 Back
BackDNA Repair and Transposition: Mechanisms and Implications in Genetics
Study Guide - Smart Notes

DNA Repair and Transposition
Overview
This study guide covers the molecular mechanisms of DNA repair and transposition, their roles in maintaining genetic integrity, and their implications for human disease. The content is directly relevant to genetics, specifically addressing gene mutation, DNA repair, and transposable elements as outlined in college-level genetics courses.
Ames Test for Mutagenicity
Principle and Procedure
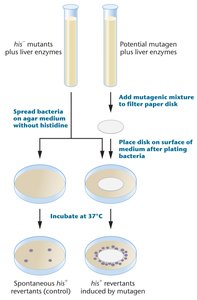
The Ames test is a biological assay used to assess the mutagenic potential of chemical compounds. It utilizes strains of Salmonella typhimurium that are his- auxotrophs (unable to synthesize histidine). Exposure to a mutagen may cause a reversion mutation, allowing bacteria to grow without histidine.
his- mutants are plated on agar lacking histidine.
Bacteria are exposed to a potential mutagen.
Reversion to his+ indicates mutagenicity.
Mutagenic compounds induce reversion at a rate much higher than spontaneous mutation.
Example: A chemical that causes a high frequency of his+ revertants is considered mutagenic.
Mutations Causing Human Disease
Types of Mutations and Associated Disorders
Single-gene mutations can cause a variety of human genetic disorders. The molecular change determines the disease phenotype.
Missense mutation: Alters a single amino acid (e.g., Achondroplasia: Glycine to arginine in FGFR3 gene).
Nonsense mutation: Introduces a premature stop codon (e.g., Marfan syndrome: Tyrosine to stop codon in fibrillin-1 gene).
Insertion: Adds extra nucleotides (e.g., Familial hypercholesterolemia: Short insertions in LDLR gene).
Deletion: Removes nucleotides (e.g., Cystic fibrosis: Three-base-pair deletion in CFTR gene).
Trinucleotide repeat expansion: Increases repeat number (e.g., Huntington disease: >40 CAG repeats in Huntingtin gene).
β-Thalassemia: Mutations in the HBB Gene
β-Thalassemia is caused by diverse mutations in the HBB gene affecting transcription, translation, splicing, and stability of mRNA.
Gene Region Affected | Number of Mutations Known | Description |
|---|---|---|
5′ upstream region | 22 | Single base-pair mutations decrease transcription (e.g., TATA box mutation). |
mRNA CAP site | 1 | Mutation at +1 reduces mRNA levels. |
5′ untranslated region | 3 | Mutations decrease transcription/translation. |
ATG translation initiation codon | 7 | Mutations prevent translation. |
Exons 1, 2, 3 | 36 | Missense/nonsense mutations, abnormal splicing. |
Introns 1, 2 | 38 | Splicing defects, abnormal splice sites. |
Polyadenylation site | 6 | Reduced mRNA stability. |
Throughout HBB gene | >100 | Insertions, deletions, duplications, frameshifts. |
DNA Repair Systems
General Features
DNA repair systems are essential for maintaining genetic integrity by counteracting spontaneous and induced DNA damage. Defective repair mechanisms are linked to genetic diseases and cancer susceptibility.
Present in all organisms
Counteract genetic damage
Maintain genome stability
Mismatch Repair (MMR)
MMR corrects base pair mismatches not fixed by DNA polymerase proofreading. The process involves endonuclease, exonuclease, DNA polymerase, and ligase.
Defective MMR is associated with hereditary colon cancer.
Post-Replication Repair
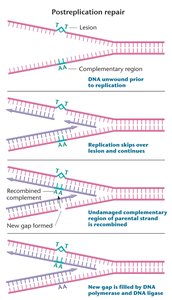
When DNA damage blocks replication, recombination with the undamaged template repairs the gap, allowing replication to finish.
Uses recombination to fill gaps left by DNA polymerase.
Critical for cell survival when DNA is damaged.
SOS Repair
SOS repair is a bacterial response to extensive DNA damage. It induces expression of multiple genes, including error-prone DNA polymerases, allowing survival at the cost of increased mutation.
Induced by unrepaired mismatches
Error-prone, allows cell survival
Photoactivation Repair
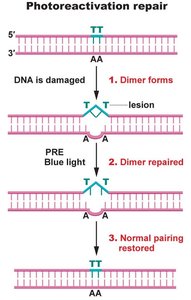
Photoactivation repair uses the photoreactivation enzyme (PRE) to cleave thymidine dimers formed by UV radiation. PRE is activated by blue light and reverses UV damage. This mechanism is not found in humans.
Cleaves thymidine dimers
Activated by blue light
Present in many organisms, absent in humans
Base Excision Repair
Base excision repair corrects minor defects affecting single bases, such as the presence of uracil in DNA. DNA glycosylase removes the base, AP endonuclease cleaves the backbone, and DNA polymerase and ligase repair the gap.
Human cells have multiple DNA glycosylases for different types of damage.
Nucleotide Excision Repair (NER)
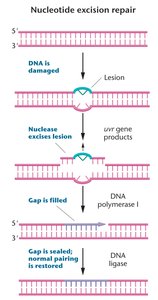
NER repairs bulky lesions that distort the DNA helix and impede replication/transcription. A region of 13-28 bp is removed, and repair is completed by DNA polymerase and ligase.
Critical for removing UV-induced damage
Defects in NER cause Xeroderma Pigmentosum (XP)
Xeroderma Pigmentosum (XP)
XP is a human disorder characterized by dry, pigmented skin and a predisposition to skin cancer. It is caused by defects in NER genes, revealed through somatic cell hybridization studies.
NER involves damage recognition, helicases, nucleases, and DNA polymerases.
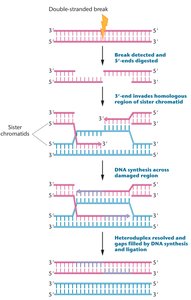
Double-Strand Break Repair
Double-strand breaks are repaired by homologous recombination or nonhomologous end joining. Defects in these systems result in X-ray hypersensitivity and predisposition to hereditary breast and ovarian cancers.
Homologous recombination uses sister chromatids as templates.
Nonhomologous end joining directly ligates broken ends.
Transposable Elements (Transposons)
Discovery and Significance
Transposable elements are DNA sequences that can move within the genome. Discovered by Barbara McClintock in 1950, they play a role in genome evolution and can cause mutations.
Movement can disrupt gene function
Example: Pigmented spots in corn kernels due to transposon excision

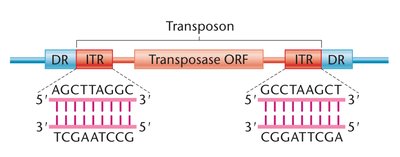
DNA Transposons
DNA transposons move without an RNA intermediate. They have inverted terminal repeats (ITRs) recognized by transposase enzyme and use a cut-and-paste mechanism.
ITRs are essential for transposase recognition
Cut-and-paste mechanism
Types of DNA Transposons
There are two main types:
Autonomous elements: Can transpose independently; mutations are unstable.
Nonautonomous elements: Require autonomous elements for transposition; insertions are stable unless an autonomous element is present.
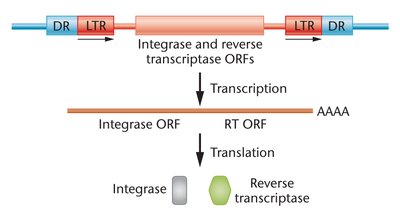
Retrotransposons
Retrotransposons move within the genome using an RNA intermediate and a copy-and-paste mechanism. They resemble retroviruses and are abundant in the human genome.
LINEs (Long Interspersed Nuclear Elements): 1-6 kb, 850,000 copies
SINEs (Short Interspersed Nuclear Elements): 100-500 bp, 1,500,000 copies
Active transposition can cause disease
Summary Table: DNA Repair Mechanisms
Repair Mechanism | Type of Damage | Key Enzymes | Organisms |
|---|---|---|---|
Mismatch Repair | Base pair mismatches | Endonuclease, exonuclease, DNA pol, ligase | All |
Post-Replication Repair | Replication-blocking lesions | Recombinase, DNA pol, ligase | All |
SOS Repair | Extensive DNA damage | Error-prone DNA pol | Bacteria |
Photoactivation Repair | Thymidine dimers | Photoreactivation enzyme | Many, not humans |
Base Excision Repair | Single base defects | DNA glycosylase, AP endonuclease, DNA pol, ligase | All |
Nucleotide Excision Repair | Bulky lesions | Damage recognition, helicase, nuclease, DNA pol, ligase | All |
Double-Strand Break Repair | Double-strand breaks | Recombinase, DNA pol, ligase | All |
Additional info: The notes expand on brief points from the original slides, providing definitions, examples, and context for each repair mechanism and transposon type. Tables are recreated for clarity and completeness.
