 Back
BackDNA Repair Mechanisms: A Comprehensive Study Guide for Genetics Students
Study Guide - Smart Notes

DNA Repair Mechanisms
Overview of DNA Repair
Cells possess multiple mechanisms to correct mutations and maintain genomic integrity. DNA repair is essential for preventing the accumulation of mutations that can lead to diseases such as cancer. The choice of repair pathway depends on the type of DNA damage encountered, including small base changes, bulky distortions, mismatches, and strand breaks.
Direct Repair: Reverses chemical damage without removing nucleotides.
Base Excision Repair (BER): Removes non-bulky, abnormal bases.
Nucleotide Excision Repair (NER): Excises bulky lesions that distort the DNA helix.
Mismatch Repair (MMR): Corrects base-pair mismatches post-replication.
Double-Strand Break Repair: Repairs breaks in both DNA strands via NHEJ or HRR.
Proofreading by DNA Polymerase
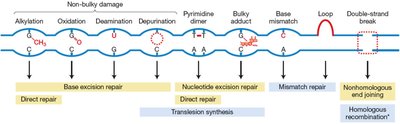
During DNA replication, most errors are corrected by the proofreading activity of DNA polymerase. The enzyme checks each newly added base for correct pairing and removes incorrect nucleotides using its 3’ exonuclease activity.
Proofreading: DNA polymerase reads the newly added base before proceeding.
Correction: Incorrect bases are excised and replaced with the correct nucleotide.
Types of DNA Damage and Corresponding Repair Pathways
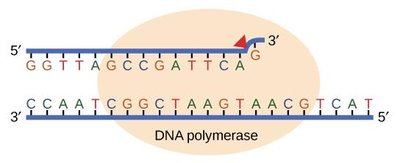
Classification of DNA Damage
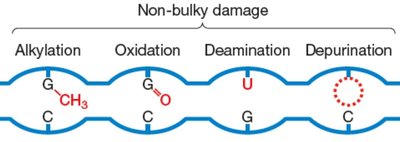
DNA damage can be classified as non-bulky (e.g., alkylation, oxidation, deamination, depurination), bulky (e.g., pyrimidine dimers, bulky adducts), mismatches, loops, and double-strand breaks. Each type is addressed by a specific repair mechanism.
Non-bulky damage: BER or direct repair
Bulky damage: NER
Base mismatch: MMR
Double-strand breaks: NHEJ or HRR

Direct Repair Mechanisms
Photoreactivation and Alkylation Repair
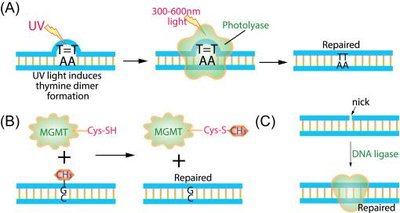
Direct repair mechanisms restore damaged bases to their original form without excising nucleotides or breaking the DNA backbone. These are highly specific and enzyme-mediated.
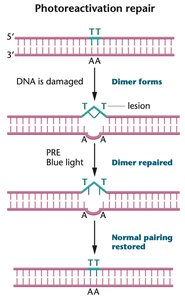
Photoreactivation: Uses light energy to repair UV-induced thymine dimers via photolyase.
Alkylation Repair: O6-methylguanine methyltransferase (MGMT) removes methyl groups from guanine.
Base Excision Repair (BER)
Mechanism and Steps
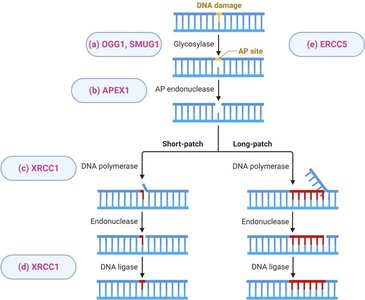
BER eliminates non-bulky damages that do not distort the double helix, such as uracil or 3-methyladenine. The process involves recognition and removal of the abnormal base, followed by repair synthesis.
DNA glycosylase: Recognizes and removes the damaged base.
AP endonuclease: Cuts the DNA backbone at the abasic site.
DNA polymerase β: Excises the incorrect nucleotide and adds the correct one.
DNA ligase: Seals the nick in the DNA backbone.
Nucleotide Excision Repair (NER)
Mechanism and Steps
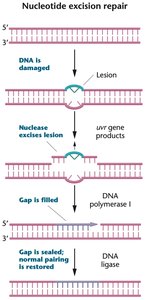
NER repairs bulky DNA lesions that distort the double helix, such as thymine dimers and bulky adducts. The damaged region is excised, and the gap is filled using the undamaged strand as a template.
Nuclease: Excises a segment of the damaged strand.
DNA polymerase I: Synthesizes new DNA to fill the gap.
DNA ligase: Seals the repaired strand.
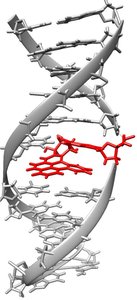
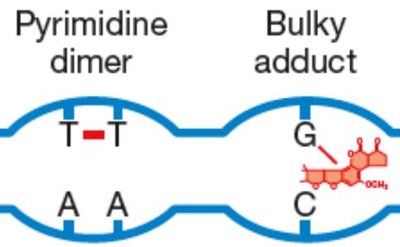
Bulky Adducts and Thymine Dimers
Bulky adducts, such as benzo[a]pyrene–guanine, and thymine dimers are classic examples of lesions repaired by NER. Failure in NER can lead to persistent DNA damage, mutations, and disorders like xeroderma pigmentosum.
Bulky adducts: Large chemical groups attached to DNA bases.
Thymine dimers: UV-induced covalent bonds between adjacent thymines.
Xeroderma pigmentosum: Caused by defective NER, leading to UV sensitivity and skin cancer risk.
Mismatch Repair (MMR)
Mechanism and Steps
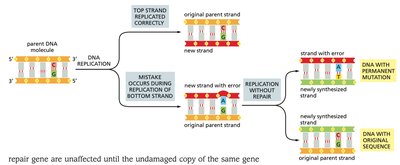
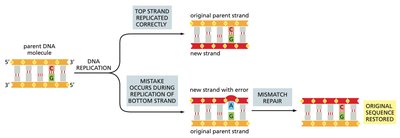
MMR corrects base-pair mismatches that escape proofreading during DNA replication. The system distinguishes the newly synthesized strand from the parental strand and removes the incorrect base.
Recognition: MMR proteins (MutS, MutL, MutH in bacteria; MSH2, MSH6, MSH3 in eukaryotes) detect mismatches.
Strand discrimination: Methylation in bacteria; nicks and replication machinery in eukaryotes.
Excision: Exonuclease removes the mismatched region.
Repair synthesis: DNA polymerase fills the gap.
Ligation: DNA ligase seals the strand.
Double-Strand Break Repair
Non-Homologous End Joining (NHEJ)
NHEJ repairs double-strand breaks by directly rejoining the broken DNA ends. This process can result in nucleotide loss or addition and does not require a homologous template.
Recognition: End-binding proteins keep broken ends together.
Processing: Nucleases may digest DNA, creating overhangs.
Gap filling: DNA polymerase fills gaps.
Ligation: DNA ligase seals the ends.
Cell cycle independence: Can occur at any phase.
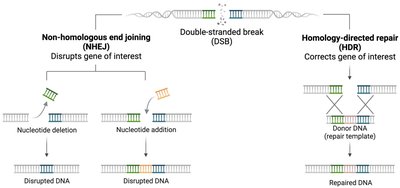
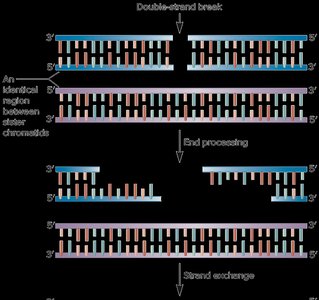
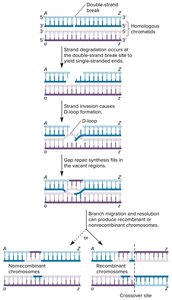
Homologous Recombination Repair (HRR)
HRR uses a sister chromatid as a template to accurately repair double-strand breaks. This mechanism is restricted to S and G2 phases when sister chromatids are available.
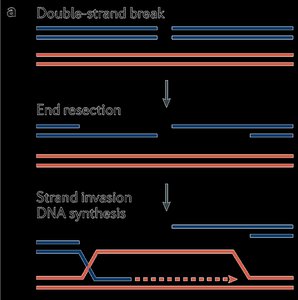
End resection: Short segments are digested to create single-stranded overhangs.
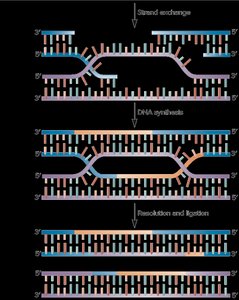
Strand invasion: The broken strand invades the sister chromatid.
DNA synthesis: Missing DNA is copied from the template.
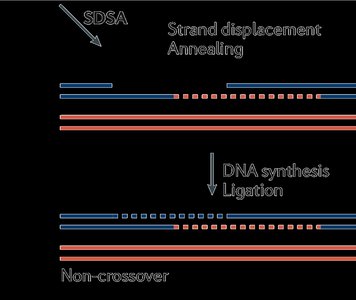
Resolution: Holliday junctions are formed and resolved, resulting in crossover or non-crossover products.
Summary Table: DNA Repair Mechanisms
Type of Damage | Repair Mechanism | Key Enzymes | Outcome |
|---|---|---|---|
Non-bulky (alkylation, oxidation, deamination, depurination) | Direct repair, BER | Photolyase, MGMT, DNA glycosylase, AP endonuclease | Restores original base or replaces abnormal base |
Bulky (thymine dimers, bulky adducts) | NER | Nuclease, DNA polymerase, DNA ligase | Removes damaged segment, fills gap |
Base mismatch | MMR | MutS, MutL, MutH (bacteria); MSH2, MSH6, MSH3 (eukaryotes) | Removes mismatched base, restores correct sequence |
Double-strand breaks | NHEJ, HRR | End-binding proteins, DNA polymerase, DNA ligase | Rejoins ends (NHEJ) or uses template for accurate repair (HRR) |
Key Terms and Concepts
DNA polymerase: Enzyme responsible for DNA synthesis and proofreading.
Photolyase: Enzyme that repairs UV-induced thymine dimers using light energy.
MGMT: Removes methyl groups from guanine bases.
DNA glycosylase: Recognizes and removes abnormal bases in BER.
AP endonuclease: Cuts DNA backbone at abasic sites.
Nuclease: Excises damaged DNA segments in NER.
MutS/MutL/MutH: Proteins involved in mismatch repair.
Holliday junction: Cross-shaped structure formed during HRR.
Equations and Formulas
DNA repair does not involve complex mathematical equations, but the following formula represents the energy required for breaking a phosphodiester bond:
Where is the change in free energy, is the gas constant, is temperature, and is the equilibrium constant.
Examples and Applications
Example: Xeroderma pigmentosum is caused by defective NER, leading to UV sensitivity.
Application: Understanding DNA repair mechanisms is crucial for cancer research and gene therapy.
Additional info: Academic context was added to clarify the molecular steps and significance of each repair pathway, as well as to provide definitions and examples for exam preparation.
