Back
BackComprehensive Study Notes: Genetic Mapping, DNA Replication, Gene Expression, and Regulation
Study Guide - Smart Notes

Genetic Linkage and Chromosome Mapping
Core Genetic Assumptions
Classical genetics often begins with simplified assumptions to facilitate understanding of inheritance patterns. These assumptions include:
Two Alleles per Gene: Each gene has only two alleles.
Complete Dominance: One allele is completely dominant over the other.
Autosomal Location: Genes are located on autosomes, not sex chromosomes.
Single-Gene Traits: Each trait is determined by a single gene.
Independent Assortment: Genes are on different chromosomes and assort independently.
Genetic Linkage and Crossing Over
Genes located close together on the same chromosome are linked and do not assort independently.
Complete Linkage: No crossing over occurs; only parental gametes are produced (100% parental type).
Incomplete Linkage: Crossing over occurs, producing both parental and recombinant gametes. Parental types are more frequent than recombinants.
Recombination Frequency and Chromosome Mapping
Recombination Frequency: The maximum frequency between two genes is 50%. Genes more than 50 map units apart behave as unlinked.
Map Unit (mu) / Centimorgan (cM): 1 mu = 1% recombination frequency.
Genetic vs. Physical Maps: Genetic maps are based on recombination frequencies, while physical maps are based on actual DNA length (base pairs).
Type of Map | Basis | Units |
|---|---|---|
Genetic Map | Recombination frequency | cM (centimorgan) |
Physical Map | DNA sequence/base pairs | bp, kb, Mb |
Mapping Methodology
Two-Point Crosses: Used to determine the order and distance between two genes.
Three-Point Mapping: Involves analyzing offspring from trihybrid crosses to determine gene order and distances.
Gene Order Determination: The gene that switches position in double crossover (DCO) progeny compared to non-crossover (NCO) is the middle gene.
Bacterial Genetics and Gene Transfer
Vertical and Horizontal Gene Transfer
Vertical Gene Transfer: DNA is passed from parent to offspring via binary fission.
Horizontal Gene Transfer (HGT): DNA is transferred between organisms without reproduction. Main mechanisms include conjugation, transformation, and transduction.
Mechanisms of Genetic Recombination in Bacteria
Conjugation: Direct transfer of DNA via a pilus. F+ cells transfer the F factor to F- cells. Hfr strains integrate the F factor into the chromosome, allowing chromosomal gene transfer.
Transformation: Uptake of free DNA from the environment by competent cells.
Transduction: Transfer of DNA via bacteriophages (viruses that infect bacteria).
Mapping Bacterial Genes
Interrupted Mating: Used to map gene order by stopping conjugation at intervals and determining which genes have transferred.
Cotransformation/Cotransduction: High cotransfer frequency indicates close proximity of genes.
Nucleic Acid Structure and DNA Replication
Nucleotide Structure
DNA: Deoxyribose sugar, phosphate group, nitrogenous bases (A, T, G, C).
RNA: Ribose sugar, phosphate group, nitrogenous bases (A, U, G, C).
Levels of Nucleic Acid Structure
Primary: Linear sequence of nucleotides joined by phosphodiester bonds.
Secondary: Hydrogen bonding between bases (double helix in DNA, hairpins in RNA).
Tertiary: Three-dimensional folding (e.g., catalytic RNA).
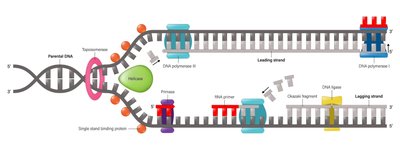
DNA Replication Mechanism
DNA replication is semiconservative and occurs during the S phase of the cell cycle. It involves a complex of enzymes and proteins to ensure accuracy and efficiency.
Directionality: DNA is synthesized in the 5' → 3' direction; new nucleotides are added to the 3' end.
Leading Strand: Synthesized continuously toward the replication fork.
Lagging Strand: Synthesized discontinuously in Okazaki fragments away from the fork.
Key Enzymes: Helicase, primase, DNA polymerase III, DNA polymerase I, DNA ligase, topoisomerase, single-stranded binding proteins.
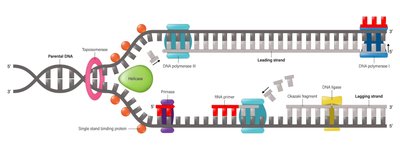
Eukaryotic DNA Replication Complexity
Larger genomes and linear chromosomes require multiple origins of replication.
DNA is packaged into chromatin, adding regulatory complexity.
Telomere replication requires the enzyme telomerase to prevent chromosome shortening.
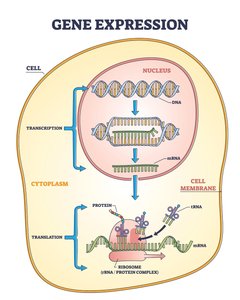
Transcription and RNA Processing
Transcription: DNA to RNA
Transcription is the synthesis of RNA from a DNA template. It involves three main stages:
Initiation: RNA polymerase binds to the promoter region (e.g., TATA box) with the help of transcription factors.
Elongation: RNA polymerase synthesizes RNA in the 5' → 3' direction, unwinding DNA as it moves.
Termination: RNA polymerase stops at a termination sequence, releasing the RNA transcript.
Eukaryotic RNA Processing
5' Capping: Addition of a modified guanine nucleotide to the 5' end for stability and ribosome binding.
3' Polyadenylation: Addition of a poly-A tail to the 3' end for stability and export.
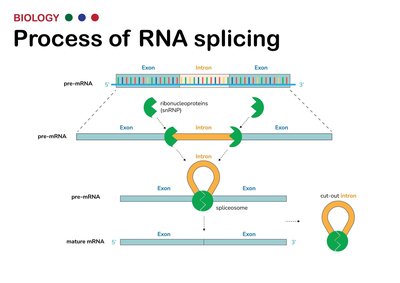
Splicing: Removal of non-coding introns and joining of exons by the spliceosome. Alternative splicing allows for multiple proteins from one gene.
Translation: RNA to Protein
Translation Mechanism
Translation is the process by which ribosomes synthesize proteins using mRNA as a template.
Ribosomes: Composed of large and small subunits with A, P, and E sites.
tRNA: Brings amino acids to the ribosome; anticodon pairs with mRNA codon.
Stages:
Initiation: Ribosome assembles at the start codon (AUG).
Elongation: Amino acids are added one by one to the growing chain.
Termination: Stop codon is reached; release factor releases the polypeptide.
The Genetic Code
Unambiguous: Each codon specifies only one amino acid.
Redundant: Most amino acids are encoded by more than one codon.
Nonoverlapping: Codons are read one at a time.
Nearly Universal: Shared by almost all organisms.
Mutations and DNA Repair
Types of Mutations
Point Mutations: Single nucleotide changes (silent, missense, nonsense).
Insertions/Deletions (Indels): Addition or loss of nucleotides, potentially causing frameshifts.
Functional Effects of Mutations
Loss-of-Function: Reduces or eliminates gene product function.
Gain-of-Function: Produces new or enhanced activity.
Null Mutation: Complete loss of function.
Neutral Mutation: No significant effect on phenotype.
DNA Repair Mechanisms
Proofreading: DNA polymerase corrects errors during replication.
Mismatch Repair (MMR): Corrects errors missed by proofreading.
Base Excision Repair (BER): Removes and replaces damaged bases.
Nucleotide Excision Repair (NER): Removes bulky lesions (e.g., thymine dimers).
Double-Strand Break Repair: Homologous recombination (accurate) or non-homologous end joining (error-prone).
Mutagen Detection: The Ames Test
Uses Salmonella typhimurium his⁻ strains to detect mutagenicity of chemicals.
Reversion to his⁺ indicates mutagenic activity.
Gene Regulation
Prokaryotic Gene Regulation: Operons
Operon Structure: Promoter, operator, and structural genes regulated together.
lac Operon (Inducible): Turned on by lactose (inducer); negative and positive control mechanisms.
trp Operon (Repressible): Turned off by tryptophan (corepressor); attenuation fine-tunes expression.
Eukaryotic Gene Regulation
Chromatin Remodeling: Histone acetylation increases transcription; DNA methylation silences genes.
Transcriptional Control: Promoters, enhancers, silencers, and transcription factors regulate gene expression.
Post-Transcriptional Regulation: Alternative splicing and RNA interference (RNAi) diversify gene products.
Summary Table: Key Genetic Concepts
Concept | Definition | Example/Application |
|---|---|---|
Linkage | Genes close together on a chromosome are inherited together | AB/ab testcross |
Map Unit (cM) | 1% recombination frequency | 15 cM = 15% recombination |
Operon | Cluster of genes under one promoter | lac operon in E. coli |
Frameshift Mutation | Indel not in multiples of three | Insertion of 1 base in coding region |
Illustrative Diagrams