 Back
BackDNA Organization in Chromosomes
Study Guide - Smart Notes

Chapter 12. DNA Organization in Chromosomes
I. Viral & Bacterial Chromosomes
Genetic material in viruses and bacteria is organized differently from that in eukaryotes, reflecting their unique replication and packaging requirements.
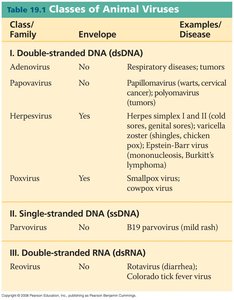
1. Viral Chromosomes
Nature of Viral Genomes: Viral chromosomes can be composed of either DNA or RNA, which may be single- or double-stranded, and can exist in circular or linear forms.
Size and Packaging: Viral DNA is highly compacted and functionally inert when packed. For example, λ-DNA is a double-stranded, linear molecule of 48,500 base pairs, while T2 phage DNA can be as long as 52 μm when released.
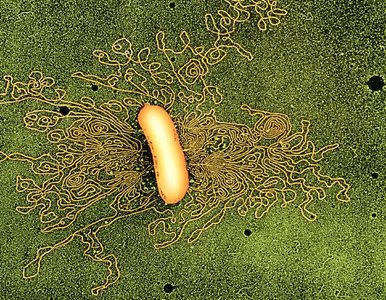
2. Bacterial Chromosomes
Structure: Bacterial chromosomes are typically double-stranded DNA, compacted into a region called the nucleoid. The E. coli genome is about 1,000 μm in length.
DNA-Binding Proteins: Proteins such as HU and H1 help organize and compact the bacterial chromosome.
Supercoiling: Most bacterial DNA is slightly underwound and supercoiled, which aids in compaction and accessibility for replication and transcription.
Topoisomerases: These enzymes cut one or both DNA strands to wind or unwind the helix, then reseal the ends, regulating DNA topology.
II. Mitochondrial & Chloroplast DNA
Mitochondria and chloroplasts contain their own DNA, which is distinct from nuclear DNA and reflects their evolutionary origins.
1. Mitochondrial (mt) DNA
Inheritance: Traits encoded by mitochondrial or chloroplast DNA are typically maternally inherited.
Structure: Mitochondrial DNA is circular, double-stranded, and protein-free. It replicates semi-conservatively.
Size Variation: Plant mtDNA (~100 kb) is much larger than animal mtDNA (5–20 kb).
Gene Content: Few or no gene repetitions; many enzymes required for mtDNA replication are encoded by nuclear genes.
Ribosomes: Mitochondrial ribosomes vary in size (55–80S).
2. Chloroplast (cp) DNA
Structure: Chloroplast DNA is circular, double-stranded, protein-free, and larger than mtDNA (~120–195 kb). It replicates semi-conservatively and lacks 5-methylcytosine.
Gene Content: Codes for rRNAs (5S, 16S, 23S), at least 25 tRNAs, ribosomal proteins, and photosynthesis-related proteins (e.g., rubisco).
Copy Number: Each chloroplast contains multiple copies of cpDNA (up to 75 copies).
Evolutionary Note: Both mtDNA and cpDNA resemble bacterial genomes, supporting the endosymbiotic theory of organelle origin.
III. Organization of Chromatin in Eukaryotes
Eukaryotic DNA is organized into chromatin, a complex of DNA and proteins, which allows for efficient packaging and regulation of gene expression.
1. Specialized Chromosomes
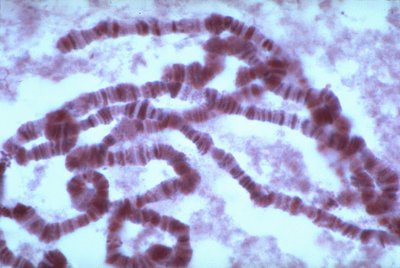
Polytene Chromosomes: Found in salivary gland cells of insect larvae (order Diptera), these chromosomes are large, have distinctive banding patterns, and are composed of many DNA strands formed by endomitosis (multiple rounds of DNA replication without cell division).
Banding: Bands are rich in DNA and may contain multiple genes. Puff regions indicate high gene activity.
Lampbrush Chromosomes: Large chromosomes with extensive DNA looping, found in oocytes of vertebrates and some insect spermatocytes. The loops are sites of active transcription during prophase I of meiosis.
2. Chromatin Structure & Nucleosomes
Chromatin: DNA is complexed with histone proteins to form chromatin. The degree of compaction changes during the cell cycle.
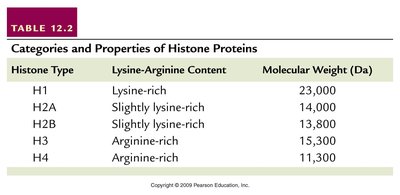
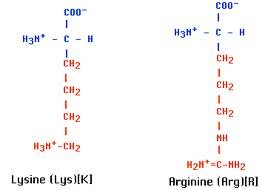
Histones: Five main types (H1, H2A, H2B, H3, H4), all positively charged due to lysine and arginine content. They play a structural role in chromatin condensation.
Histone Type | Lysine-Arginine Content | Molecular Weight (Da) |
|---|---|---|
H1 | Lysine-rich | 23,000 |
H2A | Slightly lysine-rich | 14,000 |
H2B | Slightly lysine-rich | 13,800 |
H3 | Arginine-rich | 15,300 |
H4 | Arginine-rich | 11,300 |
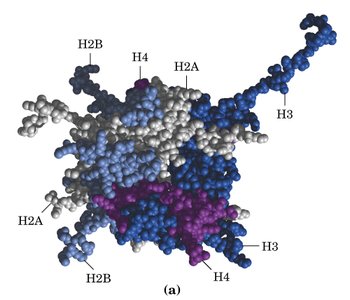
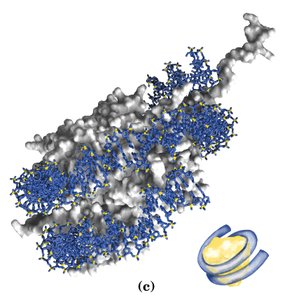
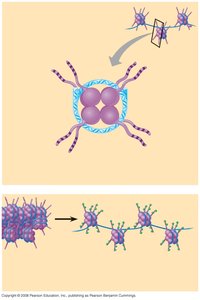
Nucleosomes: The basic unit of chromatin, consisting of an octamer of histones (two each of H2A, H2B, H3, H4) with ~146 bp of DNA wrapped around it. H1 and ~50 bp of linker DNA connect adjacent nucleosomes.
Chromatin Fiber: Nucleosomes are organized into an 11 nm fiber, which is further folded into a 30 nm solenoid structure, and ultimately into highly condensed mitotic chromosomes.
3. Chromatin Remodeling & Histone Modifications
Chromatin Remodeling: Necessary for DNA accessibility during replication and gene expression. Involves changes in nucleosome position and structure.
Histone Modifications: Acetylation (loosens chromatin, promotes gene expression), methylation (condenses chromatin, represses genes), and phosphorylation (can loosen chromatin if adjacent to methylation).
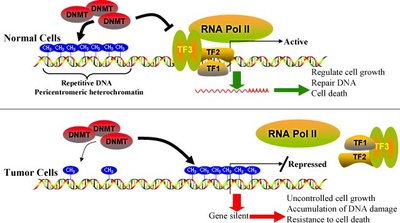
4. Euchromatin vs. Heterochromatin
Euchromatin: Uncoiled, lightly stained, and transcriptionally active.
Heterochromatin: Densely packed, heavily stained, and generally inactive. Replicates later in S phase, maintains chromosome integrity, and forms centromeres, telomeres, Barr bodies, and much of the Y chromosome.
Positional Effect: Genes relocated near heterochromatin may be silenced.
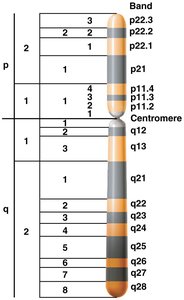
Chromosome Banding: Techniques such as C-banding (centromeres) and G-banding (differential staining) reveal chromosomal regions and complexity.
IV. DNA Sequence Organization in Eukaryotes
Eukaryotic genomes contain a variety of repetitive and unique DNA sequences, many of which have structural or regulatory roles.
1. Repetitive DNA & Satellite DNA
Repetitive DNA: Includes highly and moderately repetitive sequences, which may be non-genic or contain multiple gene copies. Up to 40% of the human genome is repetitive DNA.
Satellite DNA: Highly repetitive, short sequences with lower density than main band DNA, often found in centromeric regions.
2. Centromeric & Telomeric DNA Sequences
Centromeres: Primary constrictions on chromosomes, essential for chromosome segregation during mitosis and meiosis. The centromeric DNA (CEN) is about 225 bp, divided into three regions (I, II, III), with region III being critical for kinetochore binding.
Telomeres: Repetitive sequences at chromosome ends (e.g., TT(TT)GGGG), added by telomerase, protect chromosome integrity and prevent gene loss during replication.
3. VNTRs, Dinucleotide Repeats, SINEs & LINEs
VNTRs (Variable Number Tandem Repeats): 15–100 bp sequences repeated in clusters, highly variable among individuals, used in DNA fingerprinting.
Dinucleotide Repeats (Microsatellites): Short (e.g., (CA)n) repeats dispersed throughout the genome, also variable and useful as molecular markers.
SINEs (Short Interspersed Elements): <500 bp, present in high copy numbers (e.g., Alu family), constitute >5% of the human genome.
LINEs (Long Interspersed Elements): 5,000–7,500 bp, retrotransposons (e.g., L1 family), about 100,000 copies in the human genome. SINEs and LINEs together make up ~30% of the genome.
Protein-Coding Genes: Only 2–10% of the eukaryotic genome encodes proteins; the rest is noncoding or repetitive DNA.
