 Back
BackDNA and Chromosome Structure: Organization, Packaging, and Genome Complexity
Study Guide - Smart Notes

DNA Structure and Forms
Essential Characteristics of Hereditary Material
Hereditary material must fulfill several criteria to serve as the basis for genetic inheritance and evolution:
Localization: Found in the nucleus and as a component of chromosomes.
Stability: Maintains a stable form within cells.
Complexity: Contains sufficient information for structure, function, development, and reproduction.
Replication: Able to accurately replicate so daughter cells inherit identical information.
Mutability: Undergoes low rates of mutation, providing genetic variation for evolution.
Historical Discovery of DNA
DNA was first isolated in the 1860s by Friedrich Miescher from nuclei of white blood cells and salmon sperm. Early 20th-century experiments localized genes to chromosomes, and subsequent work identified DNA (not RNA) as the hereditary material.
Key contributors: Sutton, Boveri, Fleming, Avery, MacCleod, McCarty, Watson, Crick, Franklin, Wilkins.


Nucleotides: The Building Blocks of DNA
Nucleotide Structure
Nucleotides are the monomers of nucleic acids, forming the primary structure of DNA and RNA. Each nucleotide consists of three major parts:
Phosphate group
Pentose sugar: D-deoxyribose (DNA) or D-ribose (RNA)
Nitrogenous base: Purines (adenine, guanine) or Pyrimidines (cytosine, thymine, uracil)
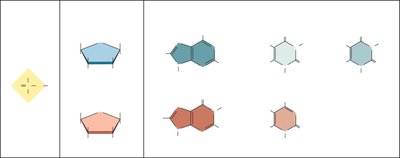
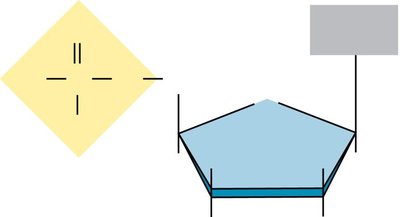
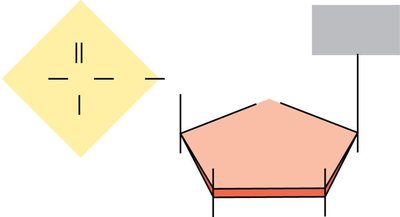
Key differences: DNA contains deoxyribose and thymine; RNA contains ribose and uracil.
The sugar carbons are numbered (1' to 5'), with the 1' carbon attached to the base, 2' distinguishing DNA from RNA, and 3' being the site for new nucleotide attachment.
Complementary Base Pairing
Base pairing is fundamental to the double helix structure:
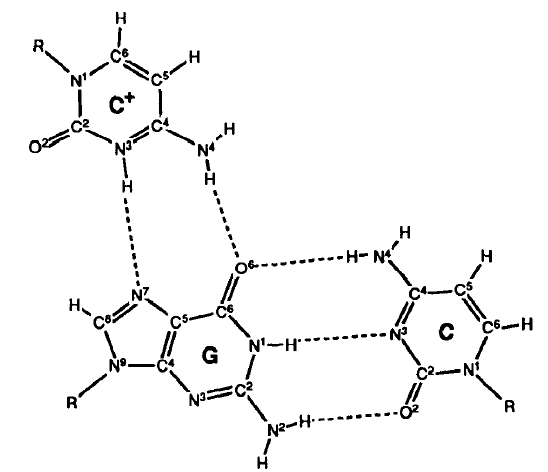
Adenine (A) pairs with Thymine (T) in DNA (2 hydrogen bonds).
Guanine (G) pairs with Cytosine (C) (3 hydrogen bonds).
In RNA, Uracil (U) replaces thymine.
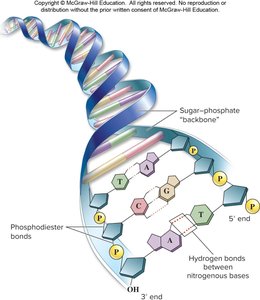
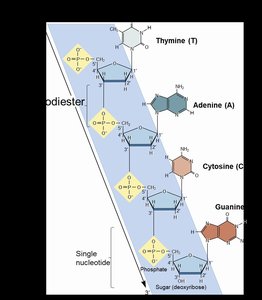
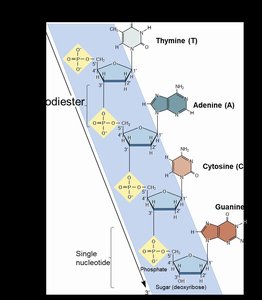
Single Strand of DNA
DNA strands have directionality based on the sugar's carbon atoms:
5' end: Terminates with a phosphate group.
3' end: Terminates with a hydroxyl (-OH) group.
New nucleotides are always added to the 3' end during synthesis.
Strands are connected by phosphodiester linkages.
DNA Double Helix and Higher-Order Structure
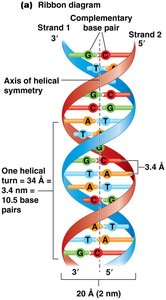
Features of the DNA Double Helix
The most common form of DNA is a right-handed double helix:
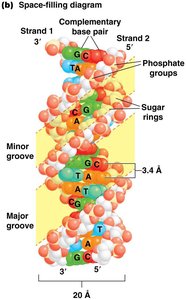
Each complete turn contains 10.5 base pairs (bp), spanning 3.4 nm.
Diameter is consistently 2 nm.
Strands are antiparallel (5' to 3' and 3' to 5').
Major and minor grooves are exposed to water and serve as protein interaction sites.
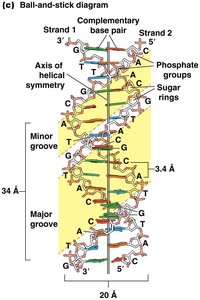
Forces stabilizing DNA: Hydrogen bonds, base-stacking/pi-stacking, hydrophobic and van der Waals interactions.
Alternative Forms of DNA
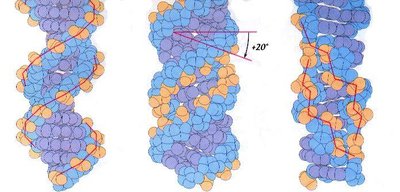
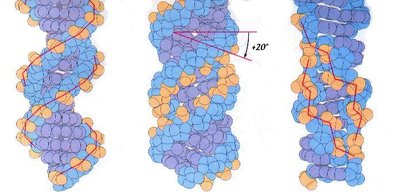
DNA can adopt several helical forms:
B-DNA: Most common, right-handed, 10.4 bp per turn, 2 nm diameter.
A-DNA: Right-handed, 11 bp per turn, wider diameter, occasional in cells.
Z-DNA: Left-handed, zig-zag appearance, found near transcription start sites.
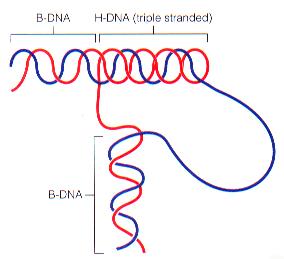
H-DNA: Triple-stranded DNA, occurs in short segments, may regulate gene expression.
Genomic Complexity and Packaging
Genome Definition and Processes
A genome is the complete set of genetic material in an organism. Four key processes are associated with genomes:
Synthesis: RNA and protein production, regulated by proteins.
Replication: DNA duplication, stabilized by proteins.
Segregation: Chromosome separation during cell division, aided by condensation.
Compaction: DNA packaging to fit within cells or nuclei.
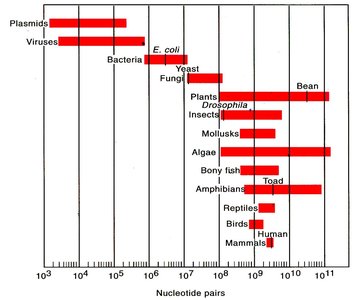
Comparison of Genome Complexity and Packaging
Genome size and packaging vary across biological domains:
Viruses: 5–200 Kbp, DNA or RNA, single/double-stranded, linear/circular.
Bacteria: 1–10 Mbp, circular chromosome, looped and supercoiled.
Archaea: 500 Kbp–6 Mbp, circular chromosome, coiled with histones.
Eukaryotes: 5 Mbp–>100 Gbp, linear chromosomes, coiled with histones, supercoiled.
Virus Structure and Genomes
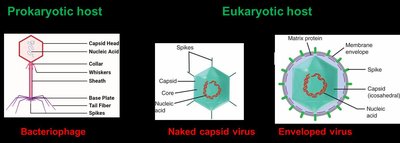
Virus Structure
Viruses range from 20 to 300 nm in diameter and consist of:
Internal nucleic acid core (DNA or RNA)
Protein coat (capsid)
Some have a lipid envelope derived from host cells
Surface spikes for receptor binding
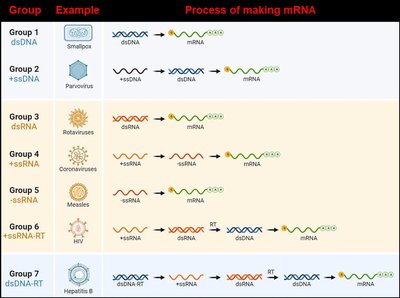
Organization of Viral Genomes
Viral genomes are highly diverse:
Double or single-stranded RNA or DNA
Circular or linear, single or multiple molecules
Viruses are obligate intracellular parasites, not cellular organisms
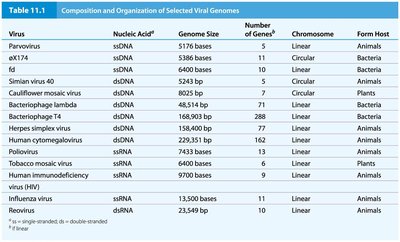
Virus | Nucleic Acid | Genome Size | Number of Genes | Chromosome | Form Host |
|---|---|---|---|---|---|
Parvovirus | ssDNA | 5176 bases | 5 | Linear | Animals |
Herpes simplex virus | dsDNA | 152,000 bp | 162 | Linear | Animals |
Influenza virus | ssRNA | 13,500 bases | 11 | Linear | Animals |
HIV | ssRNA | 9700 bases | 9 | Linear | Animals |
Bacteriophage lambda | dsDNA | 48,502 bp | 37 | Linear | Bacteria |
Cauliflower mosaic virus | dsDNA | 8245 bp | 7 | Circular | Plants |
Poliovirus | ssRNA | 7,500 bases | 11 | Linear | Animals |
Additional info: Table includes selected viral genomes, showing diversity in nucleic acid type, size, and host. |
Bacterial Genomes and Chromosome Packaging
Bacterial Genome Organization
Bacteria typically have a haploid genome with a single circular chromosome of double-stranded DNA. They may also possess plasmids—autonomously replicating DNA molecules.
Genomes carry thousands of protein-coding genes with short intergenic regions.
Repetitive sequences may be interspersed.
Nucleoid: Dense, non-membrane region where the chromosome is supercoiled.
Bacterial Chromosome Condensation
Chromosome packaging involves:
Loops anchored by proteins (HU, H-NS, SMC proteins)
Supercoiling: DNA can be negatively or positively supercoiled, with negative supercoiling being most common.
Supercoiling is regulated by DNA gyrase/topoisomerase II (introduces supercoils) and topoisomerase I (relaxes supercoils).
Eukaryotic Chromosome Organization and Packaging
Chromatin Structure
Eukaryotic chromosomes are organized into chromatin—a complex of DNA and proteins (primarily histones). Chromatin condensation varies:
Euchromatin: Less condensed, actively expressed genes.
Heterochromatin: More condensed, fewer expressed genes.
Facultative heterochromatin: Variable condensation, related to gene transcription.
Constitutive heterochromatin: Permanently condensed, found in centromeres and telomeres.
Levels of Chromatin Packaging
Nucleosome (10 nm fiber): dsDNA wrapped around histone octamer (H2A, H2B, H3, H4).
30 nm fiber: Coiling of nucleosomes into solenoid structure, stabilized by H1 histone.
300 nm fiber (radial loops): Loops of 30 nm fibers attached to chromosome scaffold.
700 nm chromatid: Further compaction mediated by condensin proteins.
Chromatin structure changes through the cell cycle, becoming maximally condensed during metaphase.
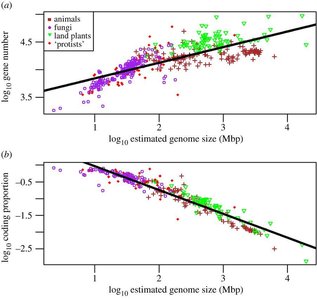
Eukaryotic Genome Complexity
Genome Size and Gene Number
Eukaryotic genome size varies greatly and does not always correlate with organismal complexity (C-value paradox). Gene number is more closely associated with complexity, but exceptions exist (e.g., humans vs. plants).
Human Genome Composition
Approximately 3.2 billion base pairs
~20,000 to 25,000 protein-coding genes
~40% unique sequence DNA, ~58% repetitive sequence DNA
Repetitive Sequence DNA
Repetitive DNA is hierarchically classified:
Transposons: DNA segments that move within the genome.
Retrotransposons: Use RNA intermediates to copy themselves.
LTR retrotransposons: Endogenous retroviruses (ERVs), flanked by long terminal repeats.
Non-LTR retrotransposons: LINEs (autonomous, code for reverse transcriptase), SINEs (non-autonomous, rely on LINEs).
Alu SINEs: Unique to primates, 1 million copies in humans (~10% of genome).
Satellite DNA: Tandem Repeats
Microsatellites: 1–6 bp, repeated many times.
Minisatellites: 10–100 bp, repeated 5–50 times.
Macrosatellites: Several thousand bp, much larger repeats.
VNTRs: Variable number tandem repeats, used in DNA fingerprinting.
Telomeres: Repetitive sequence (TTAGGG in vertebrates).
Centromeres: Imperfect satellite repeats, most common is alpha satellite DNA (171 bp).
Archaeal Chromosomes and Histones
Archaeal Genome Organization
Archaea have a single, usually circular chromosome, often with plasmids. Genome size ranges from 500 Kbp to 6 Mbp, with ~87% protein-coding sequences. Some repetitive and intergenic regions are present.
Archaeal Histones and Nucleosomes
Archaeal histone proteins show homology to eukaryotic histones and form nucleosome-like complexes. Their functional role is not fully understood but likely involves regulation of gene expression.
Summary Table: Genome Complexity Across Domains
Domain | Genome Size | Chromosome Structure | Packaging |
|---|---|---|---|
Virus | 5–200 Kbp | Linear/circular, DNA/RNA | Capsid, envelope |
Bacteria | 1–10 Mbp | Circular, dsDNA | Supercoiling, nucleoid |
Archaea | 500 Kbp–6 Mbp | Circular, dsDNA | Histones, supercoiling |
Eukaryotes | 5 Mbp–>100 Gbp | Linear, multiple chromosomes | Histones, chromatin, supercoiling |
