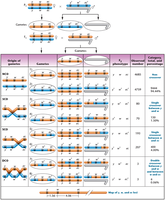
Three-point mapping requires: (1) a parent heterozygous for all three genes, (2) phenotypic classes reflecting parental gametes, and (3) a sufficient number of offspring.
Noncrossover phenotypes occur most frequently; double-crossover phenotypes occur least frequently.
Reciprocal classes are pairs of phenotypes that complement each other and are derived from heterozygotes.
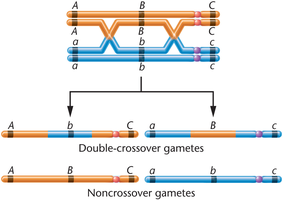
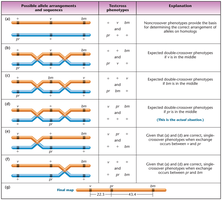
Determining Gene Sequence
Gene order can be determined by comparing double-crossover phenotypes to parental types.
Two main methods: (1) analyzing possible gene arrangements, (2) using double-crossover data to confirm gene order.
5.4 As the Distance between Two Genes Increases, Mapping Estimates Become More Inaccurate
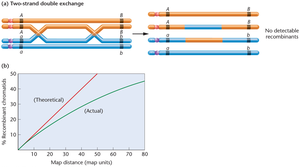
Map Distance and Interference
The expected frequency of multiple crossovers increases with the distance between genes, but some crossovers may go undetected, leading to underestimation of map distances.
Interference (I): The phenomenon where one crossover event inhibits the occurrence of another nearby crossover.
Coefficient of coincidence (C): Quantifies the difference between observed and expected double crossovers.
Positive interference: Fewer double crossovers than expected (I > 0); negative interference: more double crossovers than expected (I < 0).
5.5 Drosophila Genes Have Been Extensively Mapped
Model Organisms and Chromosome Maps
Extensive mapping has been performed in model organisms such as Drosophila, maize, and mice due to the availability of many mutants.
These maps have been instrumental in understanding gene order and genetic distances.

Key Terms and Concepts
Linkage: The tendency of genes located close together on a chromosome to be inherited together.
Recombination: The process by which genetic material is exchanged between homologous chromosomes during meiosis.
Map unit (centiMorgan): A unit of measure for genetic linkage; one map unit equals a 1% recombination frequency.
Chiasma: The physical site of crossing over between homologous chromosomes.
Interference: The phenomenon where one crossover event reduces the probability of another nearby crossover.
Summary Table: Types of Crossovers and Their Consequences
Type of Crossover | Number of Chromatids Involved | Resulting Gametes | Use in Mapping |
|---|---|---|---|
Single Crossover (SCO) | 2 | 2 parental, 2 recombinant | Distance between two genes |
Double Crossover (DCO) | 2 or more | Double recombinant types (least frequent) | Order and distance of three genes |
No Crossover | 0 | All parental | Indicates complete linkage |
Example Calculation: Map Distance
Map distance between two genes can be calculated as:
Conclusion
Chromosome mapping through the analysis of linkage and recombination is a cornerstone of classical genetics. It enables the determination of gene order, genetic distances, and the construction of genetic maps, which are essential for understanding inheritance patterns and for applications in breeding, medicine, and research.
