 Back
BackDNA Replication: Mechanisms, Enzymes, and Chromosome End Replication
Study Guide - Smart Notes

DNA Replication: Mechanisms and Evidence
Overview of DNA Replication Mechanisms
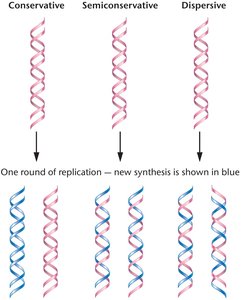
DNA replication is a fundamental process in genetics, ensuring the accurate transmission of genetic information from one generation to the next. Three main models were proposed to explain how DNA replicates: semiconservative, conservative, and dispersive.
Semiconservative Replication: Each new DNA molecule consists of one parental (old) strand and one newly synthesized strand.
Conservative Replication: The parental double helix remains intact, and an entirely new double helix is synthesized.
Dispersive Replication: Parental DNA is dispersed throughout both strands of the daughter molecules after replication.
Experimental Evidence: The Meselson-Stahl Experiment
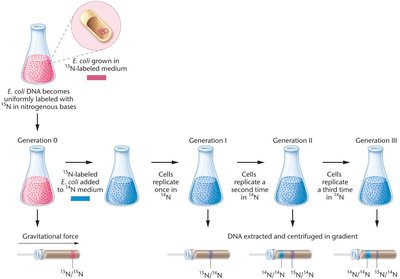
The Meselson-Stahl experiment provided definitive evidence for the semiconservative mechanism of DNA replication. By growing E. coli in media containing heavy nitrogen (15N) and then switching to light nitrogen (14N), they could distinguish old from new DNA strands using density gradient centrifugation.
Key Steps: Cells were grown in 15N, then transferred to 14N. DNA was extracted after each generation and centrifuged to separate based on density.
Results: After one replication, DNA was of intermediate density (ruling out conservative replication). After two replications, both intermediate and light DNA were observed, supporting semiconservative replication and ruling out the dispersive model.
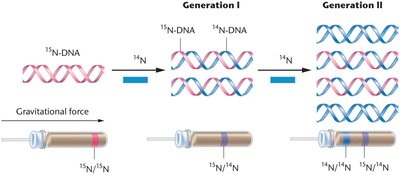
Semiconservative Replication in Eukaryotes
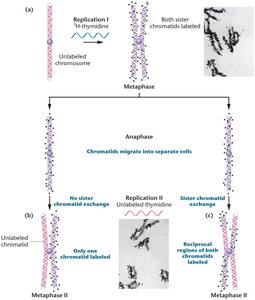
Semiconservative replication also occurs in eukaryotes. Experiments using radioactive thymidine and autoradiography demonstrated that after one replication, both chromatids are labeled, and after two replications, only one chromatid is labeled.
Initiation and Direction of DNA Replication
Origins of Replication
DNA replication begins at specific sites called origins of replication (ori). In bacteria, there is typically a single origin (oriC in E. coli), while eukaryotes have multiple origins due to their larger genomes. Replication proceeds bidirectionally from each origin, forming two replication forks.
Replicon: The length of DNA replicated from a single origin.
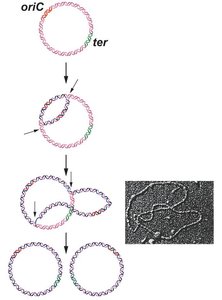
Enzymes and Requirements for DNA Replication
Key Enzymes and Substrates
DNA replication requires several enzymes and substrates:
DNA Polymerases: Enzymes that synthesize new DNA strands by adding nucleotides to a pre-existing chain.
Deoxynucleoside Triphosphates (dNTPs): The building blocks for DNA synthesis (dATP, dCTP, dGTP, dTTP).
Template and Primer: DNA polymerases require a template strand and a primer with a free 3'-OH group to initiate synthesis.
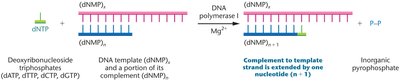
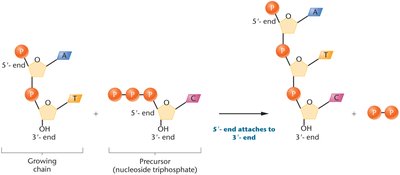
Bacterial DNA Polymerases
Bacteria possess five DNA polymerases (I-V), each with specialized functions:
DNA Polymerase I: Removes RNA primers and fills in gaps.
DNA Polymerase III: Main enzyme for DNA replication; a large holoenzyme complex.
DNA Polymerases II, IV, V: Involved in DNA repair and replication of damaged DNA.
All polymerases synthesize DNA in the 5' to 3' direction and have 3' to 5' exonuclease (proofreading) activity.
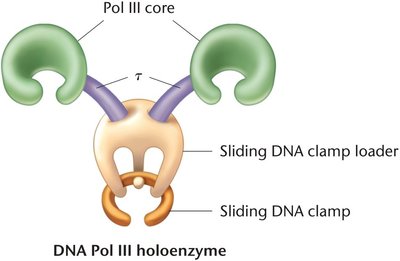
Mechanism of DNA Replication
Steps in DNA Replication
DNA replication involves several coordinated steps:
Unwinding the double helix (helicase)
Stabilizing single strands (single-strand binding proteins)
Relieving supercoiling (DNA gyrase/topoisomerase)
Synthesizing RNA primers (primase)
Elongating new DNA strands (DNA polymerases)
Removing RNA primers (DNA polymerase I)
Filling gaps and joining fragments (DNA ligase)
Proofreading and error correction
Initiation: Primer Synthesis
DNA polymerases cannot initiate synthesis de novo; they require a short RNA primer synthesized by primase. The primer provides a free 3'-OH group for DNA polymerase to extend.
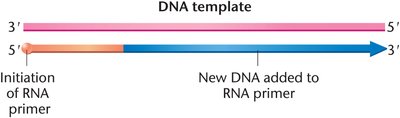
Leading and Lagging Strand Synthesis
Because DNA polymerases can only synthesize in the 5' to 3' direction, replication is continuous on the leading strand and discontinuous on the lagging strand, producing short Okazaki fragments.
Leading Strand: Synthesized continuously toward the replication fork.
Lagging Strand: Synthesized discontinuously away from the fork in short fragments.
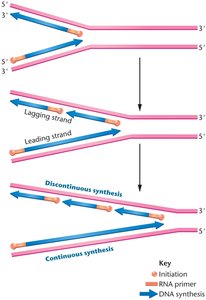
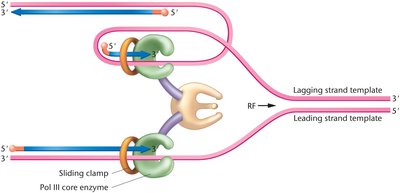
Coordination at the Replication Fork
The lagging strand template is looped so that both DNA polymerase complexes move in the same physical direction, ensuring coordinated synthesis of both strands.
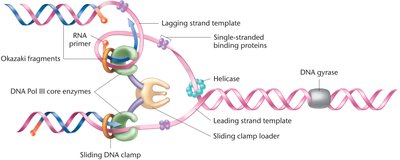

DNA Replication in Eukaryotes
Eukaryotic DNA Polymerases
Eukaryotes have multiple DNA polymerases with specialized functions:
Polymerase α: Associates with primase to synthesize RNA-DNA primers; low processivity.
Polymerase δ: Synthesizes DNA on the lagging strand and participates in repair.
Polymerase ε: Synthesizes DNA on the leading strand and participates in repair.
Other polymerases are involved in DNA repair and bypass of damaged DNA.
End Replication Problem and Telomeres
Linear chromosomes present a unique challenge: after removal of the RNA primer at the 5' end, there is no free 3'-OH group for DNA polymerase to fill in the gap, leading to progressive shortening of chromosomes with each cell division.
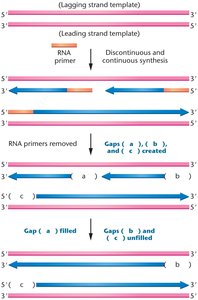
Telomeres and Telomerase
Telomeres are repetitive, non-coding DNA sequences at the ends of chromosomes (e.g., 5'-TTAGGG-3' in humans) that protect chromosome integrity. Telomerase is a ribonucleoprotein enzyme that extends telomeres by reverse transcription, using its RNA component as a template.
Telomerase binds to the G-rich overhang and adds repeats, preventing chromosome shortening.
After extension, primase and DNA polymerase fill in the complementary strand, and DNA ligase seals the gap.
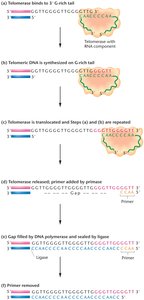
Telomeres in Disease, Aging, and Cancer
Shortened telomeres are associated with cellular aging and certain diseases. Most somatic cells lack telomerase activity, leading to eventual senescence. In contrast, cancer cells often reactivate telomerase, contributing to their immortality.
Summary Table: Key Enzymes and Functions in DNA Replication
Enzyme/Protein | Function |
|---|---|
DNA Polymerase III (prokaryotes) | Main DNA synthesis |
DNA Polymerase I (prokaryotes) | Removes RNA primers, fills gaps |
Primase | Synthesizes RNA primers |
Helicase | Unwinds DNA helix |
Single-Strand Binding Proteins | Stabilize unwound DNA |
DNA Gyrase (Topoisomerase) | Relieves supercoiling |
DNA Ligase | Joins Okazaki fragments |
Telomerase | Extends telomeres in eukaryotes |
