 Back
BackDNA Replication and Recombination: Key Concepts and Mechanisms
Study Guide - Smart Notes

DNA Replication and Recombination
Overview
This chapter explores the molecular mechanisms of DNA replication and recombination, focusing on the enzymes, processes, and regulatory features that ensure accurate duplication and maintenance of genetic information in both prokaryotic and eukaryotic cells.
DNA Synthesis in Bacteria
DNA Polymerases in Bacterial DNA Replication
DNA synthesis in bacteria is carried out by five DNA polymerases, each with distinct roles in replication and repair.
DNA Polymerase I: Discovered by Arthur Kornberg, it directs DNA synthesis, requires a DNA template, and all four deoxyribonucleoside triphosphates (dNTPs).
DNA Polymerase II and III: Also elongate DNA and possess proofreading activity.
DNA Polymerase III: The primary enzyme for DNA replication in E. coli.
DNA Polymerase IV and V: Primarily involved in DNA repair, especially after damage such as UV exposure.
All DNA polymerases require a free 3'-OH group to add nucleotides and synthesize DNA in the 5' to 3' direction.
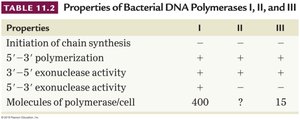
Properties | I | II | III |
|---|---|---|---|
Initiation of chain synthesis | + | - | - |
5'→3' polymerization | + | + | + |
3'→5' exonuclease activity | + | + | + |
5'→3' exonuclease activity | + | - | - |
Molecules of polymerase/cell | 400 | ? | 15 |

Chain Elongation and Proofreading
DNA chain elongation occurs by adding nucleotides to the 3' end, forming new phosphodiester bonds. DNA polymerases possess proofreading activity (3'→5' exonuclease) to remove incorrectly paired nucleotides, ensuring high fidelity during replication.
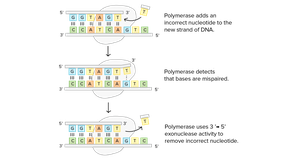
Functions of DNA Polymerases
Elongation: DNA Pol I, II, and III can elongate existing DNA strands.
Proofreading: All three have 3'→5' exonuclease activity for error correction.
Primer Removal: Only DNA Pol I has 5'→3' exonuclease activity, allowing it to remove RNA primers and fill gaps.
Structure and Function of DNA Pol III Holoenzyme
The DNA Pol III holoenzyme is a multi-subunit complex with distinct functions:
α subunit: 5'→3' polymerization
ε subunit: 3'→5' exonuclease (proofreading)
Key Steps in Bacterial DNA Replication
Seven Key Issues to Resolve
Unwinding of the helix
Reducing increased coiling (supercoiling)
Synthesis of primer for initiation
Discontinuous synthesis of the second strand
Removal of RNA primers
Joining of gap-filling DNA to adjacent strand
Proofreading
Step 1: Unwinding the Helix
DnaA protein: Binds to the origin of replication (ORI), causing the DNA to open and expose single-stranded DNA (ssDNA).
Single-stranded binding proteins (SSBPs): Stabilize the open conformation of the helix.
DNA helicase: Unwinds the DNA helix using energy from ATP hydrolysis.
Step 2: Reducing Supercoiling
As the DNA helix unwinds, supercoiling occurs ahead of the replication fork. DNA gyrase (a type of topoisomerase) relieves this tension by making transient cuts in the DNA.

Step 3: Synthesis of Primer for Initiation
Primase: An RNA polymerase that synthesizes a short RNA primer, providing a free 3'-OH group for DNA polymerase III to begin elongation.
DNA Pol I: Removes RNA primers and replaces them with DNA.
Step 4: Discontinuous Synthesis of the Second Strand
Because DNA polymerase can only synthesize in the 5'→3' direction, the two template strands are replicated differently:
Leading strand: Synthesized continuously.
Lagging strand: Synthesized discontinuously as Okazaki fragments, each initiated by an RNA primer.
Steps 5 & 6: Removal of RNA Primers and Joining of DNA Fragments
Okazaki fragments: Short DNA segments on the lagging strand.
DNA Pol I: Removes RNA primers and fills in the gaps with DNA.
DNA ligase: Seals nicks between fragments by forming phosphodiester bonds.
Step 7: Proofreading
DNA polymerases possess exonuclease activity to remove mismatched nucleotides, ensuring high fidelity of DNA replication.
Eukaryotic DNA Replication
Complexity of Eukaryotic DNA Replication
Eukaryotic DNA replication is more complex than bacterial replication due to:
Greater amount of DNA
Multiple origins of replication (ORIs) per chromosome
Linear chromosomes
Presence of telomeres
DNA packaged with histone proteins (chromatin)
Multiple Origins of Replication
Eukaryotic chromosomes contain multiple ORIs to facilitate rapid and efficient DNA synthesis.
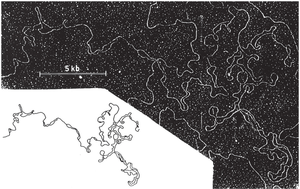
DNA and Histone Proteins
Eukaryotic DNA is tightly associated with histone proteins, forming nucleosomes. These structures must be temporarily removed or repositioned during replication to allow access for DNA polymerases.

Telomeres and Telomerase
Structure and Function of Telomeres
Telomeres are repetitive DNA sequences at the ends of linear chromosomes that protect genetic information from loss during replication. With each cell division, telomeres shorten, eventually leading to cellular aging.
Role of Telomerase
Telomerase is a ribonucleoprotein enzyme that extends telomeres by adding repetitive sequences, using an RNA template within the enzyme. This activity is crucial in stem cells and cancer cells, but is generally inactive in most somatic cells.
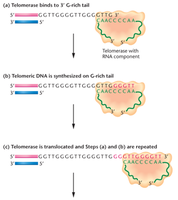
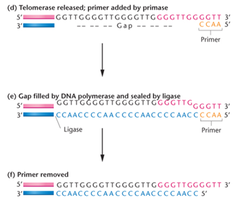
Telomerase, Aging, and Cancer
Telomerase activity is linked to cellular immortality in cancer cells.
Shortened telomeres are associated with aging and age-related diseases.
Therapeutic strategies are being developed to target telomerase in cancer treatment.
Genetic Recombination
Homologous Recombination
Genetic recombination involves the exchange of genetic material between homologous DNA molecules. This process is essential for genetic diversity, DNA repair, and proper chromosome segregation during meiosis.
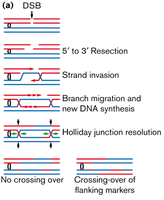
Double-strand break (DSB) repair: Involves resection, strand invasion, branch migration, and resolution of Holliday junctions.
Can result in crossing-over or non-crossing-over of flanking markers.
Summary Table: Properties of Bacterial DNA Polymerases
Property | DNA Pol I | DNA Pol II | DNA Pol III |
|---|---|---|---|
Initiation of chain synthesis | + | - | - |
5'→3' polymerization | + | + | + |
3'→5' exonuclease activity | + | + | + |
5'→3' exonuclease activity | + | - | - |
Molecules/cell | 400 | ? | 15 |
Key Equations
DNA Synthesis Directionality:
Proofreading Exonuclease Activity:
Additional info: The analogy of a telephone cord is used to help visualize DNA supercoiling, which is relieved by topoisomerases such as DNA gyrase. The electron micrographs included illustrate the physical structure of replicating DNA and nucleosome organization. The diagrams of telomerase action clarify the stepwise extension and completion of telomeres during replication.
