 Back
BackChapter 7
Study Guide - Smart Notes

DNA Replication: Mechanisms and Models
Overview of DNA Replication
DNA replication is a fundamental process in all living organisms, ensuring the accurate transmission of genetic information from one generation to the next. The process is highly regulated and involves a series of coordinated steps and specialized enzymes.
Semiconservative replication: Each daughter DNA molecule consists of one parental (old) strand and one newly synthesized strand.
Conservative replication: The parental molecule remains intact, and an entirely new molecule is synthesized (not the actual mechanism in cells).
Dispersive replication: Parental and new DNA are interspersed in both strands (not the actual mechanism in cells).
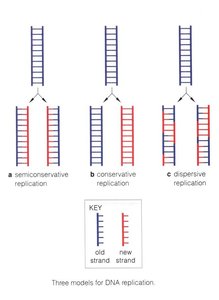
Key Point: Experimental evidence supports the semiconservative model as the mechanism of DNA replication in all organisms.
Semiconservative Replication in Detail
During semiconservative replication, the double helix unwinds, and each parental strand serves as a template for the synthesis of a complementary daughter strand. This results in two identical DNA molecules, each with one old and one new strand.
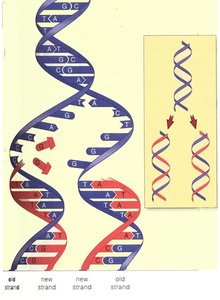
Initiation and Directionality of Replication
Replication Origins and Bubbles
Replication begins at specific sites called origins of replication. In bacteria, there is typically a single origin, while eukaryotes have multiple origins per chromosome. Replication proceeds bidirectionally, forming a replication bubble with two replication forks moving in opposite directions.
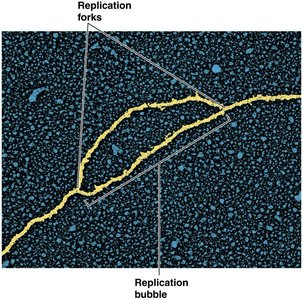
Bidirectional Replication in Circular Chromosomes
In prokaryotes such as E. coli, the circular chromosome replicates bidirectionally from a single origin (oriC) until the two replication forks meet at the terminus.
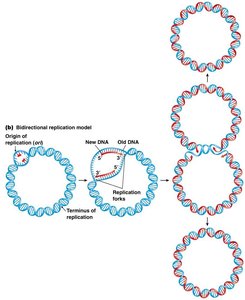
Key Enzymes and Proteins in DNA Replication
Major Proteins and Their Functions
DNA replication requires a coordinated effort of several enzymes and proteins, each with a specific role:
Protein | Role |
|---|---|
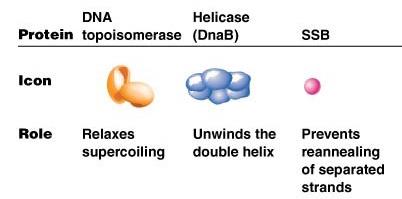
DNA topoisomerase | Relaxes supercoiling ahead of the replication fork |
Helicase (DnaB) | Unwinds the double helix |
SSB (Single-stranded binding protein) | Prevents reannealing of separated strands |
Protein | Role |
|---|---|
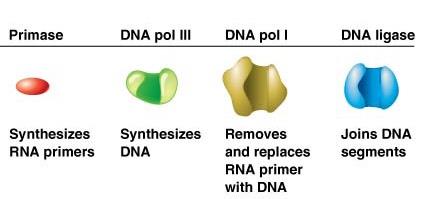
Primase | Synthesizes RNA primers |
DNA pol III | Synthesizes DNA |
DNA pol I | Removes and replaces RNA primer with DNA |
DNA ligase | Joins DNA segments |
Initiation of Replication in Bacteria
Structure of the Origin (oriC) in E. coli
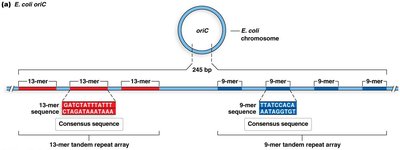
The oriC region in E. coli is about 245 base pairs and contains several conserved sequence motifs, including 13-mer and 9-mer repeats. These sequences are recognized by initiator proteins to begin replication.
Assembly of the Initiation Complex
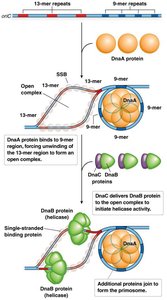
Initiation involves the binding of DnaA proteins to the 9-mer repeats, causing local unwinding at the adjacent AT-rich 13-mer repeats. DnaC then helps load DnaB helicase, which further unwinds the DNA, and SSBs stabilize the single strands.
Elongation: The Replisome and Synthesis of New Strands
The Replisome Complex
The replisome is a large protein complex that coordinates the synthesis of both leading and lagging DNA strands at the replication fork. It includes DNA polymerases, primase, helicase, SSBs, and other factors.
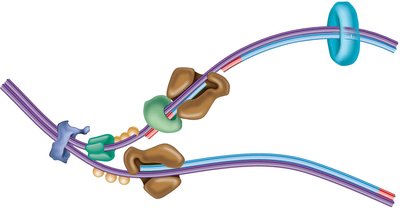
Stepwise Mechanism of DNA Synthesis
Helicase unwinds the DNA double helix.
Topoisomerase relieves supercoiling ahead of the fork.
SSBs bind to single-stranded DNA to prevent reannealing.
Primase synthesizes short RNA primers to provide a 3' OH group for DNA polymerase.
DNA polymerase III extends the primers, synthesizing new DNA in the 5' to 3' direction.
DNA polymerase I removes RNA primers and fills in with DNA.
DNA ligase seals nicks between Okazaki fragments on the lagging strand.
Replication Fork Dynamics
Leading and Lagging Strand Synthesis
DNA polymerase can only synthesize DNA in the 5' to 3' direction. This creates a continuous leading strand and a discontinuous lagging strand composed of Okazaki fragments.
Termination and Regulation
Termination in Prokaryotes
Replication ends when the forks meet at specific termination sites (ter) bound by Tus proteins, which block further progression of the replication machinery.
Comparison: Prokaryotic vs. Eukaryotic DNA Replication
Key Differences
Chromosome structure: Prokaryotes have circular chromosomes; eukaryotes have linear chromosomes.
Origins of replication: Single in prokaryotes, multiple in eukaryotes.
Enzymes: Different DNA polymerases are used for leading and lagging strand synthesis in eukaryotes (DNA pol ε and δ, respectively).
Primer removal: DNA pol I in prokaryotes; FEN1 in eukaryotes.
End-replication problem: Eukaryotes require telomeres and telomerase to complete replication of chromosome ends.
DNA Proofreading and Fidelity
Proofreading Mechanisms
DNA polymerases possess 3' to 5' exonuclease activity, allowing them to remove incorrectly paired nucleotides and maintain high fidelity during replication. Additional repair systems further reduce the error rate.
Polymerase error rate: ~1 in 100,000,000 bases after proofreading
With repair systems: ~1 in 10,000,000,000 bases
Telomeres and the End-Replication Problem
Role of Telomeres
Linear chromosomes in eukaryotes face the end-replication problem, where the lagging strand cannot be fully replicated at the ends. Telomeres, repetitive DNA sequences, protect chromosome ends, and the enzyme telomerase extends these regions in germ-line and some stem cells.
In Vitro DNA Replication: PCR and Sanger Sequencing
Polymerase Chain Reaction (PCR)
PCR is a laboratory technique that amplifies specific DNA sequences using cycles of denaturation, primer annealing, and extension. It requires a heat-stable DNA polymerase (e.g., Taq polymerase), primers, nucleotides, and a buffer.
Applications: Molecular cloning, gene detection, DNA sequencing, forensics
Steps: Denaturation (~95°C), Annealing (~45–68°C), Extension (~72°C), repeated for 10–35 cycles
Sanger DNA Sequencing
Sanger sequencing uses chain-terminating dideoxynucleotides (ddNTPs) to generate DNA fragments of varying lengths, which are then separated by electrophoresis to determine the DNA sequence.
Key concept: Incorporation of a ddNTP terminates DNA synthesis at specific bases, allowing sequence determination.
Summary Table: Major Steps and Enzymes in DNA Replication
Step | Enzyme/Protein | Function |
|---|---|---|
Initiation | DnaA, DnaB, DnaC, SSB | Origin recognition, unwinding, stabilization |
Primer synthesis | Primase (DnaG) | RNA primer synthesis |
Elongation | DNA pol III (prokaryotes), DNA pol ε/δ (eukaryotes) | DNA strand synthesis |
Primer removal | DNA pol I (prokaryotes), FEN1 (eukaryotes) | Removes RNA primers |
Joining fragments | DNA ligase | Seals nicks between Okazaki fragments |
