 Back
BackDNA Replication in Eukaryotes: Mechanisms, Challenges, and Telomere Maintenance
Study Guide - Smart Notes

DNA Replication in Eukaryotes
Overview and Comparison with Prokaryotic Replication
DNA replication is a fundamental process required for cell division and inheritance. While the core mechanisms are conserved between prokaryotes and eukaryotes, several key differences exist due to the complexity of eukaryotic genomes.
Similarities:
Replication begins at specific origins of replication where the DNA helix unwinds, forming a replication fork.
Both leading and lagging strand synthesis occur, with DNA polymerase mediating elongation in a bi-directional manner.
Differences:
Eukaryotes have much more DNA, organized into multiple linear chromosomes with centromeres and telomeres.
DNA is packaged with proteins (chromatin), requiring additional steps for replication.
Multiple origins of replication are present (hundreds in yeast, thousands in mammals), allowing for efficient duplication of large genomes.
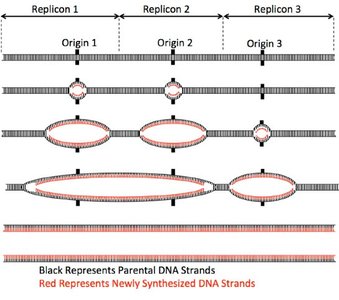
Multiple Replication Origins and Replication Bubbles
Eukaryotic chromosomes initiate replication at numerous origins, creating replication bubbles that expand bi-directionally. Each bubble contains two replication forks, ensuring rapid and complete genome duplication.
Each replicon is the DNA segment replicated from a single origin.
Replication bubbles merge as synthesis progresses, resulting in two identical DNA molecules.
Initiation of Replication: ORC and Pre-RC Complexes
The initiation of DNA replication in eukaryotes is tightly regulated to ensure each segment is replicated only once per cell cycle.
Origin Recognition Complex (ORC): A set of proteins that bind to replication origins early in the G1 phase, marking them for replication.
Pre-Replication Complex (pre-RC): Additional proteins assemble at the origins, licensing them for replication. The presence of pre-RC is required for DNA polymerase recruitment.
Once replication begins, pre-RC proteins disperse and reassemble at origins in the next G1 phase, preventing re-replication within the same cycle.
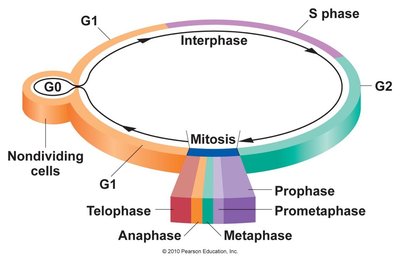
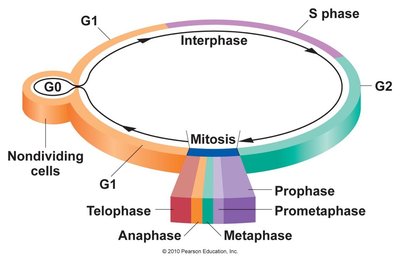
Chromatin and DNA Binding Proteins
Eukaryotic DNA is packaged with proteins, primarily histones, forming chromatin. This packaging must be temporarily disrupted for replication to proceed.
Histones and other DNA-binding proteins are removed or modified ahead of the replication fork.
After new DNA synthesis, these proteins re-associate with the daughter DNA strands, restoring chromatin structure.

Challenges of Replicating Linear Chromosomes: Telomeres
Structure and Function of Telomeres
Linear chromosomes present unique challenges at their ends, known as telomeres. These specialized structures prevent chromosome ends from being recognized as DNA breaks and protect against degradation and fusion.
Telomeres consist of short, tandemly repeated DNA sequences: 5'- (TTAGGG)n -3' on the G-rich strand and 3'- (AATCCC)n -5' on the C-rich strand.
The G-rich strand forms a 3' single-stranded overhang, which can fold back and form G-quartet structures, creating a protective loop.
Replication Problems at Telomeres
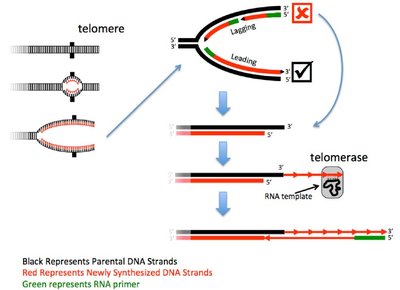
Conventional DNA polymerases cannot fully replicate the ends of linear chromosomes, leading to progressive shortening with each cell division.
After removal of the RNA primer on the lagging strand, a 3' gap remains that cannot be filled by DNA polymerase.
This results in gradual telomere shortening, which can eventually compromise chromosome integrity.
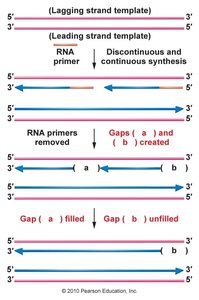
Telomerase: The Solution to the End-Replication Problem
Telomerase is a specialized ribonucleoprotein enzyme that extends the G-rich strand of telomeres, counteracting the end-replication problem.
Telomerase contains an RNA template complementary to the telomeric repeat, allowing it to add new repeats to the 3' end of the chromosome.
This process is a form of reverse transcription, as the RNA template is used to synthesize DNA.
After telomerase action, conventional DNA polymerase can fill in the complementary C-rich strand, restoring telomere length.
Summary Table: Key Differences in DNA Replication
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Genome Structure | Circular, single chromosome | Linear, multiple chromosomes |
Origins of Replication | Single origin | Multiple origins per chromosome |
DNA Packaging | Minimal (no histones) | Chromatin (histones and other proteins) |
End-Replication Problem | Absent (circular DNA) | Present (solved by telomerase) |
Key Terms
Origin of Replication: Specific DNA sequence where replication begins.
Replication Fork: The Y-shaped region where DNA is unwound and new strands are synthesized.
Telomere: Repetitive DNA sequence at the end of a linear chromosome, protecting it from degradation.
Telomerase: Enzyme that extends telomeres by adding DNA repeats using an RNA template.
Pre-Replication Complex (pre-RC): Protein assembly that licenses origins for replication.
Relevant Equations
General telomeric repeat sequence (human):
Complementary C-rich strand:
