 Back
BackDNA Organization in Chromosomes: Structure, Packing, and Chromatin Remodeling
Study Guide - Smart Notes

DNA Organization in Chromosomes
Forms of DNA Helices
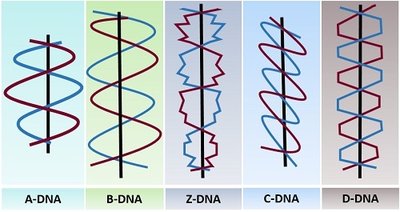
DNA can adopt several structural forms depending on environmental conditions and sequence composition. These forms differ in their handedness, helical pitch, and base pair arrangements.
A-DNA: Right-handed helix, found under dehydrating conditions, more compact than B-DNA.
B-DNA: The most common form in cells, right-handed helix, standard Watson-Crick structure.
C-DNA: Right-handed, occurs under greater dehydration than A-DNA.
D-DNA: Right-handed, lacks guanine bases.
Z-DNA: Left-handed helix, found in synthetic DNA with only G-C pairs, zigzag backbone.
Viral and Bacterial Chromosomes
Viral and bacterial chromosomes are typically simpler than those of eukaryotes, often consisting of a single nucleic acid molecule with minimal associated proteins.
Viral chromosomes: Can be DNA or RNA, single or double stranded, circular or linear. Genetic material is inert until released into a host cell.
Bacterial chromosomes: Usually double-stranded DNA (dsDNA), compacted into a region called the nucleoid. Associated with histone-like proteins (HU and H-NS) that help fold and bend DNA.

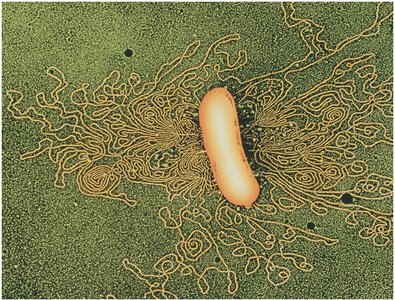
DNA Packing in Viruses and Bacteria
Both viruses and bacteria must package long DNA molecules into small volumes. This is achieved through supercoiling and association with DNA-binding proteins.
Supercoiling: DNA is twisted upon itself to reduce its volume and increase density.
DNA-binding proteins: In bacteria, HU and H-NS proteins contain positively charged amino acids that bind to the negatively charged DNA backbone, facilitating compaction.
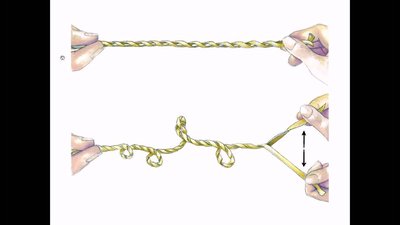
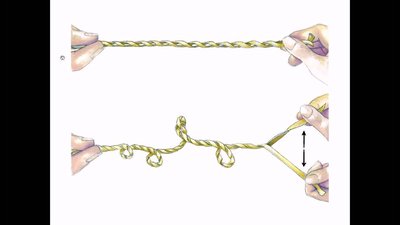
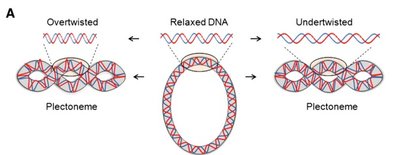
Supercoiling of DNA
Supercoiling is a process by which DNA twists upon itself, allowing for tighter packing and compaction. It occurs in both prokaryotes and eukaryotes, especially in circular chromosomes.
Positive supercoiling: DNA is overrotated, helix twists in the same direction as the helix.
Negative supercoiling: DNA is underrotated, helix twists in the opposite direction.
Advantages: Tighter packing, increased density, and compaction of DNA.
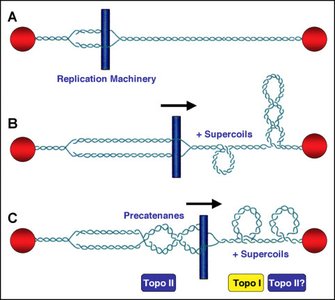
Topoisomerases
Topoisomerases are enzymes that regulate the supercoiling of DNA by cutting and resealing DNA strands.
Type I topoisomerase: Cleaves one strand of the DNA helix.
Type II topoisomerase: Cleaves both strands of the DNA helix.
These enzymes wind or unwind the helix before resealing the ends, essential for DNA replication and transcription.
Eukaryotic DNA Organization
Complexity and Packing
Eukaryotic chromosomes contain much more DNA per chromosome and are associated with many proteins, primarily histones. The DNA is highly compacted to fit within the small nuclear volume.
Human DNA: ~2 meters of DNA packed into a nucleus 5-10 μm in diameter.
Chromatin: The complex of DNA, histones, and non-histone proteins that make up uncoiled chromosomes.
Histones and Nucleosomes
Histones are basic proteins rich in lysine and arginine, which bind to the negatively charged DNA backbone. Nucleosomes are the fundamental units of chromatin structure.
Nucleosome: Segment of DNA (147 bp) wound around an octamer of histone proteins (2 copies each of H2A, H2B, H3, H4).
H1 histone: Binds to linker DNA (53 bp) between nucleosomes, involved in higher-order chromatin structure.
Histone Types and Properties
Histone Type | Lysine-Arginine Content |
|---|---|
H1 | Lysine-rich |
H2A | Slightly lysine-rich |
H2B | Slightly lysine-rich |
H3 | Arginine-rich |
H4 | Arginine-rich |
Levels of DNA Packing
DNA is packed into chromosomes through a hierarchical series of structures:
Nucleosome: 11 nm fiber, DNA reduced to 1/3 original length.
30 nm fiber: Bundle of nucleosomes coiled and stacked, separated by H1 histone, 6-fold increase in compaction.
300 nm looped domain: 30 nm fibers folded into looped domains.
Chromatid: Coiled chromatin fibers compacted into chromosome arms during metaphase.
Chromatin Remodeling
Chromatin must be dynamically remodeled to allow access to DNA for replication and transcription. This involves chemical modifications of histone tails.
Acetylation: Addition of acetyl groups to lysine residues, neutralizing positive charge and loosening DNA-histone interaction. Enzyme: histone acetyltransferase (HAT).
Methylation: Addition of methyl groups to arginine and lysine, often associated with gene activation. Enzyme: methyltransferase.
Phosphorylation: Addition of phosphate groups to serine and histidine, introducing negative charge. Enzyme: kinase.
All modifications are reversible and regulate gene expression.
Euchromatin and Heterochromatin
Eukaryotic chromosomes are not structurally uniform:
Euchromatin: Uncoiled, transcriptionally active regions.
Heterochromatin: Condensed, transcriptionally inactive regions, often lacking genes or containing repressed genes.
Telomeres: Maintain chromosome integrity.
Centromeres: Essential for chromosome movement during cell division.
Chromosome Banding
Chromosome banding techniques (C-banding, G-banding) are used to visualize specific regions of chromosomes, identify abnormalities, and study structural rearrangements.
C-banding: Stains centromeric heterochromatin.
G-banding: Produces a unique banding pattern for each chromosome, useful for detecting translocations, deletions, insertions, and inversions.
Repetitive DNA in Eukaryotic Genomes
Types of Repetitive DNA
Only a small proportion of the eukaryotic genome encodes functional genes. The majority consists of various types of repetitive DNA.
Tandem repeats: Short sequences repeated in tandem, main component of centromeres, can be up to several megabases.
Variable number tandem repeats (VNTRs): Minisatellites, 15-100 bp long, found within and between genes.
Short tandem repeats (STRs): Microsatellites, di-, tri-, tetra-, pentanucleotide repeats, e.g., (CA)n in humans.
Interspersed repeats: Scattered throughout the genome, transposable, includes SINEs (Alu family, 200-300 bp) and LINEs (L1 family, 6400 bp).
Pseudogenes: Single-copy non-coding sequences, evolutionary vestiges of duplicated genes, not transcribed.
Proportion of Functional Genes
Human genome: ~2% encodes functional genes (~20,000 genes).
Sea urchin: ~10% of genome encodes proteins.
Drosophila: 5-10% of genome encodes proteins.
~50% of human DNA is repetitive.
Summary
Phage and bacterial chromosomes are short, circular DNA free of associated proteins.
Eukaryotic DNA is organized into different levels of compaction within the nucleus.
Only a small part of eukaryotic genomes contains functional genes; the majority is repetitive DNA.
