 Back
BackDNA Packaging in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

DNA Packaging
Genome Size and the C Value Paradox
The C value refers to the total amount of DNA in the haploid genome of an organism. This value varies widely among species and does not correlate with the organism's structural or organizational complexity, a phenomenon known as the C value paradox.
Examples: Genome sizes range from 1,759 base pairs in Circovirus to 290,000,000,000 base pairs in Amoeba proteus.
Key Point: More complex organisms do not necessarily have more DNA.
Genome Packaging: The Challenge
All organisms must efficiently package their DNA to fit within the confines of a cell. The human genome, for example, would stretch 2.3 meters if fully extended, yet it fits inside a microscopic nucleus.
Genome: The complete set of chromosomes containing all DNA of an organism or organelle.
Packaging is essential for both prokaryotes and eukaryotes.
DNA Packaging in Prokaryotes
Prokaryotic Chromosome Structure
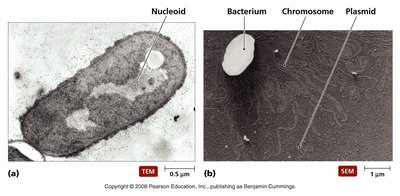
Most prokaryotes possess a single, double-stranded, circular DNA chromosome, often accompanied by smaller DNA molecules called plasmids. The DNA is located in the nucleoid region of the cell.
Nucleoid: Irregularly-shaped region within the cell where the chromosome is found.
Plasmids: Small, circular DNA molecules that replicate independently of the chromosome.
Supercoiling and DNA Loops
Prokaryotic DNA is much longer than the cell itself, necessitating compact packaging. Two main strategies are used:
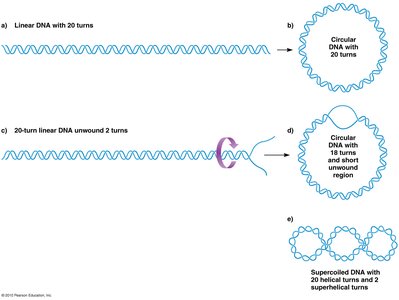
Supercoiling: The DNA double helix is twisted upon itself, introducing tension and compacting the molecule. This process is regulated by enzymes called topoisomerases.
DNA Loops: The chromosome forms loops, each attached at its base, further compacting the DNA.
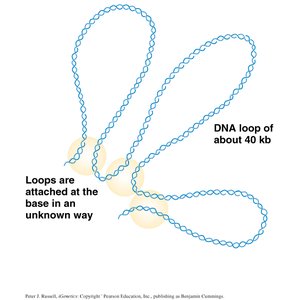
Topoisomerases: Enzymes that cut and rejoin DNA strands to relieve or introduce supercoils.
Function: Supercoiling and looping allow the genome to fit inside the small prokaryotic cell.
DNA Packaging in Eukaryotes
Histones and Nucleosomes
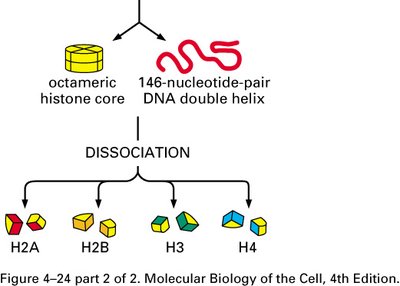
Eukaryotic DNA is packaged with proteins called histones to form a complex structure known as chromatin. There are five main classes of histones: H1, H2A, H2B, H3, and H4. The core histones (H2A, H2B, H3, H4) form an octamer around which DNA is wrapped.
Histones: Small, basic proteins with a positive charge that bind to negatively charged DNA.
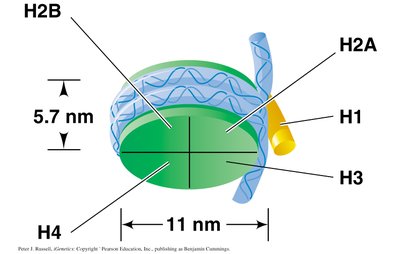
Nucleosome: The fundamental unit of chromatin, consisting of 146 base pairs of DNA wrapped twice around a histone octamer.
Linker DNA: Short stretches of DNA (8–80 bp) connecting adjacent nucleosomes.
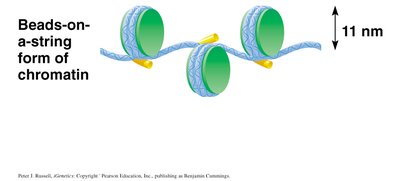
Chromatin Structure: Beads-on-a-String
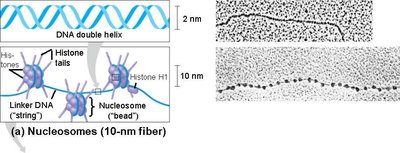
Under low magnification, chromatin appears as a series of "beads on a string," where each bead is a nucleosome. This 11 nm fiber is the first level of DNA compaction in eukaryotes.
Beads-on-a-string: Nucleosomes connected by linker DNA, forming a 10–11 nm fiber.
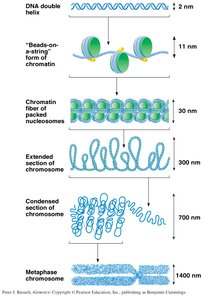
Higher-Order Chromatin Structure
Chromatin undergoes further compaction through several hierarchical levels, ultimately forming the highly condensed metaphase chromosome seen during cell division.
30 nm fiber: Nucleosomes are packed into a thicker fiber.
Looped domains: 30 nm fibers form loops attached to a protein scaffold.
Condensed chromatin: Loops are further coiled and folded to form the metaphase chromosome.
Chromatin States: Euchromatin and Heterochromatin
Chromatin exists in two main states:
Euchromatin: Less condensed, transcriptionally active, and accessible to regulatory proteins.
Heterochromatin: Highly condensed, transcriptionally inactive, and gene-poor.
Regulation of Chromatin Compaction
The degree of chromatin compaction is dynamically regulated by chemical modifications to histone tails. These modifications influence gene expression:
Acetylation: Addition of acetyl groups relaxes chromatin, allowing gene expression.
Methylation: Addition of methyl groups compacts chromatin, silencing genes.
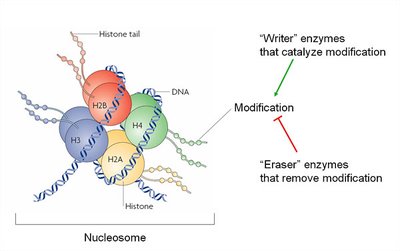
Enzymes: "Writer" enzymes add modifications, while "eraser" enzymes remove them.
Example: During interphase, chromatin is mostly euchromatin, allowing active transcription. During prophase of mitosis, chromatin condenses into heterochromatin to form visible chromosomes.
Additional info: The dynamic regulation of chromatin structure is essential for processes such as DNA replication, repair, and gene regulation, and is a major focus of epigenetics research.
