 Back
BackDNA Replication in Eukaryotes: Chromatin, Telomeres, and Telomerase
Study Guide - Smart Notes

DNA Replication in Eukaryotes
Key Differences Between Prokaryotic and Eukaryotic DNA Replication
DNA replication in eukaryotes is more complex than in prokaryotes due to differences in chromosome structure and organization.
Multiple Origins of Replication: Eukaryotic chromosomes are linear and contain multiple origins of replication, allowing for simultaneous DNA synthesis at many sites. In contrast, E. coli has a single origin on its circular chromosome.
Chromatin Structure: Eukaryotic DNA is packaged into chromatin, with DNA wrapped around histone proteins to form nucleosomes. Nucleosomes must be disassembled ahead of the replication fork and reassembled behind it, a process involving chromatin assembly factors (CAF).
Linear Chromosomes and End-Replication Problem: The linear nature of eukaryotic chromosomes creates a unique challenge at chromosome ends (telomeres), leading to progressive shortening with each replication cycle.
Chromatin and Nucleosome Structure
Organization of Chromatin
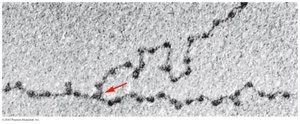
Chromatin is the complex of DNA and proteins (mainly histones) that forms chromosomes within the nucleus of eukaryotic cells. During interphase, chromatin appears as "beads on a string" under electron microscopy, where the beads are nucleosomes.
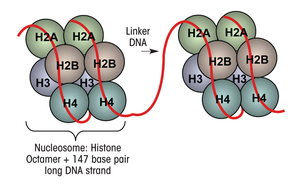
Nucleosome: The fundamental unit of chromatin, consisting of an octamer of histones (two each of H2A, H2B, H3, and H4) wrapped by 147 base pairs of DNA.
Linker DNA: The stretch of DNA (typically 20–100 bp, average ~53 bp) between nucleosomes, associated with histone H1 (the linker histone).
Histone-DNA Interaction: Histones are rich in positively charged amino acids, which interact with the negatively charged phosphate backbone of DNA, facilitating tight wrapping.
Size: Each nucleosome is about 11 nm in diameter.

Telomeres: Structure and Function
Telomere Sequence and Protection
Telomeres are repetitive DNA sequences at the ends of linear chromosomes, essential for protecting chromosome ends from degradation and fusion.
Sequence: In vertebrates, the telomeric repeat is 5' TTAGGG 3', repeated thousands of times.
G-rich and C-rich Strands: Each telomere has a G-rich strand (with a 3' overhang) and a complementary C-rich strand.
T-loop/D-loop Model: The G-rich overhang invades the double-stranded region of the telomere, forming a T-loop (large loop) and a D-loop (displacement loop), which "hide" the chromosome ends from DNA repair machinery.
Telomere-Binding Proteins: Proteins such as TRF1 and TRF2 stabilize the telomere structure (not covered in detail here).
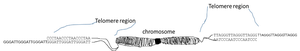
The End-Replication Problem and Telomere Shortening
Why Replication Causes Telomere Shortening
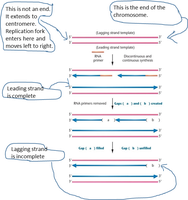
DNA polymerases require a primer to initiate synthesis and cannot fully replicate the 3' ends of linear chromosomes. This results in progressive telomere shortening with each cell division, particularly on the lagging strand.
Lagging Strand Synthesis: After removal of the final RNA primer, a gap remains at the chromosome end, leading to loss of telomeric DNA.
Consequences: Over time, telomere shortening can lead to loss of protective structure, chromosome instability, and cell senescence or apoptosis.
Telomerase: Structure and Mechanism
Components and Function of Telomerase
Telomerase is a ribonucleoprotein enzyme that extends telomeres, counteracting the shortening that occurs during DNA replication.
TERT (Telomerase Reverse Transcriptase): The protein component with reverse transcriptase activity, synthesizing DNA from an RNA template.
TERC (Telomerase RNA Component): The RNA component that serves as a template for adding telomeric repeats (e.g., 5' CUAACCCUAAC 3').
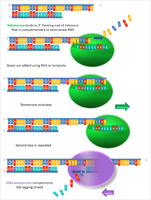
Mechanism: Telomerase binds to the 3' overhang of the G-rich strand, uses TERC as a template to extend the telomere, and then translocates to repeat the process.
Telomeres, Aging, and Disease
Cellular Senescence and the Hayflick Limit
Most human somatic cells lack active telomerase and can only divide a limited number of times (~50 divisions), a phenomenon known as the Hayflick limit. This is primarily due to telomere shortening.
Evidence: Experiments show that expressing telomerase in cultured cells allows them to divide indefinitely, while normal cells senesce after telomeres become critically short.
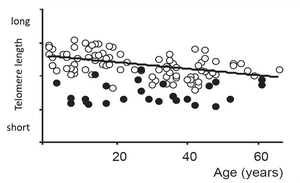
Role in Aging: Telomere shortening is associated with cellular aging and tissue dysfunction.
Dyskeratosis Congenita (DC): A Telomere-Related Disorder
Dyskeratosis congenita is a genetic disorder caused by mutations in the TERC gene, leading to defective telomerase and abnormally short telomeres.
Symptoms: Classic features include nail dystrophy, oral leukoplakia, abnormal skin pigmentation, and bone marrow failure. Other symptoms may include premature aging signs and organ dysfunction.
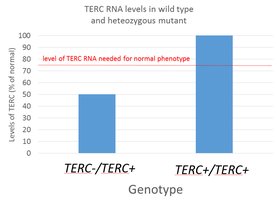
Genetics: The wild-type TERC+ allele is incompletely dominant to the null TERC- allele due to haploinsufficiency (one normal allele does not produce enough TERC RNA for normal function).
Penetrance: Not all individuals with the TERC+/TERC- genotype develop DC, indicating incomplete penetrance.
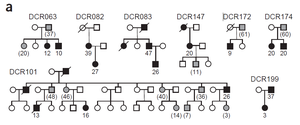
Genetic Anticipation: Symptoms appear at younger ages in successive generations, likely due to inheritance of both mutant alleles and already-shortened telomeres.
Telomeres, Telomerase, and Cancer
Balancing Aging and Cancer Risk
While telomerase activation could theoretically slow or reverse aging by maintaining telomere length, it also poses a significant cancer risk. Most cancer cells reactivate telomerase, allowing unlimited division and tumor progression.
Natural Defense: Telomere shortening acts as a "last line of defense" against uncontrolled cell proliferation, forcing cells into senescence before they can become cancerous.
Cancer Cells: Overcome this barrier by reactivating telomerase, enabling them to maintain telomere length and divide indefinitely.
Summary Table: Key Features of Eukaryotic DNA Replication, Telomeres, and Telomerase
Feature | Description |
|---|---|
Origin of Replication | Multiple per chromosome in eukaryotes; single in prokaryotes |
Chromatin Structure | DNA wrapped around histone octamers to form nucleosomes |
Telomere Sequence | 5' TTAGGG 3' repeats in vertebrates |
Telomerase Components | TERT (enzyme), TERC (RNA template) |
Telomere Shortening | Occurs with each cell division due to incomplete lagging strand replication |
Associated Diseases | Dyskeratosis congenita, cancer, aging-related tissue dysfunction |
