 Back
BackDNA Repair, Transposable Elements, and Eukaryotic Gene Regulation
Study Guide - Smart Notes

Double-Strand Break Repair
Overview of Double-Strand Break (DSB) Repair
Double-strand breaks (DSBs) in DNA are among the most severe types of genetic damage. If unrepaired, they can lead to chromosomal rearrangements, cancer, or cell death. Cells have evolved two primary pathways to repair DSBs: homologous recombination repair and nonhomologous end joining.
Homologous recombination repair (HRR): Uses a homologous sequence as a template for accurate repair.
Nonhomologous end joining (NHEJ): Directly ligates the broken DNA ends, often resulting in small insertions or deletions.
Homologous Recombination Repair
HRR is a high-fidelity repair mechanism that operates primarily during the late S and early G2 phases of the cell cycle, when a sister chromatid is available as a template.
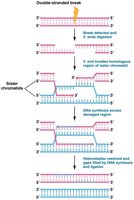
The 5' ends at the break are digested, leaving 3' single-stranded overhangs.
The 3' overhang invades the homologous sequence on the sister chromatid, forming a displacement loop.
DNA synthesis extends the invading strand, using the intact chromatid as a template.
The newly synthesized DNA is ligated, restoring the integrity of the chromosome.
Nonhomologous End Joining (NHEJ)
NHEJ is an error-prone repair pathway that is active throughout the cell cycle but is especially important in G1, before DNA replication. It does not require a homologous template.
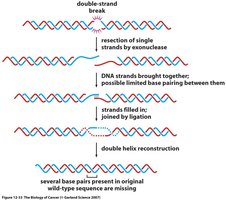
Proteins recognize and bind to the broken DNA ends.
Limited resection of single strands may occur.
The ends are processed and directly ligated together, which can result in the loss or addition of a few nucleotides.
Example: NHEJ is crucial for repairing DSBs caused by ionizing radiation or during V(D)J recombination in immune cells.
Transposable Elements
Introduction to Transposable Elements (TEs)
Transposable elements, or "jumping genes," are DNA sequences that can move within and between chromosomes. They are found in all organisms and can impact genome structure and function.
Insertion sequences (IS elements): Simple transposons found in bacteria, capable of causing mutations by inserting into genes or regulatory regions.
Bacterial transposons: Larger elements that can carry antibiotic resistance genes and move between plasmids and chromosomes.
Structure of aTransposon
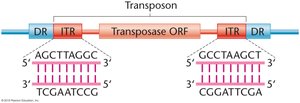
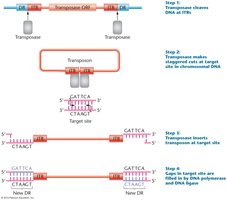
Most transposons are flanked by inverted terminal repeats (ITRs) and direct repeats (DRs) generated during insertion. The central region often encodes a transposase enzyme required for movement.
Mechanism of Transposition
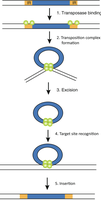
Transposition involves several steps, including binding of transposase, excision of the element, recognition of a target site, and insertion into a new location.
Ac-Ds System in Maize
The Ac-Ds system in maize is a classic example of transposable elements in plants. The Dissociation (Ds) element is nonautonomous and requires the Activator (Ac) element, which encodes transposase, for movement.
Ac (Activator): Autonomous; can move itself and Ds elements.
Ds (Dissociation): Nonautonomous; requires Ac for transposition.
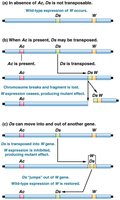
Example: In maize, insertion of Ds into the C gene disrupts anthocyanin production, resulting in colorless kernels. Excision of Ds restores color in some cells, producing spotted kernels.
Retrotransposons
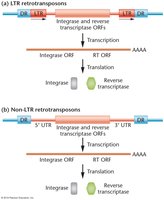
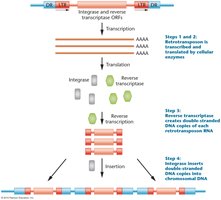
Retrotransposons are transposable elements that move via an RNA intermediate. They are divided into two main types: LTR (long terminal repeat) and non-LTR retrotransposons. Both types can be autonomous or nonautonomous.
Transcription by RNA polymerase produces an RNA copy.
Reverse transcriptase synthesizes DNA from the RNA template.
Integrase inserts the new DNA copy into the genome.
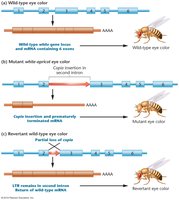
Copia Elements in Drosophila
Copia elements are a family of retrotransposons in Drosophila that are widely dispersed and can transpose to different chromosomal locations. They are characterized by direct terminal repeats (DTRs) and inverted terminal repeats (ITRs).

Example: Insertion of copia elements can disrupt gene function, such as altering eye color in Drosophila.
Transposable Elements in Humans
Nearly half of the human genome consists of transposable elements, primarily LINEs (long interspersed elements) and SINEs (short interspersed elements). While most are inactive, some can still transpose and cause mutations.
LINEs and SINEs can contribute to genetic diversity and evolution.
Approximately 0.2% of detectable human mutations are due to TE insertions.
Regulation of Gene Expression in Eukaryotes
Levels of Regulation and Chromatin Remodeling
Gene expression in eukaryotes is regulated at multiple levels, including chromatin structure, transcription, RNA processing, translation, and post-translational modification.
Chromatin remodeling and histone modifications play a central role in controlling access to DNA for transcription.
Acetylation of histones generally activates transcription, while methylation can repress it.
Chromatin Structure and Territories
Eukaryotic DNA is packaged into chromatin, which can be organized into distinct chromosome territories within the nucleus. Chromatin can be either heterochromatin (condensed, inactive) or euchromatin (less condensed, active).
Transcriptionally active genes are often located at the edges of chromosome territories.
Transcription Factories
Transcription factories are nuclear sites containing high concentrations of RNA polymerase and transcription factors, facilitating efficient gene expression.
Histone Modification and Chromatin Remodeling
Histone proteins can be covalently modified by acetylation, methylation, or phosphorylation, affecting chromatin structure and gene expression.
Histone acetyltransferases (HATs) add acetyl groups, reducing histone-DNA affinity and promoting transcription.
Chromatin remodeling complexes, such as SWI/SNF, reposition or remove nucleosomes to expose regulatory DNA sequences.
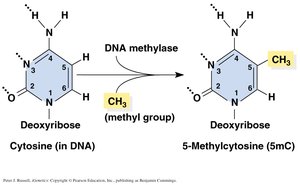
DNA Methylation and Genetic Imprinting
Direct methylation of cytosine bases in DNA is a mechanism for gene silencing. Methylated DNA recruits proteins that remove acetyl groups from histones and block transcription factor binding.
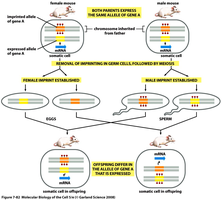
Genetic imprinting involves methylation patterns that result in expression of only one parental allele.
Regulation of Transcription
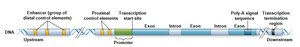
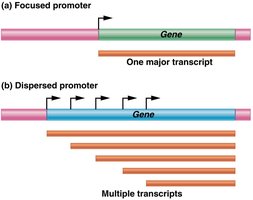
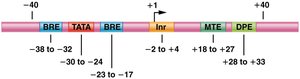
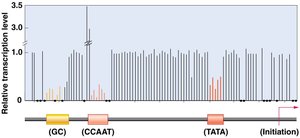
Eukaryotic genes contain promoters, enhancers, and silencers that regulate transcription. Promoters are required for transcription initiation, while enhancers and silencers modulate the level and tissue specificity of expression.
Promoters: Core and proximal elements; recognized by transcription machinery.
Enhancers: Increase transcription from a distance; can be upstream, downstream, or within genes.
Silencers: Repress transcription initiation.
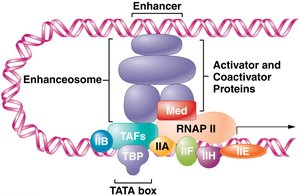
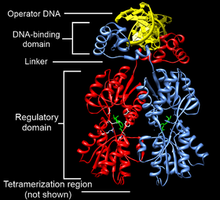
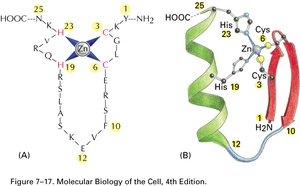
Transcription Factors and the Pre-Initiation Complex
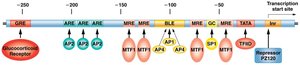
Transcription factors are proteins that bind to specific DNA sequences (cis-acting elements) to regulate gene expression. They contain DNA-binding and trans-activating domains. The assembly of general transcription factors and RNA polymerase II at the promoter forms the pre-initiation complex (PIC).
Coactivators and the enhanceosome facilitate interactions between activators, repressors, and the transcription machinery.
GAL Gene System in Yeast
The GAL gene system in yeast is a model for inducible gene expression. GAL genes are transcribed only in the presence of galactose, regulated by the interplay of structural and regulatory genes, and the upstream activating sequence (UASG).
GAL4 protein binds UASG to activate transcription; GAL80 inhibits GAL4 in the absence of galactose.
Post-Transcriptional Regulation
Alternative Splicing
Alternative splicing allows a single gene to produce multiple mRNA and protein isoforms, greatly expanding the proteome. Spliceopathies are diseases caused by splicing defects.
Example: The Dscam gene in Drosophila can generate over 38,000 isoforms through alternative splicing.
Control of mRNA Stability
The steady-state level of mRNA is determined by the balance between transcription and degradation. mRNA stability is regulated by poly-A tail length, decapping, and endonucleolytic cleavage.
anslational Tand Post-Translational Regulation
Protein levels are further regulated by translation efficiency and protein degradation. Ubiquitin tags proteins for degradation by the proteasome. The p53 protein is a key example of a transcription factor regulated by both synthesis and degradation in response to cellular stress.
Summary Table: Main DNA Repair Pathways
Pathway | Template Required? | Accuracy | Cell Cycle Phase |
|---|---|---|---|
Homologous Recombination Repair | Yes (sister chromatid) | High | Late S/G2 |
Nonhomologous End Joining | No | Low (error-prone) | All, especially G1 |
