 Back
BackChapter 6
Study Guide - Smart Notes

DNA and Chromosome Structure
Introduction
Understanding the structure and organization of DNA is fundamental to genetics. DNA's molecular architecture underpins heredity, gene expression, and genome packaging across all domains of life. This section explores DNA's chemical structure, its various forms, and the complexity of genome organization in viruses, bacteria, eukaryotes, and archaea.
Essential Characteristics of Hereditary Material
Localization: DNA is found in the nucleus and is a key component of chromosomes.
Stability: DNA maintains a stable form within cells.
Complexity: DNA contains the information necessary for the structure, function, development, and reproduction of organisms.
Replication: DNA can accurately replicate, ensuring genetic continuity between generations.
Mutability: DNA can undergo mutations, providing genetic variation essential for evolution.
Historical Perspective: Discovery of DNA
DNA was first isolated by Friedrich Miescher in the 1860s from white blood cells and salmon sperm.
By the early 1900s, genes were localized to chromosomes (Sutton, Boveri, Fleming).
Experiments from 1928-1952 (Avery, MacLeod, McCarty) established DNA, not RNA, as the hereditary material.
The physical structure of DNA was elucidated in the 1950s by Watson, Crick, Franklin, and Wilkins.


Levels of DNA Organization
Monomer: Nucleotides
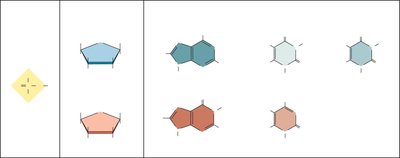
DNA is a polymer composed of nucleotide monomers. Each nucleotide consists of a phosphate group, a deoxyribose sugar, and a nitrogenous base (adenine, thymine, cytosine, or guanine).
Pyrimidines: Single-ring structures (cytosine, thymine)
Purines: Double-ring structures (adenine, guanine)
Single Strand of DNA
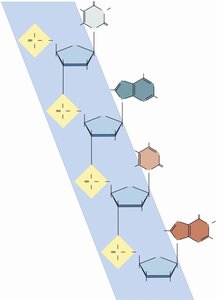
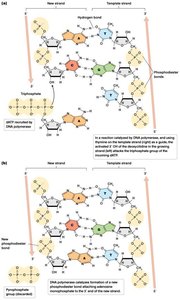
DNA strands have directionality, defined by the 5' (phosphate-terminated) and 3' (hydroxyl-terminated) ends. New nucleotides are always added to the 3' end during synthesis.
Phosphodiester bonds link nucleotides, forming the sugar-phosphate backbone.
Base pairing: Adenine pairs with thymine (A-T), and guanine pairs with cytosine (G-C).
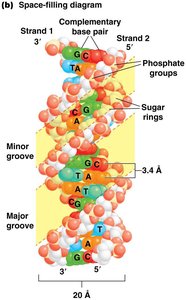
Double Helix: Secondary Structure
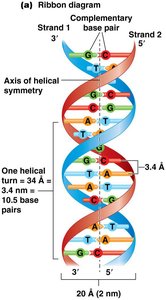
The most common form of DNA is the right-handed double helix (B-DNA). Two antiparallel strands are held together by hydrogen bonds between complementary bases and by base-stacking interactions.
Major and minor grooves are structural features important for protein-DNA interactions.
Helical parameters: 10 base pairs per turn, 3.4 nm per turn, 2 nm diameter.
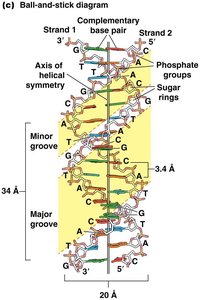
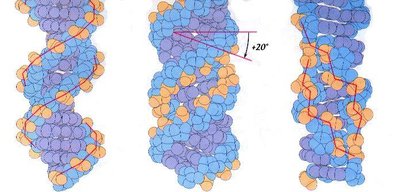
Alternative Forms of DNA
DNA can adopt several structural forms:
B-DNA: Most common, right-handed, 10.4 bp/turn, 2 nm diameter.
A-DNA: Right-handed, 11 bp/turn, wider diameter, found in some bacteriophages.
Z-DNA: Left-handed, zig-zag appearance, found near transcription start sites.
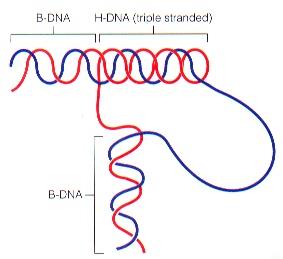
H-DNA: Triple-stranded DNA, may play a role in gene regulation.
Genomic Complexity and Packaging
Genome Definition and Processes
A genome is the complete set of genetic material in an organism. Key processes include:
Transcription: Synthesis of RNA and proteins.
Replication: Copying of DNA for cell division.
Segregation: Distribution of chromosomes during cell division.
Compaction: Packaging DNA to fit within the cell or nucleus.
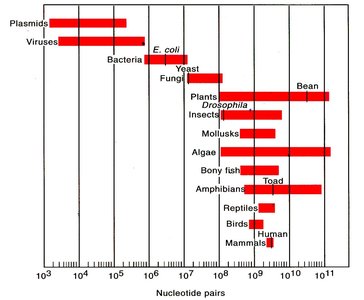
Comparison of Genome Complexity
Organism | Genome Size | Structure | Packing |
|---|---|---|---|
Viruses | 5–200 Kbp | Varied (ss/ds, RNA/DNA, circular/linear) | Protein shell (capsid) |
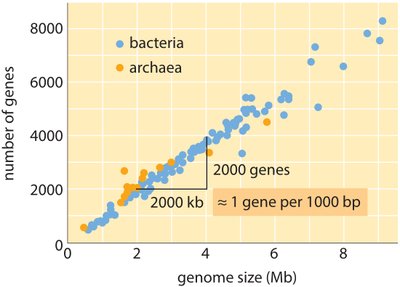
Bacteria | 1–10 Mbp (avg. 4 Mbp) | Circular, supercoiled | Looped, supercoiled, nucleoid |
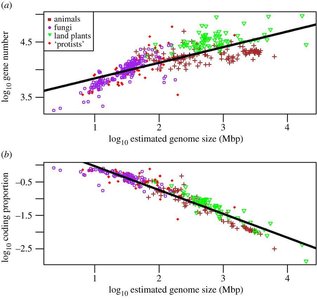
Eukaryotes | 5 Mbp – >100 Gbp | Linear, multiple chromosomes | Histones, chromatin, supercoiled |
Archaea | 500 Kbp – 6 Mbp | Circular | Histones, coiled |
Organization of Viral Genomes
Viral genomes are highly diverse, including double- or single-stranded RNA or DNA, and can be circular or linear. Viruses are not considered living organisms and rely on host cells for replication.
Non-enveloped (naked) viruses: Enclosed only in a protein shell (capsid).
Enveloped viruses: Enclosed in a capsid and an external membrane derived from the host cell.
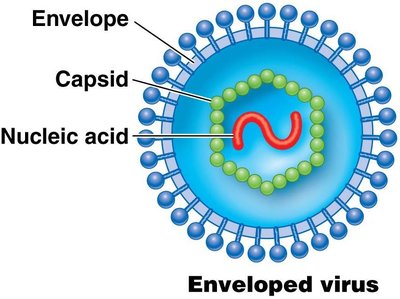
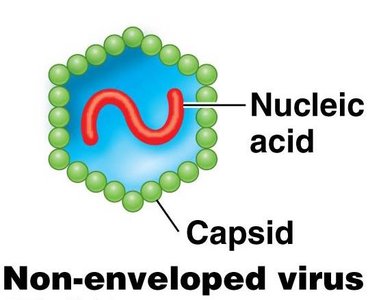
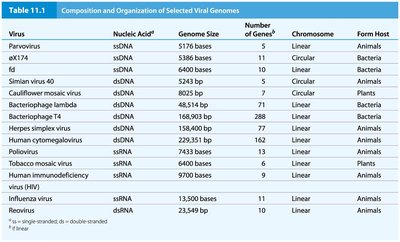
Table: Selected Viral Genomes
Virus | Nucleic Acid | Genome Size | Genes | Chromosome | Host |
|---|---|---|---|---|---|
Parvovirus | ssDNA | 5176 bases | 5 | Linear | Animals |
Bacteriophage T4 | dsDNA | 168,903 bp | 288 | Linear | Bacteria |
Herpes simplex virus | dsDNA | 152,000 bp | 80 | Linear | Animals |
Influenza virus | ssRNA | 13,500 bases | 11 | Linear | Animals |
Reovirus | dsRNA | 23,549 bp | 10 | Linear | Animals |
Retroviruses
Retroviruses are RNA viruses that encode reverse transcriptase, allowing them to convert their RNA genome into DNA, which integrates into the host genome. Not all RNA viruses are retroviruses, but all retroviruses are RNA viruses.
Bacterial Genomes
Bacteria typically have a single, circular, double-stranded DNA chromosome. They may also contain plasmids—small, autonomously replicating DNA molecules. The bacterial chromosome is densely packed in a region called the nucleoid, and supercoiling is a key feature of bacterial DNA organization.
Supercoiling: Controlled by DNA gyrase (introduces negative supercoils) and topoisomerase I (relaxes supercoils).
Gene density: Bacterial genomes are highly coding, with short intergenic regions and some repetitive sequences.
Eukaryotic Chromosomes and Chromatin Organization
Eukaryotic genomes are much larger and more complex than those of prokaryotes. DNA is packaged with histone proteins into chromatin, which undergoes multiple levels of compaction to fit within the nucleus.
Nucleosome: DNA wrapped around a histone octamer (2 each of H2A, H2B, H3, H4), forming the 10 nm fiber.
30 nm fiber: Nucleosomes coil into a solenoid structure, stabilized by histone H1.
300 nm fiber (radial loops): 30 nm fibers form loops attached to a protein scaffold.
Metaphase chromosome: Maximum compaction during cell division.
Chromatin can be classified as:
Euchromatin: Less condensed, transcriptionally active.
Heterochromatin: Highly condensed, transcriptionally inactive. Includes constitutive (permanently condensed, e.g., centromeres, telomeres) and facultative (variable condensation) forms.
Repetitive DNA in Eukaryotes
Retrotransposons: LTR and non-LTR elements (LINEs and SINEs) make up a large portion of the human genome.
Satellite DNA: Includes telomeres (TTAGGG repeats) and centromeres (alpha satellite DNA).
VNTRs: Variable number tandem repeats, used in DNA fingerprinting.
Archaeal Genomes and Chromosome Organization
Archaea possess a single, usually circular chromosome, often with plasmids. Their genome size ranges from 500 Kbp to 6 Mbp, with a high proportion of protein-coding DNA. Archaeal histones are homologous to eukaryotic histones and likely play a role in gene regulation, though their function is not fully understood.
Summary Table: DNA and Chromosome Organization Across Life
Domain | Genome Structure | Packing Proteins | Key Features |
|---|---|---|---|
Viruses | ss/ds RNA or DNA, linear/circular | Capsid proteins | Diverse, not cellular |
Bacteria | Circular dsDNA | HU, H-NS, SMC proteins | Supercoiled, nucleoid |
Eukaryotes | Linear dsDNA, multiple chromosomes | Histones, SMC proteins | Chromatin, complex packaging |
Archaea | Circular dsDNA | Histone-like proteins | Intermediate features |
Additional info: This guide covers content from Chapters 7 and 10, focusing on DNA structure, forms, and genome organization in various domains of life, as outlined in the provided course chapters.
