 Back
BackDNA Replication: Mechanisms, Enzymes, and Experimental Evidence
Study Guide - Smart Notes

DNA Replication: Mechanisms and Experimental Evidence
Overview of DNA Replication
DNA replication is a fundamental process in genetics, ensuring that genetic information is accurately transmitted from one generation to the next. This process occurs during the S phase of the cell cycle and involves the duplication of chromosomes, followed by their segregation during cell division.
Structure of DNA
Watson and Crick Model
The structure of DNA was elucidated by Watson and Crick in 1953, building on the work of Chargaff and Franklin. DNA is a right-handed double helix with a sugar-phosphate backbone on the outside and complementary base pairs (A-T and G-C) on the inside, held together by hydrogen bonds. The two strands are antiparallel and serve as templates for replication.
Properties of Genetic Material
Essential Features
Accurate Replication: Genetic material must replicate precisely to ensure progeny inherit the same information as the parent.
Information Storage: DNA contains instructions for cellular structure, function, and development.
Capacity for Change: DNA must be capable of mutation to allow for heritable variation and evolution.
Mechanisms of DNA Replication
Models of Replication
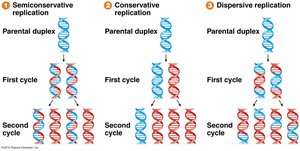
Three models were proposed for DNA replication:
Semi-conservative: Each daughter DNA molecule consists of one parental and one newly synthesized strand.
Conservative: The parental molecule remains intact, and an entirely new molecule is synthesized.
Dispersive: Parental and new DNA are interspersed in both strands.
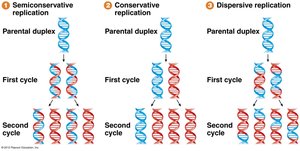
Experimental Evidence: Meselson-Stahl Experiment
The Meselson-Stahl experiment (1958) provided strong evidence for the semi-conservative model. By growing E. coli in media containing heavy (15N) and light (14N) nitrogen isotopes, they demonstrated that after one replication cycle, DNA molecules were of intermediate density, ruling out the conservative model. After a second cycle, both intermediate and light DNA were observed, ruling out the dispersive model.


Enzymology of DNA Replication
DNA Polymerases
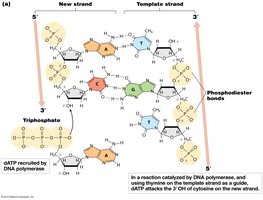
DNA polymerases are the enzymes responsible for synthesizing new DNA strands. They catalyze the formation of phosphodiester bonds between the 3' OH group of the last nucleotide and the 5' phosphate of the incoming deoxynucleoside triphosphate (dNTP).
Template Requirement: DNA polymerases require a template strand and a primer with a free 3' OH group.
Directionality: DNA synthesis occurs in the 5' to 3' direction only.
Proofreading: Many DNA polymerases possess 3' to 5' exonuclease activity, allowing them to remove incorrectly paired bases and increase fidelity.
Leading and Lagging Strand Synthesis
At the replication fork, one strand (the leading strand) is synthesized continuously, while the other (the lagging strand) is synthesized discontinuously in short fragments called Okazaki fragments. These fragments are later joined by DNA ligase.
Initiation of DNA Replication in Prokaryotes
Origin of Replication (oriC)
In E. coli, replication begins at a single origin (oriC). Initiator proteins (DnaA) unwind the DNA, forming a replication bubble. DNA helicases further unwind the DNA, and primase synthesizes short RNA primers to provide free 3' OH groups for DNA polymerase III.
Replication Fork Dynamics
Single-stranded binding proteins stabilize the unwound DNA, and DNA polymerase III extends the new DNA strand from the RNA primer. On the lagging strand, DNA polymerase I replaces RNA primers with DNA, and DNA ligase seals the nicks between Okazaki fragments.
DNA Replication in Eukaryotes
Multiple Origins of Replication
Eukaryotic chromosomes are linear and contain multiple origins of replication to ensure timely duplication of large genomes. The basic enzymatic mechanisms are conserved between prokaryotes and eukaryotes.
Telomeres and Telomerase
Linear chromosomes have telomeres—repetitive, non-coding sequences at their ends that protect against loss of genetic information during replication. The enzyme telomerase extends telomeres using an RNA template, preventing chromosome shortening in germ cells and some stem cells.
Summary Table: Key Enzymes and Functions in DNA Replication
Enzyme/Protein | Function |
|---|---|
DNA Polymerase III | Main enzyme for DNA synthesis (prokaryotes) |
DNA Polymerase I | Removes RNA primers and fills gaps with DNA |
Primase | Synthesizes RNA primers |
DNA Ligase | Joins Okazaki fragments |
Helicase | Unwinds DNA double helix |
Single-Stranded Binding Protein | Stabilizes unwound DNA |
Topoisomerase | Relieves supercoiling ahead of replication fork |
Telomerase | Extends telomeres in eukaryotes |
Key Equations
Phosphodiester Bond Formation:
Base Pairing Rules:
Conclusion
DNA replication is a highly regulated, accurate process essential for genetic continuity. The semi-conservative mechanism, elucidated by the Meselson-Stahl experiment, and the coordinated action of multiple enzymes ensure faithful duplication of genetic material in both prokaryotes and eukaryotes.
