 Back
BackChromosome Structure and DNA Sequence Organization: Study Guide
Study Guide - Smart Notes

Chromosome Structure and DNA Sequence Organization
Overview
This chapter explores the structural organization of chromosomes and the arrangement of DNA sequences within them. It covers the differences between viral, prokaryotic, and eukaryotic chromosomes, specialized chromosome types, DNA packaging, chromatin remodeling, and the classification of repetitive DNA.
Studying Chromosome Structure
Methods of Study
Chromosome structure is investigated using various techniques:
Microscopy: Allows visualization of chromosome morphology and banding patterns.
Specialized Chromosomes: Polytene and lampbrush chromosomes are used for detailed study.
Enzyme Digestion: Endonucleases help reveal nucleosome structure.
Dyes and Isotope Uptake: Used to highlight specific regions and activities within chromosomes.
Comparing Chromosomes Across Life Forms
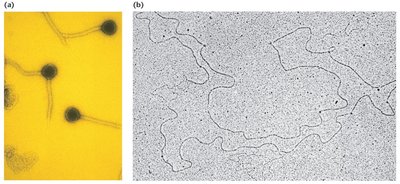
Viruses
Contain single- or double-stranded nucleic acid (RNA or DNA).
Genomes can be circular or linear.
Bacteria
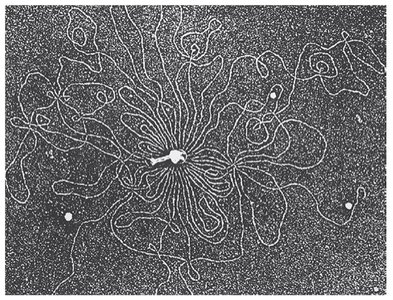
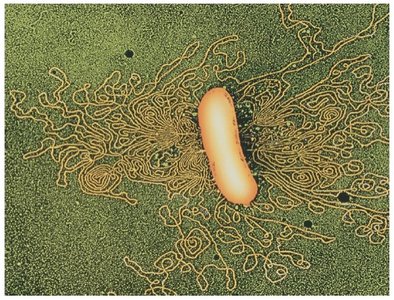
Double-stranded DNA, usually circular.
DNA associates with histone-like proteins for structural support.

Mitochondria and Chloroplasts
Double-stranded, circular DNA.
No DNA-associated proteins.
Typically exhibit maternal inheritance.
Mitochondrial DNA strands have different densities; mitochondrial DNA is smaller than chloroplast DNA.
Specialized Eukaryotic Chromosomes
Polytene Chromosomes
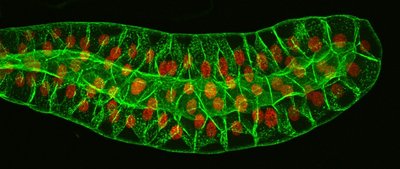
Polytene chromosomes are unusually large and easy to study by microscopy. They are found in certain cells, such as Drosophila salivary glands, and display a distinct banding pattern.
Clusters of identical homologous chromosomes (1000–5000 DNA strands in precise alignment).
Result from many rounds of replication without strand separation or cell division.
Display distinctive banding (chromomere) patterns.
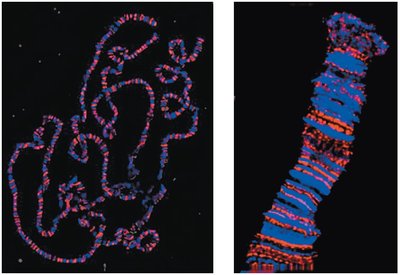
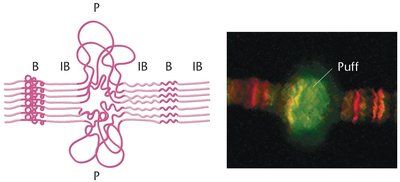
Puff Regions
Localized uncoiled regions of DNA.
Sites of active transcription.
Lampbrush Chromosomes
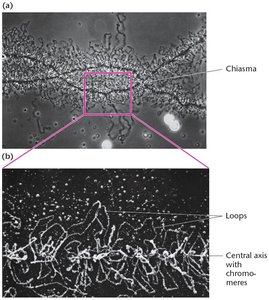
Lampbrush chromosomes are found in most vertebrate oocytes during prophase I of meiosis. They are synapsed but not condensed, and their loops are sites of active gene expression.
~10,000 loops, each corresponding to an uncoiled region of one chromatid.
Central axis composed of two chromatids.
Loops are sites of active transcription.
DNA Packaging in Eukaryotes
Chromatin Structure
Chromatin is DNA associated with proteins, visible in the nucleus under a microscope. During interphase, chromatin is decondensed; during cell division, it condenses for easier chromosome movement.
Histones: Five main types, overall positive charge due to lysine and arginine content.
Non-histones: Various structural and regulatory proteins.
Nucleosome: Basic unit of eukaryotic chromosome structure; DNA wound around an octet of histone proteins.
Nucleosome Structure
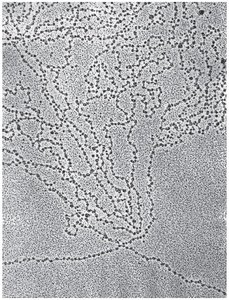
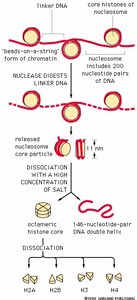
Discovered after digesting DNA with endonuclease.
Contain about 200 base pairs (bp) of DNA; core particle has 147 bp.
Electron microscopy reveals a "beads on a string" structure.
Histone tetramers: H2A2:H2B2 and H32:H42.
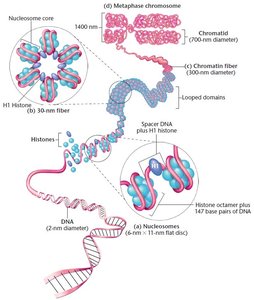
Multiple Levels of DNA Packaging
DNA associates with histones and undergoes subsequent coiling.
H1 histone binds to linker DNA between nucleosomes.
30 nm fiber is the packaging state during interphase.
Chromatin Remodeling and Regulation
Chromatin Remodeling
Chromatin must decondense for DNA replication and gene expression.
Tightly packaged chromatin is inaccessible to regulatory proteins.
Histone tails are targets for chemical modifications (acetylation, methylation, phosphorylation).
These modifications regulate gene expression.
Epigenetics: Heritable yet reversible changes in gene expression not due to changes in DNA sequence.
Heterochromatin vs Euchromatin
Heterochromatin: Condensed, appears darker during interphase, mostly inactive for gene expression.
Euchromatin: Uncoiled, appears lighter during interphase, active for gene expression.
Chromosome Banding
Mitotic chromosomes exhibit characteristic banding patterns visualized by differential staining techniques.
G-Banding: Proteolytic digestion followed by Giemsa staining; reveals A-T rich regions.
C-Banding: Heat denatured DNA followed by Giemsa staining; reveals only the centromere region.
DNA Sequence Organization
Satellite DNA
Eukaryotic DNA can be analyzed by its density.
Majority is seen as one visible peak; satellite DNA differs in density.
Highly repetitive DNA, found in heterochromatic centromeric regions.
Not found in prokaryotes.
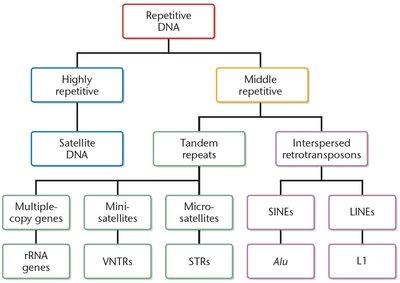
Categories of Eukaryotic Repetitive DNA
Highly repetitive: Centromeres and telomeres.
Moderately (middle) repetitive: VNTRs, STRs, duplicated tRNA and rRNA genes, transposable elements (SINEs and LINEs).
Transposable elements make up 45% of the human genome and are interspersed throughout.
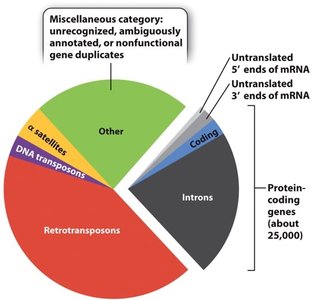
Human Genome Sequence Composition
Protein-coding genes constitute a small fraction (~25,000 genes).
Introns, retrotransposons, DNA transposons, α-satellites, and other non-coding regions make up the majority.
Untranslated regions (5' and 3' ends of mRNA) are also present.
Summary Table: Types of Repetitive DNA
Type | Examples | Location/Function |
|---|---|---|
Highly repetitive | Satellite DNA | Centromeres, telomeres |
Moderately repetitive | VNTRs, STRs, rRNA/tRNA genes, SINEs, LINEs | Interspersed throughout genome, structural and regulatory roles |
Key Equations
DNA Packaging Ratio
The packaging ratio describes how much DNA is compacted:
Histone-DNA Interaction
Each nucleosome core contains:
Conclusion
Understanding chromosome structure and DNA sequence organization is fundamental to molecular genetics. The packaging and modification of chromatin, the presence of specialized chromosomes, and the abundance of repetitive DNA all contribute to the complexity and regulation of the genome.
