 Back
BackDNA Replication: Mechanisms, Enzymes, and Experimental Evidence
Study Guide - Smart Notes

DNA Replication: Mechanisms, Enzymes, and Experimental Evidence
Introduction to DNA Replication
DNA replication is a fundamental process in all living organisms, ensuring the accurate transmission of genetic information from one generation to the next. The structure of DNA, as elucidated by Watson and Crick, immediately suggested a mechanism for its replication, where each strand serves as a template for the synthesis of a new complementary strand.
Modes of DNA Replication
Conservative, Semiconservative, and Dispersive Replication
There are three theoretical models for how DNA could replicate:
Semi-conservative replication: Each daughter DNA molecule consists of one parental (old) strand and one newly synthesized strand.
Conservative replication: The parental DNA molecule remains intact, and an entirely new molecule is synthesized.
Dispersive replication: Both daughter molecules contain interspersed segments of old and new DNA.

The Meselson-Stahl Experiment
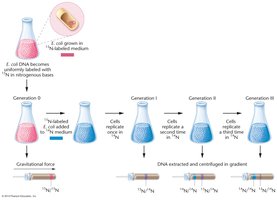
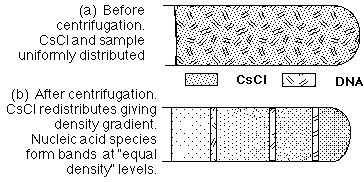
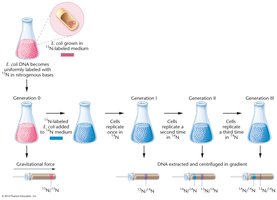
The Meselson-Stahl experiment provided definitive evidence for the semiconservative model of DNA replication. They used isotopic labeling with 15N and 14N to distinguish old and new DNA strands in E. coli and analyzed the DNA using cesium chloride (CsCl) density gradient centrifugation.
Generation 0: Bacteria grown in 15N medium; all DNA is heavy.
Generation 1: After one division in 14N medium, all DNA is of intermediate density (one old and one new strand).
Generation 2: After a second division, two bands appear: one light (14N/14N) and one intermediate (15N/14N), consistent only with semiconservative replication.

Enzymes and Steps in DNA Replication
Key Enzymes Involved
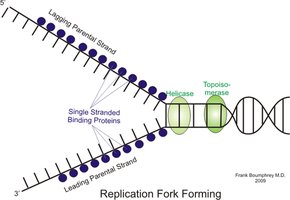
Helicase: Unwinds the DNA double helix at the origin of replication.
Single-Stranded Binding Proteins (SSBPs): Stabilize unwound DNA strands.
Topoisomerase (DNA gyrase in E. coli): Relieves supercoiling tension ahead of the replication fork.
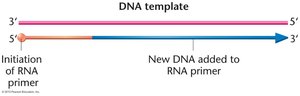
Primase: Synthesizes short RNA primers to provide a 3'-OH group for DNA polymerases.
DNA Polymerase III: Main enzyme for DNA synthesis; extends the new DNA strand from the RNA primer.
DNA Polymerase I: Removes RNA primers and fills in the gaps with DNA.
DNA Ligase: Seals nicks in the DNA backbone by forming phosphodiester bonds.
Six Steps of DNA Replication in E. coli
DNA is unwound: Helicase unwinds the DNA at the origin, and SSBPs stabilize the single strands.
Tension is reduced: Topoisomerase (DNA gyrase) relieves supercoiling ahead of the fork.
RNA primers are synthesized: Primase synthesizes short RNA primers on both strands.
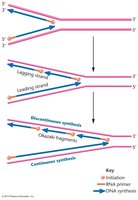
Leading and lagging strands are polymerized: DNA pol III synthesizes the leading strand continuously and the lagging strand discontinuously (Okazaki fragments).
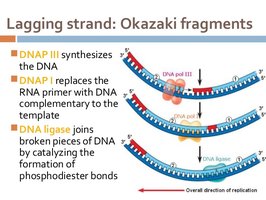
RNA primers are removed and gaps filled: DNA pol I removes RNA primers and fills in with DNA.
DNA nick sealing: DNA ligase seals the nicks between Okazaki fragments, completing the backbone.
Properties of DNA Polymerases
Comparison of DNA Polymerase I and III
DNA polymerases are essential for DNA synthesis, but they have distinct properties and functions:
Property | DNA Pol I | DNA Pol III |
|---|---|---|
Initiation of chain synthesis | – | – |
5'→3' polymerization | + | + |
3'→5' exonuclease activity | + | + |
5'→3' exonuclease activity | + | – |
Molecules of polymerase/cell | 400 | 15 |
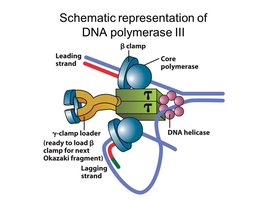
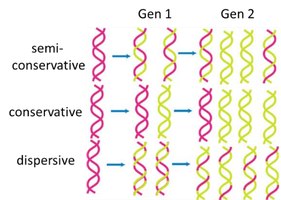
DNA Polymerase III Holoenzyme Structure
DNA pol III is a holoenzyme composed of multiple subunits, each with a specific function. It is responsible for the bulk of DNA synthesis during replication.

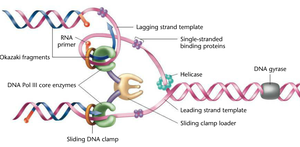
Mechanics of Leading and Lagging Strand Synthesis
Continuous vs. Discontinuous Synthesis
Leading strand: Synthesized continuously in the direction of the replication fork movement.
Lagging strand: Synthesized discontinuously in short segments called Okazaki fragments, which are later joined together.

Okazaki Fragments, DNA Ligase, and Primer RNA
Okazaki fragments: Short DNA fragments synthesized on the lagging strand.
DNA ligase: Enzyme that joins Okazaki fragments by forming phosphodiester bonds.
Primer RNA: Short RNA sequence synthesized by primase to provide a starting point for DNA synthesis.

Bidirectional Replication and the Replicon
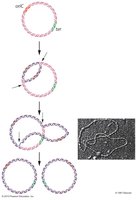
Bidirectional Synthesis in Circular Chromosomes
In E. coli, DNA replication begins at a single origin (oriC) and proceeds bidirectionally, forming two replication forks. The entire circular chromosome acts as a single replicon.
Summary Table: Key Terms and Functions
Term | Definition/Function |
|---|---|
Helicase | Unwinds DNA double helix |
SSBP | Stabilizes single-stranded DNA |
Topoisomerase (Gyrase) | Relieves supercoiling tension |
Primase | Synthesizes RNA primers |
DNA pol III | Main DNA synthesizing enzyme |
DNA pol I | Removes RNA primers, fills gaps |
DNA ligase | Seals nicks in DNA backbone |
Additional info:
DNA replication is highly accurate due to proofreading activities of DNA polymerases.
Replication in eukaryotes is more complex, involving multiple origins of replication and additional regulatory mechanisms.
