 Back
BackDNA Replication: Mechanisms, Enzymes, and Applications
Study Guide - Smart Notes

DNA Replication: Fundamental Concepts
Three Models of DNA Replication
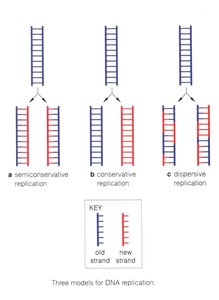
DNA replication is a central process in genetics, ensuring the faithful transmission of genetic information from one generation to the next. Three theoretical models were proposed to explain how DNA is duplicated:
Semiconservative replication: Each daughter DNA molecule consists of one parental (old) strand and one newly synthesized strand.
Conservative replication: The parental DNA remains intact, and an entirely new molecule is synthesized.
Dispersive replication: Both daughter molecules contain interspersed segments of old and new DNA.
Semiconservative replication is the model supported by experimental evidence and is the mechanism used by all organisms.
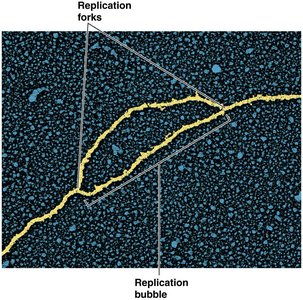
Replication Bubble and Forks
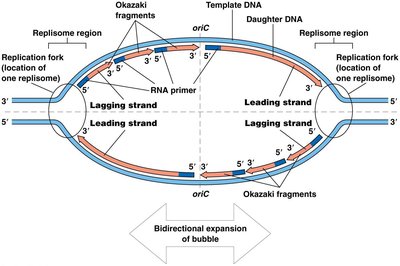
DNA replication begins at specific sites called origins of replication. The DNA unwinds locally, forming a replication bubble with two replication forks where new DNA synthesis occurs.
Replication proceeds bidirectionally from the origin.
Each fork moves outward, synthesizing new DNA strands.
Bacterial DNA Replication
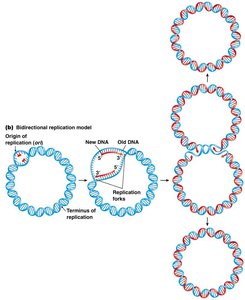
Bidirectional Replication in Circular Chromosomes
Bacterial chromosomes are typically circular and possess a single origin of replication. Replication proceeds in both directions, creating two replication forks that eventually meet at the terminus.
Replication is bidirectional and semiconservative.
Each daughter cell receives one old and one new DNA strand.
Key Proteins in DNA Replication
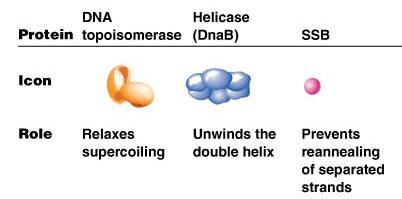
DNA replication requires the coordinated action of several proteins, each with a specific function:
Protein | Icon | Role |
|---|---|---|
DNA topoisomerase | Orange structure | Relaxes supercoiling |
Helicase (DnaB) | Blue structure | Unwinds the double helix |
SSB | Pink sphere | Prevents reannealing of separated strands |
Protein | Icon | Role |
|---|---|---|
Primase | Red structure | Synthesizes RNA primers |
DNA pol III | Green structure | Synthesizes DNA |
DNA pol I | Yellow structure | Removes and replaces RNA primer with DNA |
DNA ligase | Blue structure | Joins DNA segments |
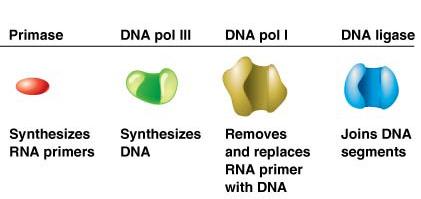
Structure of the Origin of Replication (oriC) in E. coli
The origin of replication in E. coli (oriC) is a well-defined region containing specific sequence motifs:
AT-rich regions: Easier to unwind due to fewer hydrogen bonds.
DnaA boxes: Binding sites for initiator protein DnaA.
GATC methylation sites: Involved in regulation.
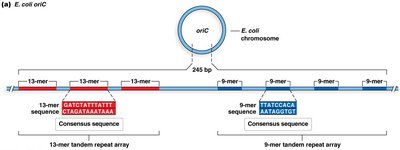
Initiation of Replication at oriC
Replication initiation involves the binding of DnaA proteins to DnaA boxes, bending and unwinding the AT-rich region, and recruitment of helicase (DnaB) by DnaC. Helicase unwinds the DNA, and single-stranded binding proteins (SSBs) stabilize the unwound strands.
Formation of the replisome at the replication fork.
Accessory proteins assist in complex assembly.
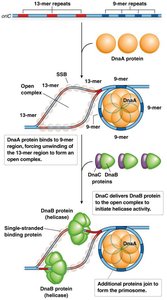
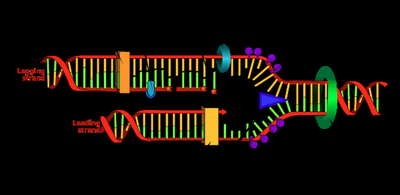
The Replisome: Multi-Enzyme Complex
The replisome is a large protein complex responsible for DNA synthesis at the replication fork. It includes:
DNA helicase: Unwinds DNA.
DNA polymerases: Synthesize new DNA strands.
DNA primase: Synthesizes RNA primers.
Clamp loaders and other accessory proteins.
DNA polymerase synthesizes the leading strand continuously and the lagging strand discontinuously in short fragments called Okazaki fragments.
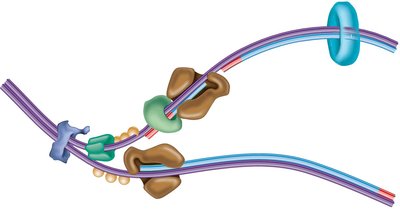
Replication Fork Maintenance
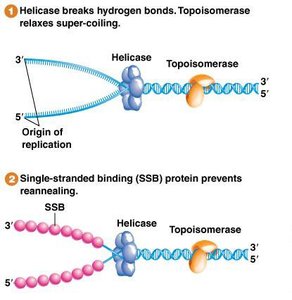
Helicase and SSBs are essential for maintaining the replication fork:
Helicase: Unwinds the DNA double helix.
SSBs: Bind to single-stranded DNA to prevent reannealing.
Topoisomerase Function
Unwinding DNA creates torsional stress and supercoiling. Topoisomerases relieve this stress by making controlled cuts in the DNA:
Type I: Single-stranded breaks.
Type II: Double-stranded breaks.
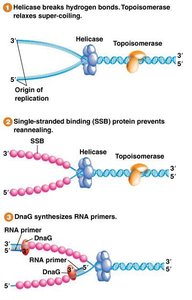
RNA Primers and Primase
DNA polymerase cannot initiate synthesis de novo; it requires a 3' end provided by an RNA primer. Primase synthesizes these primers.
Primers are short RNA sequences complementary to the template.
Primase is a specialized RNA polymerase.
Bacterial DNA Polymerases
Three main DNA polymerases function in bacteria:
DNA pol III: Main replicative enzyme, synthesizes new DNA.
DNA pol I: Removes RNA primers and replaces them with DNA.
DNA pol II, IV, V: Involved in DNA repair.
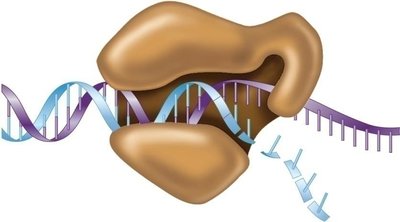
Mechanism of DNA Polymerase III
DNA polymerase III catalyzes the addition of nucleotides in the 5' to 3' direction, using dNTPs as substrates. The enzyme's structure resembles a right hand, with the template DNA passing through the palm.
Can only extend an existing strand.
Requires a primer to start synthesis.
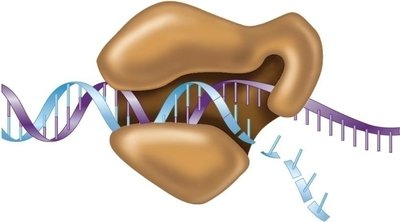
Processivity and Speed of DNA Polymerase III
The holoenzyme remains attached to the DNA, allowing rapid synthesis. The beta subunit is crucial for processivity, enabling the enzyme to synthesize up to 750 nucleotides per second.
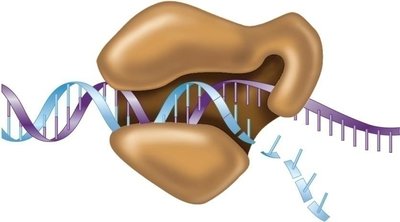
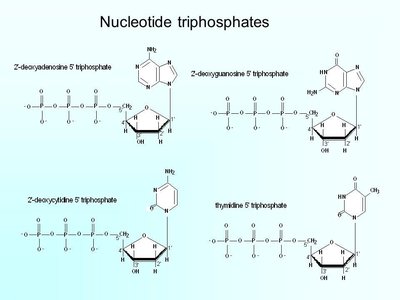
Powering DNA Synthesis: Nucleotide Triphosphates
DNA polymerase catalyzes the formation of phosphodiester bonds between nucleotides, powered by the hydrolysis of nucleotide triphosphates (dNTPs). The energy released drives the reaction:
Bond formation between 3' OH of growing strand and 5' phosphate of incoming dNTP.
Release of pyrophosphate.
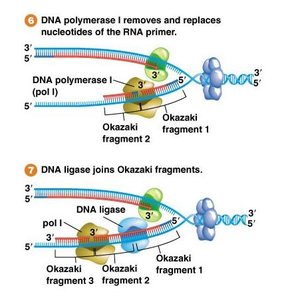
Primer Removal and DNA Ligase
After DNA synthesis, RNA primers are removed and replaced with DNA by DNA pol I. DNA ligase seals the nicks between fragments, creating a continuous strand.
Okazaki Fragments and the Lagging Strand Problem
The lagging strand is synthesized discontinuously as Okazaki fragments due to the antiparallel nature of DNA and the unidirectional activity of DNA polymerase.
Fragments are joined by DNA ligase.
Ensures complete replication of both strands.
Eukaryotic DNA Replication
Multiple Origins and Linear Chromosomes
Eukaryotic chromosomes are linear and contain multiple origins of replication, each forming its own replication bubble. Replication proceeds bidirectionally from each origin.
Replication forks merge, resulting in sister chromatids.
Origins are less well characterized in higher eukaryotes.
Eukaryotic DNA Polymerases
Eukaryotes utilize a specialized set of DNA polymerases:
Pol α: Initiates DNA synthesis with RNA/DNA primer.
Pol δ: Extends lagging strand.
Pol ε: Extends leading strand.
Pol γ: Replicates mitochondrial DNA.
Pol β: Functions in base excision repair.
PrimPol: Combines primase and polymerase activities.
Primer Replacement in Eukaryotes
On the lagging strand, DNA polymerase δ displaces the RNA primer, creating a flap. Flap endonuclease (FEN1) removes the flap, and DNA ligase joins the fragments.
Telomeres and the End-Replication Problem
Linear chromosomes pose a challenge for lagging strand synthesis at the ends. Telomeres are repetitive sequences that protect chromosome ends. Telomerase extends telomeres using an internal RNA template, allowing complete replication.
Telomere sequence in vertebrates: 5'-TTAGGG-3'
Telomerase is active in germ-line and some stem cells.
Telomere shortening is associated with aging and cancer.
DNA Proofreading and Error Correction
Polymerase Fidelity and Proofreading
DNA polymerases have high fidelity due to proofreading activity (3' to 5' exonuclease). Additional repair systems further reduce error rates.
Polymerase error rate: ~1 in 100,000 bases.
With repair: ~1 in 10 billion bases.
In Vitro DNA Replication: PCR and Sanger Sequencing
Polymerase Chain Reaction (PCR)
PCR is an automated method to amplify specific DNA segments. It involves repeated cycles of denaturation, primer annealing, and extension using a heat-stable DNA polymerase (e.g., Taq polymerase).
Applications: molecular cloning, gene detection, sequencing.
Exponential amplification: DNA doubles each cycle.
Separation and Visualization of PCR Products
Amplified DNA fragments are separated by gel electrophoresis and visualized by DNA-specific staining. Smaller fragments migrate faster in the gel.
Sanger Sequencing
Sanger sequencing uses chain-terminating dideoxynucleotides (ddNTPs) to generate DNA fragments of varying lengths. Fragments are separated by electrophoresis, and the sequence is determined by the order of termination.
Modern methods use fluorescently labeled ddNTPs and capillary electrophoresis.
Summary Table: Prokaryotic vs Eukaryotic DNA Replication
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Chromosome shape | Circular | Linear |
Origin of replication | Single | Multiple |
Replication direction | Bidirectional (5' to 3') | Bidirectional (5' to 3') |
Replication termination | ter sequences, Tus proteins | Chromosome ends, telomeres |
Leading strand synthesis | DNA pol III | DNA pol ε |
Lagging strand synthesis | DNA pol III | DNA pol δ |
Primer removal | DNA pol I | FEN1 |
Topoisomerase, helicase, SSB, primase, ligase | Yes | Yes |
Additional info: These notes expand on the original lecture content, providing definitions, examples, and context for each step and enzyme involved in DNA replication. The included images directly reinforce the explanations and mechanisms described.
